Article
|
Open Access
Featured
-
-
Article
| Open Access A revised biosynthetic pathway for the cofactor F420 in prokaryotes
A revised biosynthetic pathway for the cofactor F420 in prokaryotes
Cofactor F420 plays crucial roles in bacterial and archaeal metabolism, but its biosynthetic pathway is not fully understood. Here, the authors present the structure of one of the enzymes and provide experimental evidence for a substantial revision of the pathway, including the identification of a new intermediate.
- Ghader Bashiri
- , James Antoney
- & Colin J. Jackson
-
Article
| Open Access A TetR-family transcription factor regulates fatty acid metabolism in the archaeal model organism Sulfolobus acidocaldarius
A TetR-family transcription factor regulates fatty acid metabolism in the archaeal model organism Sulfolobus acidocaldarius
Certain archaea appear to metabolize fatty acids, but the regulation of these pathways is unclear. Here, Wang et al. provide genetic, functional and structural evidence supporting that a TetR-family transcriptional regulator is involved in regulation of fatty acid metabolism in Sulfolobus acidocaldarius.
- Kun Wang
- , David Sybers
- & Eveline Peeters
-
Article
| Open Access Mixing of meteoric and geothermal fluids supports hyperdiverse chemosynthetic hydrothermal communities
Mixing of meteoric and geothermal fluids supports hyperdiverse chemosynthetic hydrothermal communities
Chemosynthetic microbial communities in hydrothermal environments receiving meteoric and geothermal fluids are understudied. Here, Colman et al. use metagenomics to study one such community from a hot spring at Yellowstone National Park, revealing exceptional biodiversity and unique functional potential.
- Daniel R. Colman
- , Melody R. Lindsay
- & Eric S. Boyd
-
Article
| Open Access Metabolic diversity within the globally abundant Marine Group II Euryarchaea offers insight into ecological patterns
Metabolic diversity within the globally abundant Marine Group II Euryarchaea offers insight into ecological patterns
The physiology and ecology of many yet-uncultured microorganisms, such as the Marine Group II Euryarchaea (MGII), is unclear. Here, Benjamin Tully analyses 250 MGII genomes, identifies 17 distinct subclades, and provides a detailed view of the metabolic potential and distribution of these archaea.
- Benjamin J. Tully
-
Article
| Open Access CRISPR analysis suggests that small circular single-stranded DNA smacoviruses infect Archaea instead of humans
CRISPR analysis suggests that small circular single-stranded DNA smacoviruses infect Archaea instead of humans
Smacoviruses are found in the intestinal tract of humans and animals but their precise host remains elusive. Here, the authors identify smacovirus-matching CRISPR spacer sequences in the faecal archaeon Candidatus Methanomassiliicoccus intestinalis, implicating Archaea as a potential host.
- César Díez-Villaseñor
- & Francisco Rodriguez-Valera
-
Article
| Open Access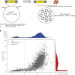 The essential genome of the crenarchaeal model Sulfolobus islandicus
The essential genome of the crenarchaeal model Sulfolobus islandicus
Sulfolobus islandicus is a model organism within the TACK superphylum of the Archaea. Here, the authors perform a genome-wide analysis of essential genes in this organism, show that the proteinaceous S-layer is not essential, and explore potential stages of evolution of the essential gene repertoire in Archaea.
- Changyi Zhang
- , Alex P. R. Phillips
- & Rachel J. Whitaker
-
Article
| Open Access Genomic inference of the metabolism and evolution of the archaeal phylum Aigarchaeota
Genomic inference of the metabolism and evolution of the archaeal phylum Aigarchaeota
The phylum of archaea Aigarchaeota is poorly characterized due to limited genomic sampling. Here, Hua and colleagues use genome-resolved metagenome sequencing to reconstruct six hot spring strains of Aigarchaeota and then infer their metabolism and evolutionary history.
- Zheng-Shuang Hua
- , Yan-Ni Qu
- & Wen-Jun Li
-
Article
| Open Access Rpn11-mediated ubiquitin processing in an ancestral archaeal ubiquitination system
Rpn11-mediated ubiquitin processing in an ancestral archaeal ubiquitination system
Ubiquitin modification also occurs in archaea. Here, the authors characterize an archaeal ancestral ubiquitination system, present the crystal structure of the archaeal deubiquitinase Rpn11 from Caldiarchaeum subterraneum bound to ubiquitin and provide insights into evolutionary relationships.
- Adrian C. D. Fuchs
- , Lorena Maldoner
- & Jörg Martin
-
Article
| Open Access Unifying the global phylogeny and environmental distribution of ammonia-oxidising archaea based on amoA genes
Unifying the global phylogeny and environmental distribution of ammonia-oxidising archaea based on amoA genes
Ammonia-oxidising archaea (AOA) were only discovered a little over a decade ago and remain poorly characterized despite their ubiquity and importance for nitrogen cycling. Here, the authors define a taxonomy of AOA based on a resolved amoA phylogeny and describe emergent global patterns in AOA diversity.
- Ricardo J. Eloy Alves
- , Bui Quang Minh
- & Christa Schleper
-
Article
| Open Access Assembly of 913 microbial genomes from metagenomic sequencing of the cow rumen
Assembly of 913 microbial genomes from metagenomic sequencing of the cow rumen
Microbes in the cow rumen are crucial for the breakdown of plant material. Here, Stewart et al. assemble over 900 bacterial and archaeal genomes from the cow rumen microbiome, revealing new species and genes encoding enzymes with potential roles in carbohydrate metabolism.
- Robert D. Stewart
- , Marc D. Auffret
- & Mick Watson
-
Article
| Open Access Biological methane production under putative Enceladus-like conditions
Biological methane production under putative Enceladus-like conditions
Many methanogenic archaea use H2 and CO2 to produce methane. Here, Taubner et al. show that Methanothermococcus okinawensis produces methane under conditions extrapolated for Saturn’s icy moon, Enceladus, and estimate that serpentinization may produce sufficient H2 for biological methane production.
- Ruth-Sophie Taubner
- , Patricia Pappenreiter
- & Simon K.-M. R. Rittmann
-
Article
| Open Access The transcript cleavage factor paralogue TFS4 is a potent RNA polymerase inhibitor
The transcript cleavage factor paralogue TFS4 is a potent RNA polymerase inhibitor
Transcript cleavage factors such as eukaryotic TFIIS assist the resumption of transcription following RNA pol II backtracking. Here the authors find that one of the Sulfolobus solfataricus TFIIS homolog—TFS4—has evolved into a potent RNA polymerase inhibitor potentially involved in antiviral defense.
- Thomas Fouqueau
- , Fabian Blombach
- & Finn Werner
-
Article
| Open Access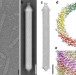 Unique architecture of thermophilic archaeal virus APBV1 and its genome packaging
Unique architecture of thermophilic archaeal virus APBV1 and its genome packaging
The rod-shaped virus APBV1 is among the most thermostable viruses known. Here, Ptchelkine et al. determine its structure at near-atomic resolution, show that the DNA is packed as left-handed superhelix and identify extended hydrophobic interfaces that likely contribute to the extreme thermostability of the capsid.
- Denis Ptchelkine
- , Ashley Gillum
- & Juha T. Huiskonen
-
Article
| Open Access ‘ARMAN’ archaea depend on association with euryarchaeal host in culture and in situ
‘ARMAN’ archaea depend on association with euryarchaeal host in culture and in situ
In the absence of complete genomes, the metabolic capabilities of uncultured ARMAN-like archaea have been uncertain. Here, Golyshina et al. apply an enrichment culture technique and find that the ungapped genome of the ARMAN-like archaeon Mia14 has lost key metabolic pathways, suggesting dependence on the host archaeon Cuniculiplasma divulgatum.
- Olga V. Golyshina
- , Stepan V. Toshchakov
- & Peter N. Golyshin
-
Article
| Open Access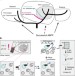 Activity-based protein profiling as a robust method for enzyme identification and screening in extremophilic Archaea
Activity-based protein profiling as a robust method for enzyme identification and screening in extremophilic Archaea
Activity-based protein profiling (ABPP) is a chemical proteomics method to profile activity states of enzymes under physiological conditions. Here the authors show that ABPP can be applied to archaeal serine hydrolases in the model organismSulfolobus acidocaldariusand can be used to identify novel putative serine hydrolases.
- Susanne Zweerink
- , Verena Kallnik
- & Markus Kaiser
-
Article
| Open Access A RuBisCO-mediated carbon metabolic pathway in methanogenic archaea
A RuBisCO-mediated carbon metabolic pathway in methanogenic archaea
Although not photosynthetic, some archaea possess RuBisCO, one of the enzymes characteristic of the photosynthetic Calvin-Benson cycle, but apparently lack another one, phosphoribulokinase (PRK). Here the authors describe a carbon metabolic pathway in methanogenic archaea, involving RuBisCO and PRK.
- Takunari Kono
- , Sandhya Mehrotra
- & Hiroki Ashida
-
Article
| Open Access Repression of RNA polymerase by the archaeo-viral regulator ORF145/RIP
Repression of RNA polymerase by the archaeo-viral regulator ORF145/RIP
How archaeal viruses perturb host transcription machinery is poorly understood. Here, the authors provide evidence that the archaeo-viral transcription factor ORF145/RIP targets host RNA polymerase, repressing its activity.
- Carol Sheppard
- , Fabian Blombach
- & Finn Werner
-
Article
| Open Access An archaeal ADP-dependent serine kinase involved in cysteine biosynthesis and serine metabolism
An archaeal ADP-dependent serine kinase involved in cysteine biosynthesis and serine metabolism
Archaea metabolism has unique adaptations to hostile environments. Here Makino et al. describe an unusual ADP-dependent kinase that phosphorylates free serine to O-phosphoserine and participates in an additional cysteine biosynthetic pathway in the archaeon Thermococcus kodakarensis.
- Yuki Makino
- , Takaaki Sato
- & Haruyuki Atomi
-
Article
| Open Access Genomics-informed isolation and characterization of a symbiotic Nanoarchaeota system from a terrestrial geothermal environment
Genomics-informed isolation and characterization of a symbiotic Nanoarchaeota system from a terrestrial geothermal environment
Many microbial lineages have not yet been cultured, which hampers our understanding of their physiology. Here, Wurch et al. use single-cell genomics to infer cultivation conditions for the isolation of a tiny ectosymbiotic nanoarchaeon and its crenarchaeota host from a geothermal spring.
- Louie Wurch
- , Richard J. Giannone
- & Mircea Podar
-
Article
| Open Access Genomic and transcriptomic evidence for scavenging of diverse organic compounds by widespread deep-sea archaea
Genomic and transcriptomic evidence for scavenging of diverse organic compounds by widespread deep-sea archaea
The contribution of marine archaea to the ocean's carbon cycle is unclear. Here, Li et al. analyse the genomes and transcriptomes from five deep-sea archaeal groups to reveal their metabolic characteristics, suggesting a crucial role in modulating the carbon cycle in deep oceans.
- Meng Li
- , Brett J. Baker
- & Gregory J. Dick
-
Article
| Open Access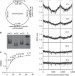 Activation of a dormant replication origin is essential for Haloferax mediterranei lacking the primary origins
Activation of a dormant replication origin is essential for Haloferax mediterranei lacking the primary origins
Archaea use multiple origins for chromosome replication, but some potential origins appear to be inactive during the genome replication. Here, Yang et al. show that when active origins are deleted from Haloferax mediterranei, a dormant origin is activated and essential, suggesting an origin-dependent replication.
- Haibo Yang
- , Zhenfang Wu
- & Hua Xiang
-
Article
| Open Access Targeted diversity generation by intraterrestrial archaea and archaeal viruses
Targeted diversity generation by intraterrestrial archaea and archaeal viruses
Diversity-generating retroelements (DGRs) are genetic elements that introduce sequence variation within target genes in bacteria and their viruses. Here, Paul et al. report the discovery of DGRs in an archaeal virus and in two archaea from marine and terrestrial subsurface environments, respectively.
- Blair G. Paul
- , Sarah C. Bagby
- & David L. Valentine
-
Article |
 Diverse uncultivated ultra-small bacterial cells in groundwater
Diverse uncultivated ultra-small bacterial cells in groundwater
Little is known about certain bacterial phyla because of our current inability to grow them in the lab. Here, Luef et al.combine metagenomics and ultrastuctural analyses to show that some of these bacteria have a very small cell size, tightly packed DNA, few ribosomes and diverse pili-like structures.
- Birgit Luef
- , Kyle R. Frischkorn
- & Jillian F. Banfield
-
Article |
 Complete architecture of the archaeal RNA polymerase open complex from single-molecule FRET and NPS
Complete architecture of the archaeal RNA polymerase open complex from single-molecule FRET and NPS
The archaeal RNA transcription machinery does not have a dedicated helicase factor. Here, the authors report the three-dimensional architecture of the open complex of DNA, RNA polymerase and its associated factors from M. jannaschii, providing a possible mechanism for promoter DNA melting.
- Julia Nagy
- , Dina Grohmann
- & Jens Michaelis
-
Article |
 Biology of a widespread uncultivated archaeon that contributes to carbon fixation in the subsurface
Biology of a widespread uncultivated archaeon that contributes to carbon fixation in the subsurface
Research on microbes that inhabit the Earth's subsurface is mostly based on metagenomic information only. Here, Probst et al. combine metagenomics with ultrastructural and functional analyses to study the biology of a group of uncultivated subsurface archaea, the SM1 Euryarchaeon lineage.
- Alexander J. Probst
- , Thomas Weinmaier
- & Christine Moissl-Eichinger
-
Article |
 The X-ray crystal structure of the euryarchaeal RNA polymerase in an open-clamp configuration
The X-ray crystal structure of the euryarchaeal RNA polymerase in an open-clamp configuration
Archaeal and eukaryotic RNA polymerases (RNAP) have conserved functional and structural similarities. Here, Jun et al.solve the first structure of a euryarchaeal RNAP in the open clamp conformation and identify insertions that may have evolved in eukaryotic Pol II to bind unique transcription factors.
- Sung-Hoon Jun
- , Akira Hirata
- & Katsuhiko S. Murakami
-
Article
| Open Access Methylotrophic methanogenic Thermoplasmata implicated in reduced methane emissions from bovine rumen
Methylotrophic methanogenic Thermoplasmata implicated in reduced methane emissions from bovine rumen
Rumen methanogenic archaea are major sources of methane emissions and potential targets for methane mitigation strategies. Poulsen et al.now show that dietary rapeseed oil (RSO) supplementation can reduce the abundance of methanogenic Thermoplasmata archaea inhabiting the bovine rumen.
- Morten Poulsen
- , Clarissa Schwab
- & Tim Urich
-
Article |
 Identification and characterization of a multidomain hyperthermophilic cellulase from an archaeal enrichment
Identification and characterization of a multidomain hyperthermophilic cellulase from an archaeal enrichment
Archaea are microorganisms that use a wide range of carbon and energy sources. Grahamet al. describe an archaeal consortium that can grow at temperatures above 90 °C using crystalline cellulose as a carbon source, with potential applications in enzymatic degradation under extreme conditions.
- Joel E. Graham
- , Melinda E. Clark
- & Frank T. Robb

 A new type of DNA phosphorothioation-based antiviral system in archaea
A new type of DNA phosphorothioation-based antiviral system in archaea