Featured
-
-
Article
| Open Access Exposure to environmental pollutants selects for xenobiotic-degrading functions in the human gut microbiome
Exposure to environmental pollutants selects for xenobiotic-degrading functions in the human gut microbiome
In this study, the authors employ metagenomics to explore the impact of environmental pollution on the human gut microbiome using samples from a cohort living in a very polluted area in Southern Italy, showing that pollutants degradation genes are more abundant in subjects with higher blood levels of those specific xenobiotics.
- Francesca De Filippis
- , Vincenzo Valentino
- & Danilo Ercolini
-
Article
| Open Access Relative dispersion ratios following fecal microbiota transplant elucidate principles governing microbial migration dynamics
Relative dispersion ratios following fecal microbiota transplant elucidate principles governing microbial migration dynamics
Microbial migration profoundly impacts ecosystems. Here, the authors introduce a statistical approach to explore microbial dispersion following fecal microbiota transplant, uncovering dependencies between colonizing taxa, with insights into community dynamics.
- Yadid M. Algavi
- & Elhanan Borenstein
-
Article
| Open Access Gut microbiome composition and metabolic activity in women with diverticulitis
Gut microbiome composition and metabolic activity in women with diverticulitis
Here, the authors present a multi-omics examination of stool samples obtained from individuals with diverticulitis and controls, uncovering disruptions in the balance of microbial composition and metabolites, as well as co-occurring microbe-metabolite associations relevant to the disease.
- Wenjie Ma
- , Yiqing Wang
- & Andrew T. Chan
-
Article
| Open Access CoCas9 is a compact nuclease from the human microbiome for efficient and precise genome editing
CoCas9 is a compact nuclease from the human microbiome for efficient and precise genome editing
Cas9 nucleases hold clinical significance for genome editing therapies. Here the authors characterize CoCas9, a compact, efficient and precise Cas9 from the human microbiome, and show that delivery via AAV vectors enables efficient editing in the mouse retina, expanding the genome editing toolbox.
- Eleonora Pedrazzoli
- , Michele Demozzi
- & Anna Cereseto
-
Article
| Open Access Freshwater genome-reduced bacteria exhibit pervasive episodes of adaptive stasis
Freshwater genome-reduced bacteria exhibit pervasive episodes of adaptive stasis
Here, by applying evolutionary genomics approaches to metagenomics data of lake microbiomes, the authors reveal that freshwater species with small genomes face extended periods with their niche adaptation capabilities frozen.
- Lucas Serra Moncadas
- , Cyrill Hofer
- & Adrian-Stefan Andrei
-
Article
| Open Access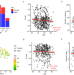 Consistent signatures in the human gut microbiome of old- and young-onset colorectal cancer
Consistent signatures in the human gut microbiome of old- and young-onset colorectal cancer
The rising incidence of young-onset sporadic colorectal cancer (yCRC) is global concern. Here, leveraging a substantial number of deep sequencing metagenomes, the authors show striking similarities in gut microbial patterns at both the taxonomic and selected gene marker levels between yCRC and old-onset CRC.
- Youwen Qin
- , Xin Tong
- & Pei-Rong Ding
-
Article
| Open Access Integrating taxonomic signals from MAGs and contigs improves read annotation and taxonomic profiling of metagenomes
Integrating taxonomic signals from MAGs and contigs improves read annotation and taxonomic profiling of metagenomes
Metagenomic taxonomic profiling usually relies either on reads or assembled contigs/MAGs. Here, authors present RAT, a tool that integrates taxonomic signals from reads, contigs, and MAGs into one profile with high precision and sensitivity. RAT provides a comprehensive view of the microbiome.
- Ernestina Hauptfeld
- , Nikolaos Pappas
- & F. A. Bastiaan von Meijenfeldt
-
Article
| Open Access Diversity and potential host-interactions of viruses inhabiting deep-sea seamount sediments
Diversity and potential host-interactions of viruses inhabiting deep-sea seamount sediments
Little is known about viral communities in deep-sea seamounts. In this study, the authors performed metagenomic and virome analysis from sediments in the western Pacific Ocean and characterize the diversity, distribution and potential ecological roles of viruses in deep-sea seamount sediments.
- Meishun Yu
- , Menghui Zhang
- & Min Jin
-
Comment
| Open Access All-inclusive nitrifiers in Antarctic soils
All-inclusive nitrifiers in Antarctic soils
Multidisciplinary culture-dependent and -independent techniques elucidate the unique microbial nitrogen cycle in nutrient-poor coastal Antarctica soils and reveal the contribution of novel key microbes to their nitrogen budget.
- Maximiliano Ortiz
-
Article
| Open Access Comparative characterization of the infant gut microbiome and their maternal lineage by a multi-omics approach
Comparative characterization of the infant gut microbiome and their maternal lineage by a multi-omics approach
Here, the authors employ multi-omics on a cohort comprising three generations of family members, showing that fecal microbiota populations, functions, and metabolome of infants vary greatly from their maternal lineage, exhibiting a less diverse microbiota and differences in various metabolite classes including short- and branched-chain fatty acids.
- Tomás Clive Barker-Tejeda
- , Elisa Zubeldia-Varela
- & Alma Villaseñor
-
Article
| Open Access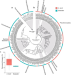 A genome-centric view of the role of the Acropora kenti microbiome in coral health and resilience
A genome-centric view of the role of the Acropora kenti microbiome in coral health and resilience
This study provides insights into the functional roles of microbial symbionts within the reef-building coral Acropora kenti. The findings reveal molecular mechanisms underpinning coral health and adaptation to local environmental stressors, which may support host resilience in the face of anthropogenic climate change and pollution.
- Lauren F. Messer
- , David G. Bourne
- & Gene W. Tyson
-
Article
| Open Access Genome-scale community modelling reveals conserved metabolic cross-feedings in epipelagic bacterioplankton communities
Genome-scale community modelling reveals conserved metabolic cross-feedings in epipelagic bacterioplankton communities
Identifying the metabolic interactions that underlie microbial communities is challenging. Here, the authors combine Tara Oceans -omics data with co-activity networks and genome-scale metabolic models to predict biotic interactions among planktonic prokaryotes in the upper ocean.
- Nils Giordano
- , Marinna Gaudin
- & Samuel Chaffron
-
Article
| Open Access Multi-omic integration of microbiome data for identifying disease-associated modules
Multi-omic integration of microbiome data for identifying disease-associated modules
Here, Muller et al. introduce MintTea, a method for analyzing multi-omic microbiome data and identifying disease-associated modules comprising mixed sets of features that collectively shift in disease, offering insights into microbiome-disease interactions.
- Efrat Muller
- , Itamar Shiryan
- & Elhanan Borenstein
-
Article
| Open Access Targeted metagenomics reveals association between severity and pathogen co-detection in infants with respiratory syncytial virus
Targeted metagenomics reveals association between severity and pathogen co-detection in infants with respiratory syncytial virus
The impact of other pathogens on disease outcome was studied in European infants with RSV infection. Additional viruses were commonly co-detected during infection but were weakly linked to severity. However, presence of Haemophilus bacteria strongly associated with severe cases.
- Gu-Lung Lin
- , Simon B. Drysdale
- & Andrew J. Pollard
-
Article
| Open Access BASALT refines binning from metagenomic data and increases resolution of genome-resolved metagenomic analysis
BASALT refines binning from metagenomic data and increases resolution of genome-resolved metagenomic analysis
Binning is an essential step in genome-resolved metagenomic analysis in which assembled contigs originating from the same source population are clustered. However it is challenging, especially for low abundance microbial species. Here the authors introduce a toolkit that integrates multiple prominent binning tools and AI for efficient and high-resolution recovery of non-redundant bins from short- and long-read metagenomic sequencing datasets.
- Zhiguang Qiu
- , Li Yuan
- & Ke Yu
-
Article
| Open Access The defensome of complex bacterial communities
The defensome of complex bacterial communities
Bacteria have evolved numerous innate and adaptive defence mechanisms. Here, Beavogui et al characterise the impact of biogeography, genetic mobility, and clustering in defense islands, on the defence systems of soil, marine, and human gut bacterial populations genomes.
- Angelina Beavogui
- , Auriane Lacroix
- & Pedro H. Oliveira
-
Article
| Open Access A metagenomic catalog of the early-life human gut virome
A metagenomic catalog of the early-life human gut virome
Here, the authors present a metagenomic catalogue of the early-life human gut virome including 160,478 non-redundant DNA and RNA viral sequences from 8,130 gut virus-like particles enriched or bulk metagenomes in the first three years of life.
- Shuqin Zeng
- , Alexandre Almeida
- & Shaopu Wang
-
Article
| Open Access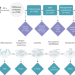 Exploring the gut DNA virome in fecal immunochemical test stool samples reveals associations with lifestyle in a large population-based study
Exploring the gut DNA virome in fecal immunochemical test stool samples reveals associations with lifestyle in a large population-based study
Here, the authors use fecal immunochemical test (FIT) samples from around 1000 individuals to characterize their gut virome, showing a diverse viral community, indicative of the individual lifestyle (smoking, fiber consumption and physical activity), thus highlighting FIT samples as a useful alternative for virome analyses.
- Paula Istvan
- , Einar Birkeland
- & Trine B. Rounge
-
Article
| Open Access Influence of microbiota-associated metabolic reprogramming on clinical outcome in patients with melanoma from the randomized adjuvant dendritic cell-based MIND-DC trial
Influence of microbiota-associated metabolic reprogramming on clinical outcome in patients with melanoma from the randomized adjuvant dendritic cell-based MIND-DC trial
MIND-DC was a randomized, placebo-controlled, phase 3 trial of adjuvant blood-derived natural dendritic cell (nDC)-based therapy in patients with stage III melanoma, showing that nDC-induced immune responses did not translate into survival benefit. Here the authors report that, despite randomization, baseline differences in fecal metagenomics and serum metabolomics profiles between treatment arms might have influenced the clinical outcome of the trial.
- Carolina Alves Costa Silva
- , Gianmarco Piccinno
- & I. Jolanda M. de Vries
-
Article
| Open Access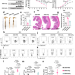 Porphyromonas gingivalis aggravates colitis via a gut microbiota-linoleic acid metabolism-Th17/Treg cell balance axis
Porphyromonas gingivalis aggravates colitis via a gut microbiota-linoleic acid metabolism-Th17/Treg cell balance axis
Periodontitis is closely linked with inflammatory bowel disease (IBD) and may have overlapping characteristics. Here the authors show that a periodontal pathogen P. gingivalis promotes intestinal inflammation by affecting the microbiome metabolite linoleic acid and Th17/Treg cell balance in the intestine.
- Lu Jia
- , Yiyang Jiang
- & Yi Liu
-
Article
| Open Access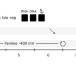 Mechanisms of extracellular electron transfer in anaerobic methanotrophic archaea
Mechanisms of extracellular electron transfer in anaerobic methanotrophic archaea
Anaerobic methanotrophic (ANME) archaea are uncultivated microbes that oxidize the greenhouse gas methane and engage in extracellular electron transfer with other microbes, metal oxides, and electrodes. Here, Ouboter et al. observe strong methane-dependent current associated with high enrichment of ANME archaea on the anode, and provide insights into the mechanisms underlying extracellular electron transfer.
- Heleen T. Ouboter
- , Rob Mesman
- & Cornelia U. Welte
-
Article
| Open Access A genome and gene catalog of the aquatic microbiomes of the Tibetan Plateau
A genome and gene catalog of the aquatic microbiomes of the Tibetan Plateau
The Tibetan Plateau is the largest plateau in the world and hosts a variety of aquatic ecosystems. Here, the authors present a gene and genome catalogue of Tibetan Plateau aquatic microbiomes, greatly expanding known taxonomic and functional diversity for the region and giving insights into its microbial biogeography.
- Mingyue Cheng
- , Shuai Luo
- & Kang Ning
-
Article
| Open Access HIV-associated gut microbial alterations are dependent on host and geographic context
HIV-associated gut microbial alterations are dependent on host and geographic context
Here, the authors compare the fecal microbial community of individuals in the U.S., Uganda, and Botswana, and identify significant bacterial taxa alterations with both treated and untreated HIV infection although with a high degree of uniqueness in each cohort, and also significant differences between populations that report men who have sex with men (MSM) behavior and non-MSM populations.
- Muntsa Rocafort
- , David B. Gootenberg
- & Douglas S. Kwon
-
Article
| Open Access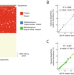 Longitudinal quantification of Bifidobacterium longum subsp. infantis reveals late colonization in the infant gut independent of maternal milk HMO composition
Longitudinal quantification of Bifidobacterium longum subsp. infantis reveals late colonization in the infant gut independent of maternal milk HMO composition
Here, the authors develop a high-throughput method to quantify Bifidobacterium longum subsp. infantis (BL. infantis), a proficient HMO-utilizer, from metagenomic sequencing, and applied it to a longitudinal cohort consisting of 21 mother-infant dyads, suggesting BL. infantis colonization to start late in the breast-feeding period.
- Dena Ennis
- , Shimrit Shmorak
- & Moran Yassour
-
Article
| Open Access Microbial decomposition of biodegradable plastics on the deep-sea floor
Microbial decomposition of biodegradable plastics on the deep-sea floor
It is unclear whether microbes can efficiently degrade biodegradable plastics in the extreme environmental conditions of the seafloor. Here, Omura et al. show that biodegradable plastics can be degraded by the action of microorganisms on the deep-sea floor, although with much less efficiency than in coastal settings.
- Taku Omura
- , Noriyuki Isobe
- & Tadahisa Iwata
-
Article
| Open Access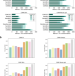 Effective binning of metagenomic contigs using contrastive multi-view representation learning
Effective binning of metagenomic contigs using contrastive multi-view representation learning
Here, the authors present COMEBin, a metagenomics binning method based on contrastive multi-view representation learning that uses data augmentation to generate multiple fragments (views) of each contig, resulting in high-quality embeddings of heterogeneous features. COMEBin outperforms state-of-the art binning methods, particularly in recovering near-complete genomes from real environmental samples.
- Ziye Wang
- , Ronghui You
- & Shanfeng Zhu
-
Article
| Open Access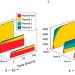 Host- plasmid network structure in wastewater is linked to antimicrobial resistance genes
Host- plasmid network structure in wastewater is linked to antimicrobial resistance genes
Authors apply theory and microbial ecology modelling to a wastewater sample, and show that antimicrobial resistance carrying plasmids interact with a higher number and more diverse range of bacteria than plasmids that do not carry resistance genes.
- Alice Risely
- , Arthur Newbury
- & Dirk Sanders
-
Article
| Open Access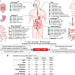 Defining the biogeographical map and potential bacterial translocation of microbiome in human ‘surface organs’
Defining the biogeographical map and potential bacterial translocation of microbiome in human ‘surface organs’
Given that the human body is composed of many microbial niches, and there have been few reports on the biogeography of the microbiome, the authors analyse the intra-individual inter-organ and intra-organ microbiome of seven surface organs of deceased individuals.
- Jun-Jun She
- , Wei-Xin Liu
- & Jun Yu
-
Article
| Open Access Functional host-specific adaptation of the intestinal microbiome in hominids
Functional host-specific adaptation of the intestinal microbiome in hominids
Here, Rühlemann et al. analyze the gut microbiome of wild-living African great apes (Gorillas, Bonobos, Chimpanzees) in comparison to that of humans, identifying host specific patterns and shared evolutionary conserved traits disrupted in humans.
- M. C. Rühlemann
- , C. Bang
- & A. Franke
-
Article
| Open Access The antibiotic resistance reservoir of the lung microbiome expands with age in a population of critically ill patients
The antibiotic resistance reservoir of the lung microbiome expands with age in a population of critically ill patients
Here, by performing tracheal aspirate RNA sequencing of critically ill patients, the authors find that older age associates with a greater number of detectably expressed antimicrobial resistance genes in the lower respiratory tract microbiome.
- Victoria T. Chu
- , Alexandra Tsitsiklis
- & Charles R. Langelier
-
Article
| Open Access Differential responses of the gut microbiome and resistome to antibiotic exposures in infants and adults
Differential responses of the gut microbiome and resistome to antibiotic exposures in infants and adults
Knowledge on how the gut microbiome and resistome responds to antibiotics across age remains limited. Here, using metagenomics data from Danish infants and young adults, the authors show that antibiotics have a more lasting impact on adults compared to infants.
- Xuanji Li
- , Asker Brejnrod
- & Søren Johannes Sørensen
-
Article
| Open Access Resin acids play key roles in shaping microbial communities during degradation of spruce bark
Resin acids play key roles in shaping microbial communities during degradation of spruce bark
The bark is the outermost defense of trees against microbial attack, largely due to toxicity of extractive compounds. Here, Ristinmaa et al. study microbial community dynamics and chemical changes during degradation of spruce bark over six months, showing that the microbial degradation of extractive compounds, such as resin acids, has a major role in shaping the microbial community.
- Amanda Sörensen Ristinmaa
- , Albert Tafur Rangel
- & Johan Larsbrink
-
Article
| Open Access Airway environment drives the selection of quorum sensing mutants and promote Staphylococcus aureus chronic lifestyle
Airway environment drives the selection of quorum sensing mutants and promote Staphylococcus aureus chronic lifestyle
This study by Ding et al reveals that the quorum-sensing dysfunction typically encountered in lung-adapted Staphylococcus aureus isolates could be selected by an enhanced ability to consume sialic acid released from airway mucins by the microbiota.
- Xiongqi Ding
- , Catherine Robbe-Masselot
- & Anne Jamet
-
Article
| Open Access Health-related quality of life is linked to the gut microbiome in kidney transplant recipients
Health-related quality of life is linked to the gut microbiome in kidney transplant recipients
Here, Swarte et al. use metagenomics to investigate the association between the gut microbiome and Health-Related Quality of Life (HRQoL) in kidney transplant recipients, showing evidence for the association of multiple taxonomic, metabolic and neuroactive pathways (gut brain modules) with lower HRQoL, together suggesting potential modifiable gut microbial factors to improve HRQoL.
- J. Casper Swarte
- , Tim J. Knobbe
- & Rinse K. Weersma
-
Article
| Open Access Bacterial genome size and gene functional diversity negatively correlate with taxonomic diversity along a pH gradient
Bacterial genome size and gene functional diversity negatively correlate with taxonomic diversity along a pH gradient
Bacterial functional diversity does not necessarily correlate with taxonomic diversity because average genome size may vary by community. Here, Wang et al. investigate bacterial communities along a natural pH gradient in forest soils, and find that average genome size and functional diversity decrease, whereas taxonomic diversity increases, as soil pH rises from acid to neutral.
- Cong Wang
- , Qing-Yi Yu
- & Cheng Gao
-
Article
| Open Access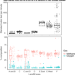 Microbial methane cycling in a landfill on a decadal time scale
Microbial methane cycling in a landfill on a decadal time scale
Microbial degradation of organic matter in landfills generates methane, a potent greenhouse gas. Here, Grégoire et al. use metagenomic approaches to study microbial methane cycling in waste landfilled over 39 years, highlighting the importance of specific microbial lineages and methane oxidation in the absence of oxygen.
- Daniel S. Grégoire
- , Nikhil A. George
- & Laura A. Hug
-
Article
| Open Access Infant microbiome cultivation and metagenomic analysis reveal Bifidobacterium 2’-fucosyllactose utilization can be facilitated by coexisting species
Infant microbiome cultivation and metagenomic analysis reveal Bifidobacterium 2’-fucosyllactose utilization can be facilitated by coexisting species
Here, Lou et al. apply metagenomics and microbiome cultivation to infant fecal samples and uncover co-existing members encoding extracellular fucosidases that initiate 2’-fucosyllactose (2’FL) breakdown and can promote extensive growth of Bifidobacterium breve.
- Yue Clare Lou
- , Benjamin E. Rubin
- & Jillian F. Banfield
-
Article
| Open Access A genomic catalogue of soil microbiomes boosts mining of biodiversity and genetic resources
A genomic catalogue of soil microbiomes boosts mining of biodiversity and genetic resources
Soil conceals a vast realm of unexplored microbes, often referred to as the “microbial dark matter.” This hidden universe boasts a rich tapestry of microbial and genetic biodiversity. Here, the authors introduce the SMAG catalogue, comprising of 40,039 metagenome-assembled genomes from 3304 soil metagenomes, and uncovering 21,077 species-level genome bins.
- Bin Ma
- , Caiyu Lu
- & Jianming Xu
-
Article
| Open Access Wastewater sequencing reveals community and variant dynamics of the collective human virome
Wastewater sequencing reveals community and variant dynamics of the collective human virome
Tisza et al. carry out a sequencing-based analysis of wastewater samples from major cities, to detect and quantify hundreds of distinct pathogenic viruses, finding striking correlations between virus abundance and local clinical cases.
- Michael Tisza
- , Sara Javornik Cregeen
- & Anthony W. Maresso
-
Article
| Open Access Globally distributed Myxococcota with photosynthesis gene clusters illuminate the origin and evolution of a potentially chimeric lifestyle
Globally distributed Myxococcota with photosynthesis gene clusters illuminate the origin and evolution of a potentially chimeric lifestyle
Photosynthesis is thought to be restricted to a few bacterial and eukaryotic phyla. Here, Li et al. provide evidence of photosynthetic abilities in uncultivated bacteria within the phylum Myxococcota, suggesting that some of these organisms may combine predatory and photosynthetic abilities.
- Liuyang Li
- , Danyue Huang
- & Yinzhao Wang
-
Article
| Open Access MetaCC allows scalable and integrative analyses of both long-read and short-read metagenomic Hi-C data
MetaCC allows scalable and integrative analyses of both long-read and short-read metagenomic Hi-C data
The authors develop an integrative and scalable framework to eliminate systematic biases and retrieve high-quality metagenome-assembled genomes using either long-read or short-read metagenomic Hi-C data.
- Yuxuan Du
- & Fengzhu Sun
-
Article
| Open Access Social and psychological adversity are associated with distinct mother and infant gut microbiome variations
Social and psychological adversity are associated with distinct mother and infant gut microbiome variations
Here, in a cohort of mother-child dyads, the authors show that maternal prenatal social disadvantage and psychosocial stressors associate with distinct gut microbiome taxonomic and functional diversity in both the mothers and their four-month children.
- Barbara B. Warner
- , Bruce A. Rosa
- & Makedonka Mitreva
-
Article
| Open Access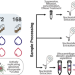 A subset of viruses thrives following microbial resuscitation during rewetting of a seasonally dry California grassland soil
A subset of viruses thrives following microbial resuscitation during rewetting of a seasonally dry California grassland soil
Rewetting of seasonally dry soils induces dramatic shifts in viral biomass and diversity. Combining stable isotope probing, metagenomics, and viromics Nicolas et al. provide evidence that viral lysis contributes to microbial turnover and the associated CO2 efflux.
- Alexa M. Nicolas
- , Ella T. Sieradzki
- & Steven J. Blazewicz
-
Article
| Open Access Ecophysiology and interactions of a taurine-respiring bacterium in the mouse gut
Ecophysiology and interactions of a taurine-respiring bacterium in the mouse gut
Authors utilise a multi-omics approach for the ecophysiological characterization of a taurine-respiring mouse gut bacterium.
- Huimin Ye
- , Sabrina Borusak
- & Alexander Loy
-
Article
| Open Access Phage-microbe dynamics after sterile faecal filtrate transplantation in individuals with metabolic syndrome: a double-blind, randomised, placebo-controlled clinical trial assessing efficacy and safety
Phage-microbe dynamics after sterile faecal filtrate transplantation in individuals with metabolic syndrome: a double-blind, randomised, placebo-controlled clinical trial assessing efficacy and safety
Bacteriophages (phages) can modify the gut microbiome to benefit human health. Here, the authors report the results of a double blind, randomized, placebo-controlled trial, showing that faecal filtrate transplantation (FFT), containing phages from lean healthy donors, is safe and improves glycemic variability in patients with metabolic syndrome, while shifting the gut phage composition.
- Koen Wortelboer
- , Patrick A. de Jonge
- & Hilde Herrema
-
Article
| Open Access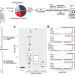 Ancient Clostridium DNA and variants of tetanus neurotoxins associated with human archaeological remains
Ancient Clostridium DNA and variants of tetanus neurotoxins associated with human archaeological remains
The analysis of microbial genomes from human archaeological samples offers a snapshot of ancient pathogens. Here, Hodgins et al. analyze metagenomic datasets from 38 human archaeological samples and identify bacterial genomic sequences related to modern-day Clostridium tetani, encoding tetanus neurotoxins.
- Harold P. Hodgins
- , Pengsheng Chen
- & Andrew C. Doxey
-
Article
| Open Access Genome-resolved correlation mapping links microbial community structure to metabolic interactions driving methane production from wastewater
Genome-resolved correlation mapping links microbial community structure to metabolic interactions driving methane production from wastewater
Anaerobic digestion of municipal mixed sludge is a microbial-mediated process that produces renewable natural gases such as methane. Here, Kieft et al. present the results of a two-year study of microbial community structure and function at a wastewater treatment plant, shedding light on metabolic interactions between microorganisms in relation with methane production.
- Brandon Kieft
- , Niko Finke
- & Steven J. Hallam
-
Article
| Open Access Removal of false positives in metagenomics-based taxonomy profiling via targeting Type IIB restriction sites
Removal of false positives in metagenomics-based taxonomy profiling via targeting Type IIB restriction sites
Here, leveraging species-specific Type IIB restriction endonuclease digestion sites as reference instead of universal markers or whole microbial genomes, the authors introduce MAP2B, a metagenomic profiler, showing it can significantly remove false-positive identification and generate highly accurate taxonomic profiling results.
- Zheng Sun
- , Jiang Liu
- & Yang-Yu Liu
-
Article
| Open Access Interrogating the viral dark matter of the rumen ecosystem with a global virome database
Interrogating the viral dark matter of the rumen ecosystem with a global virome database
Here, by mining 975 published rumen metagenomes for viral sequences, the authors construct a global rumen virome database (RVD), providing a resource for characterization of viral diversity, virus-host linkages, and potential roles in affecting rumen functions.
- Ming Yan
- , Akbar Adjie Pratama
- & Zhongtang Yu

 Exploring high-quality microbial genomes by assembling short-reads with long-range connectivity
Exploring high-quality microbial genomes by assembling short-reads with long-range connectivity