Article
|
Open Access
Featured
-
-
Article
| Open Access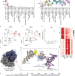 RPL3L-containing ribosomes determine translation elongation dynamics required for cardiac function
RPL3L-containing ribosomes determine translation elongation dynamics required for cardiac function
RPL3L is a paralog of the ribosomal protein RPL3 and specifically expressed in heart and skeletal muscle. Here, the authors show that RPL3L-containing ribosomes regulate translation elongation dynamics especially for the transcripts related to cardiac muscle contraction.
- Chisa Shiraishi
- , Akinobu Matsumoto
- & Keiichi I. Nakayama
-
Article
| Open Access Copy number variation in tRNA isodecoder genes impairs mammalian development and balanced translation
Copy number variation in tRNA isodecoder genes impairs mammalian development and balanced translation
Enigmatically tRNA genes exist in several hundred copies in mammalian genomes. Here the authors find a precipitous failure of development and increased amino acid misincorporation upon systematic elimination of tRNA-Phe genes in mice using CRISPR.
- Laetitia A. Hughes
- , Danielle L. Rudler
- & Aleksandra Filipovska
-
Article
| Open Access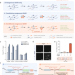 Autocatalytic base editing for RNA-responsive translational control
Autocatalytic base editing for RNA-responsive translational control
Genetic circuits that control transgene expression in response to pre-defined transcriptional cues would enable the development of smart therapeutics. Here the authors engineer programmable RNA sensors, DART VADARs, in which ADARs autocatalytically convert target hybridization into a translational output, thus amplifying editing by endogenous ADAR via positive feedback and conferring high dynamic range and a small genetic footprint.
- Raphaël V. Gayet
- , Katherine Ilia
- & James J. Collins
-
Article
| Open Access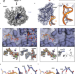 rRNA methylation by Spb1 regulates the GTPase activity of Nog2 during 60S ribosomal subunit assembly
rRNA methylation by Spb1 regulates the GTPase activity of Nog2 during 60S ribosomal subunit assembly
Regulation of 60S biogenesis remains poorly understood. Using cryo-EM, the authors show that failure of Spb1 to methylate the A-loop nucleotide G2922 prematurely activates the GTPase Nog2, suggesting that Spb1 and Nog2 form a kinetic checkpoint during ribosome maturation.
- Kamil Sekulski
- , Victor Emmanuel Cruz
- & Jan P. Erzberger
-
Article
| Open Access The translating bacterial ribosome at 1.55 Å resolution generated by cryo-EM imaging services
The translating bacterial ribosome at 1.55 Å resolution generated by cryo-EM imaging services
Developments in cryo-EM sample preparation and data collection are pivotal for structure determination. Fromm et al. present a 1.55 Å structure of the translating bacterial ribosome that provides new insights on its function and may allow for more precise structure-based drug design.
- Simon A. Fromm
- , Kate M. O’Connor
- & Simone Mattei
-
Article
| Open Access Structural basis for clearing of ribosome collisions by the RQT complex
Structural basis for clearing of ribosome collisions by the RQT complex
Ribosome collisions serve as proxy for aberrant translation to initiate rescue and quality control pathways. Here, authors elucidate the molecular mechanism of collided ribosome clearance by the ribosome quality control trigger complex.
- Katharina Best
- , Ken Ikeuchi
- & Roland Beckmann
-
Article
| Open Access Antibiotic thermorubin tethers ribosomal subunits and impedes A-site interactions to perturb protein synthesis in bacteria
Antibiotic thermorubin tethers ribosomal subunits and impedes A-site interactions to perturb protein synthesis in bacteria
Thermorubin is a ribosome-targeting antibiotic. Here, using fast-kinetics and cryoEM, the authors reveal that thermorubin primarily blocks ribosome-recycling by tethering the ribosomal subunits besides impeding translation elongation and termination steps.
- Narayan Prasad Parajuli
- , Andrew Emmerich
- & Suparna Sanyal
-
Article
| Open Access Reanalysis of ribosome profiling datasets reveals a function of rocaglamide A in perturbing the dynamics of translation elongation via eIF4A
Reanalysis of ribosome profiling datasets reveals a function of rocaglamide A in perturbing the dynamics of translation elongation via eIF4A
The compound Rocaglamide A (RocA) is known for repressing translation initiation. Here the authors identify a dual mode of action for RocA in blocking translation initiation and elongation via eIF4A using previous datasets and new analyses.
- Fajin Li
- , Jianhuo Fang
- & Xuerui Yang
-
Article
| Open Access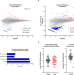 CPEB1-dependent disruption of the mRNA translation program in oocytes during maternal aging
CPEB1-dependent disruption of the mRNA translation program in oocytes during maternal aging
In the absence of transcription, gene expression required for oocyte maturation is regulated by translation of preexisting mRNAs. Here the authors show that a defective program of mRNA translation in oocyte is associated with maternal aging.
- Nozomi Takahashi
- , Federica Franciosi
- & Marco Conti
-
Article
| Open Access RSL24D1 sustains steady-state ribosome biogenesis and pluripotency translational programs in embryonic stem cells
RSL24D1 sustains steady-state ribosome biogenesis and pluripotency translational programs in embryonic stem cells
Pluripotency is coordinated at multiple levels of gene expression. Here the authors show that ribosome biogenesis is tightly regulated in embryonic stem cells (ESC) to control the translation of transcription and chromatin factors and dictate ESC fate.
- Sébastien Durand
- , Marion Bruelle
- & Mathieu Gabut
-
Article
| Open Access Tuning phenylalanine fluorination to assess aromatic contributions to protein function and stability in cells
Tuning phenylalanine fluorination to assess aromatic contributions to protein function and stability in cells
Aromatic amino acids in proteins support ligand binding and protein stability. To parse the physiocochemical roles of aromatic interactions, here Galles, Infield and co-authors identify pyrrolysine-based aminoacyl-tRNA synthetases that enable the encoding of fluorinated phenylalanine amino acids.
- Grace D. Galles
- , Daniel T. Infield
- & Christopher A. Ahern
-
Article
| Open Access Human mitochondria require mtRF1 for translation termination at non-canonical stop codons
Human mitochondria require mtRF1 for translation termination at non-canonical stop codons
The mammalian mitochondrial genome contains two noncanonical stop codons, AGA and AGG. Here, the authors show that mtRF1, a mitochondrial protein of unknown function, triggers translation termination at AGA or AGG, placing mtRF1 among mitochondrial translation factors.
- Annika Krüger
- , Cristina Remes
- & Joanna Rorbach
-
Article
| Open Access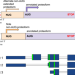 Thousands of human non-AUG extended proteoforms lack evidence of evolutionary selection among mammals
Thousands of human non-AUG extended proteoforms lack evidence of evolutionary selection among mammals
Analysis of a large number of Ribo-seq datasets and genomic alignments led to detection of novel non-AUG proteoforms. Unexpectedly the number of non-AUG proteoforms identified with Ribo-seq greatly exceeds those with strong phylogenetic support.
- Alla D. Fedorova
- , Stephen J. Kiniry
- & Pavel V. Baranov
-
Article
| Open Access DAP5 enables main ORF translation on mRNAs with structured and uORF-containing 5′ leaders
DAP5 enables main ORF translation on mRNAs with structured and uORF-containing 5′ leaders
RNA structure and upstream ORFs can modulate the translation of human transcripts. Here, the authors show that structured mRNAs with pervasive uORF translation require DAP5, an eIF4G-like protein, for translation of the main ORF.
- Ramona Weber
- , Leon Kleemann
- & Cátia Igreja
-
Article
| Open Access Structures of the eukaryotic ribosome and its translational states in situ
Structures of the eukaryotic ribosome and its translational states in situ
The translational states of eukaryotic ribosomes have so far been only investigated in vitro. Here, authors obtained the 3.8 Å in situ 80S ribosome structure, the distribution of translational states and unique arrangement of rRNA expansion segments.
- Patrick C. Hoffmann
- , Jan Philipp Kreysing
- & Martin Beck
-
Article
| Open Access Nascent peptide-induced translation discontinuation in eukaryotes impacts biased amino acid usage in proteomes
Nascent peptide-induced translation discontinuation in eukaryotes impacts biased amino acid usage in proteomes
Here the authors find that nascent chains enriched in acidic amino acids destabilize the translating ribosome, eventually leading to stochastic premature termination in eukaryotes as in bacteria. Such risk of premature termination influences the amino acid distribution in the proteomes.
- Yosuke Ito
- , Yuhei Chadani
- & Hideki Taguchi
-
Article
| Open Access Varying strength of selection contributes to the intragenomic diversity of rRNA genes
Varying strength of selection contributes to the intragenomic diversity of rRNA genes
Ribosomal RNA genes are abundant in eukaryotic genomes and code for the universal and essential RNA components of the ribosome. This study uncovers high sequence diversity of the genes within a single species and discusses the contribution of selection in the evolution of ribosomal RNA.
- Daniel Sultanov
- & Andreas Hochwagen
-
Article
| Open Access A nascent peptide code for translational control of mRNA stability in human cells
A nascent peptide code for translational control of mRNA stability in human cells
Here, the authors measure the mRNA levels of thousands of coding sequence motifs in human cells to find that nascent peptides with a combination of β-strand structures and bulky and positively charged sequences reduce mRNA levels by slowing translation.
- Phillip C. Burke
- , Heungwon Park
- & Arvind Rasi Subramaniam
-
Article
| Open Access Structural dynamics of AAA + ATPase Drg1 and mechanism of benzo-diazaborine inhibition
Structural dynamics of AAA + ATPase Drg1 and mechanism of benzo-diazaborine inhibition
Drg1 is a cytoplasmic AAA + ATPase responsible for releasing Rlp24 during ribosome assembly. Here, the authors show that Drg1 structurally and functionally shares many regulatory features with that of Cdc48/p97, suggesting a unfoldase role for Drg1.
- Chengying Ma
- , Damu Wu
- & Ning Gao
-
Article
| Open Access Regulation of BRCA1 stability through the tandem UBX domains of isoleucyl-tRNA synthetase 1
Regulation of BRCA1 stability through the tandem UBX domains of isoleucyl-tRNA synthetase 1
Aminoacyl-tRNA synthetases possess unique domains. In this study the structure of the vertebrate IARS1 and EARS1 complex reveals that vertebrate IARS1 protects the DNA repair factor BRCA1 from proteolytic degradation via its UBX-fold domain.
- Scisung Chung
- , Mi-Sun Kang
- & Yunje Cho
-
Article
| Open Access Translation regulatory factor BZW1 regulates preimplantation embryo development and compaction by restricting global non-AUG Initiation
Translation regulatory factor BZW1 regulates preimplantation embryo development and compaction by restricting global non-AUG Initiation
It is unclear how translational efficiency and codon stringency affect zygotic genome activation and early embryonic development. Here they show that the zygotic gene BZW1 increases start codon stringency, which is important for preimplantation development.
- Jue Zhang
- , Shuai-Bo Pi
- & Heng-Yu Fan
-
Article
| Open Access Tumor suppressor mediated ubiquitylation of hnRNPK is a barrier to oncogenic translation
Tumor suppressor mediated ubiquitylation of hnRNPK is a barrier to oncogenic translation
Cytoplasmic accumulation of RNA binding protein, hnRNPK in cancer cells is associated with poor prognosis. Here the authors show that SCFFbxo4 E3 ubiquitin ligase-mediated polyubiquitylation of hnRNPK restricts c-Myc translation and limits cancer progression.
- Bartosz Mucha
- , Shuo Qie
- & J. Alan Diehl
-
Article
| Open Access Cap-dependent translation initiation monitored in living cells
Cap-dependent translation initiation monitored in living cells
mRNA translation is tightly regulated to preserve cellular homeostasis. Here live-cell spectroscopy and single-particle tracking are combined to interrogate the binding dynamics of endogenous initiation factors to mRNAs 5’cap, revealing the sequence of translation initiation factors assembly and disassembly; and the clustering of translation in neurons.
- Valentina Gandin
- , Brian P. English
- & Robert H. Singer
-
Article
| Open Access A distinct mammalian disome collision interface harbors K63-linked polyubiquitination of uS10 to trigger hRQT-mediated subunit dissociation
A distinct mammalian disome collision interface harbors K63-linked polyubiquitination of uS10 to trigger hRQT-mediated subunit dissociation
Collided ribosomes are marked by ubiquitination to induce quality control mechanisms. Here, authors show that mammalian disomes form a distinct structural interface, in which uS10 K63-linked polyubiquitination is critical for ribosome dissociation.
- Momoko Narita
- , Timo Denk
- & Toshifumi Inada
-
Article
| Open Access Human mtRF1 terminates COX1 translation and its ablation induces mitochondrial ribosome-associated quality control
Human mtRF1 terminates COX1 translation and its ablation induces mitochondrial ribosome-associated quality control
How translation termination is achieved for the non-conventional mtDNA-encoded COX1 and ND6 was so far unknown. Here, Nadler et al. address this question by assessing the functions and specificity of the mitochondrial release factors mtRF1 and mtRF1a.
- Franziska Nadler
- , Elena Lavdovskaia
- & Ricarda Richter-Dennerlein
-
Article
| Open Access Translational reprogramming in response to accumulating stressors ensures critical threshold levels of Hsp90 for mammalian life
Translational reprogramming in response to accumulating stressors ensures critical threshold levels of Hsp90 for mammalian life
The molecular chaperone Hsp90 decreases with age but whether organisms can physiologically tune expression is unclear. Here, the authors show that mammalian cells can adjust Hsp90 levels to accumulating cellular stress by translational reprogramming.
- Kaushik Bhattacharya
- , Samarpan Maiti
- & Didier Picard
-
Article
| Open Access Structure of a mitochondrial ribosome with fragmented rRNA in complex with membrane-targeting elements
Structure of a mitochondrial ribosome with fragmented rRNA in complex with membrane-targeting elements
Structural and evolutionary analysis of a diverged mitoribosome with rRNA that is split in 13 pieces reveals a non-canonical reduced form of the 5 S, permutation of domain I, and how the fragmentation pattern links to the concept of a primordial ribosome.
- Victor Tobiasson
- , Ieva Berzina
- & Alexey Amunts
-
Article
| Open Access A stem cell roadmap of ribosome heterogeneity reveals a function for RPL10A in mesoderm production
A stem cell roadmap of ribosome heterogeneity reveals a function for RPL10A in mesoderm production
How ribosomes differ in composition and function to regulate gene expression is poorly understood. Here, the authors show that ribosome composition changes during stem cell differentiation and identify a ribosomal protein that regulates production of the mesoderm lineage.
- Naomi R. Genuth
- , Zhen Shi
- & Maria Barna
-
Article
| Open Access Single-cell transcriptome and translatome dual-omics reveals potential mechanisms of human oocyte maturation
Single-cell transcriptome and translatome dual-omics reveals potential mechanisms of human oocyte maturation
Development of methods for simultaneous single cell analysis of transcription and translation is still underway. Here, Hu et al. develop single-cell transcriptome and translatome dual-omics on human oocytes, which enables them to identify OOSP2 as an induction factor during human oocyte maturation.
- Wenqi Hu
- , Haitao Zeng
- & Kehkooi Kee
-
Article
| Open Access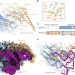 Structural basis for shape-selective recognition and aminoacylation of a D-armless human mitochondrial tRNA
Structural basis for shape-selective recognition and aminoacylation of a D-armless human mitochondrial tRNA
Mitochondrial tRNAs are indispensable and yet underwent an extreme mutational erosion. The authors report the structures of a mitochondrial aaRS-tRNA complex and show how the most degenerated of all human mtRNAs is recognized by its cognate synthetase to maintain mitochondrial gene expression.
- Bernhard Kuhle
- , Marscha Hirschi
- & Paul Schimmel
-
Article
| Open Access Oligodendrocyte differentiation alters tRNA modifications and codon optimality-mediated mRNA decay
Oligodendrocyte differentiation alters tRNA modifications and codon optimality-mediated mRNA decay
Mutations in tRNA processing factors can lead to myelin disorders. This study shows that differentiated oligodendrocytes, cells that make the myelin, are characterized by different tRNA modifications and mRNA decay compared to their precursor cells.
- Sophie Martin
- , Kevin C. Allan
- & Jeff Coller
-
Article
| Open Access Duplicated ribosomal protein paralogs promote alternative translation and drug resistance
Duplicated ribosomal protein paralogs promote alternative translation and drug resistance
Most yeast ribosomal protein genes are duplicated but the functional significance of this duplication remains unclear. This study identifies a natural program where changing the ratio of proteins produced from duplicated genes modifies translation in response to drugs regardless of ribosome number.
- Mustafa Malik Ghulam
- , Mathieu Catala
- & Sherif Abou Elela
-
Article
| Open Access CHIKV infection reprograms codon optimality to favor viral RNA translation by altering the tRNA epitranscriptome
CHIKV infection reprograms codon optimality to favor viral RNA translation by altering the tRNA epitranscriptome
Viruses completely depend on the host translational machinery, but their genomes are often enriched in rare codons and therefore should be translated with poor efficiency. Here, Jungfleisch et al. apply Ribo-Seq and RNASeq to provide a global view on the translational changes occurring during Chikungunya virus (CHIKV) infection. CHIKV infection induces a codon-specific reprogramming of the host translation machinery to favor the translation of viral RNA genomes over host mRNAs via tRNA modification.
- Jennifer Jungfleisch
- , René Böttcher
- & Juana Díez
-
Article
| Open Access Ribosome impairment regulates intestinal stem cell identity via ZAKɑ activation
Ribosome impairment regulates intestinal stem cell identity via ZAKɑ activation
Intestinal stem cells are responsible for replenishing cells within the high-turnover intestinal epithelium. Here they show that ribosome dynamics affect intestinal stem cell identity through a mechanism that is triggered by changes in nutrient availability.
- Joana Silva
- , Ferhat Alkan
- & William James Faller
-
Article
| Open Access Modulating co-translational protein folding by rational design and ribosome engineering
Modulating co-translational protein folding by rational design and ribosome engineering
The narrow exit tunnel of the ribosome is important for cotranslational protein folding. Here, authors show that their rationally designed and engineered exit tunnel protein loops modulate the free energy of nascent chain dynamics and folding.
- Minkoo Ahn
- , Tomasz Włodarski
- & John Christodoulou
-
Article
| Open Access Altered tRNA dynamics during translocation on slippery mRNA as determinant of spontaneous ribosome frameshifting
Altered tRNA dynamics during translocation on slippery mRNA as determinant of spontaneous ribosome frameshifting
Slippery sequences in mRNA can cause the ribosome to change its reading frame. Using smFRET, Poulis et al. show how reversible fluctuations of peptidyl-tRNA slow down translocation, alter ribosome dynamics, and favor spontaneous ribosome frameshifting.
- Panagiotis Poulis
- , Anoshi Patel
- & Sarah Adio
-
Article
| Open Access Pervasive translation of circular RNAs driven by short IRES-like elements
Pervasive translation of circular RNAs driven by short IRES-like elements
Unbiased screen of random sequences identified many short IRES-like elements to drive circular RNA translation and hundreds of rolling circle translation events, suggesting a pervasive cap-independent translation in human transcriptome.
- Xiaojuan Fan
- , Yun Yang
- & Zefeng Wang
-
Article
| Open Access Structural remodeling of ribosome associated Hsp40-Hsp70 chaperones during co-translational folding
Structural remodeling of ribosome associated Hsp40-Hsp70 chaperones during co-translational folding
Ribosome associated complex (RAC)- HSP70 (Ssb in yeast) is a eukaryotic chaperone system involved in co-translational folding. Here, authors report structures of RAC-containing ribosomal complexes, which suggest a working model for the dynamic actions of RAC-Ssb during the process.
- Yan Chen
- , Bin Tsai
- & Ning Gao
-
Article
| Open Access Compact IF2 allows initiator tRNA accommodation into the P site and gates the ribosome to elongation
Compact IF2 allows initiator tRNA accommodation into the P site and gates the ribosome to elongation
Initiation factor 2 (IF2) guides the ribosome to the elongation phase of protein synthesis. Here, Basu et al. provide structural insights into how compact IF2-GDP makes way for initiator tRNA accommodation into the peptidyl (P) site of the ribosome.
- Ritwika S. Basu
- , Michael B. Sherman
- & Matthieu G. Gagnon
-
Article
| Open Access Codon-specific Ramachandran plots show amino acid backbone conformation depends on identity of the translated codon
Codon-specific Ramachandran plots show amino acid backbone conformation depends on identity of the translated codon
Genetic code redundancies are considered inconsequential to protein structure. This study uncovers a dependence between local amino acid conformation in folded proteins and the identity of the codon from which that amino acid was translated.
- Aviv A. Rosenberg
- , Ailie Marx
- & Alex M. Bronstein
-
Article
| Open Access Ribosome inhibition by C9ORF72-ALS/FTD-associated poly-PR and poly-GR proteins revealed by cryo-EM
Ribosome inhibition by C9ORF72-ALS/FTD-associated poly-PR and poly-GR proteins revealed by cryo-EM
The expansion of GGGGCC repeats in the C9ORF72 gene results in the production of disease causing abnormal proteins with polymeric glycine-arginine (poly-GR) and polymeric proline-arginine (poly-PR). Here the authors demonstrate a structural mechanism of how poly-GR and poly-PR inhibit translation and how they might also perturb ribosome assembly.
- Anna B. Loveland
- , Egor Svidritskiy
- & Andrei A. Korostelev
-
Article
| Open Access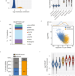 HDLBP binds ER-targeted mRNAs by multivalent interactions to promote protein synthesis of transmembrane and secreted proteins
HDLBP binds ER-targeted mRNAs by multivalent interactions to promote protein synthesis of transmembrane and secreted proteins
RNA binding protein HDLBP (or Vigilin) localizes in the endoplasmic reticulum (ER) membrane. Here the authors show that HDLBP contributes to translation of ER-targeted mRNAs.
- Ulrike Zinnall
- , Miha Milek
- & Markus Landthaler
-
Article
| Open Access Substrate recognition and cryo-EM structure of the ribosome-bound TAC toxin of Mycobacterium tuberculosis
Substrate recognition and cryo-EM structure of the ribosome-bound TAC toxin of Mycobacterium tuberculosis
Toxin-antitoxin systems are widespread in bacteria. Here the authors present structures of M. tuberculosis HigBTAC alone and bound to the ribosome in the presence of native cspA mRNA, shedding light on its mechanism of translation inhibition.
- Moise Mansour
- , Emmanuel Giudice
- & Pierre Genevaux
-
Article
| Open Access Ataluren binds to multiple protein synthesis apparatus sites and competitively inhibits release factor-dependent termination
Ataluren binds to multiple protein synthesis apparatus sites and competitively inhibits release factor-dependent termination
Ataluren is the only nonsense suppressor drug currently approved for clinical use. Here, the authors determine where ataluren binds to the ribosome and how it inhibits termination at nonsense codons.
- Shijie Huang
- , Arpan Bhattacharya
- & Barry S. Cooperman
-
Article
| Open Access Regulation of A-to-I RNA editing and stop codon recoding to control selenoprotein expression during skeletal myogenesis
Regulation of A-to-I RNA editing and stop codon recoding to control selenoprotein expression during skeletal myogenesis
Stop codon recoding is a unique mechanism for selenoprotein biogenesis. Here, the authors report regulatory systems for selenoprotein expression during skeletal muscle differentiation by regulating A-to-I RNA editing and stop codon recoding.
- Yuta Noda
- , Shunpei Okada
- & Tsutomu Suzuki
-
Article
| Open Access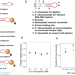 A multiplex platform for small RNA sequencing elucidates multifaceted tRNA stress response and translational regulation
A multiplex platform for small RNA sequencing elucidates multifaceted tRNA stress response and translational regulation
This paper develops a multiplex RNA-seq method that reports tRNA abundance, modification, charging, and fragmentation. Results show stress-induced regulation in translational elongation and association of modification and tRNA fragment biogenesis.
- Christopher P. Watkins
- , Wen Zhang
- & Tao Pan
-
Article
| Open Access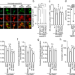 Pantothenate kinase 2 interacts with PINK1 to regulate mitochondrial quality control via acetyl-CoA metabolism
Pantothenate kinase 2 interacts with PINK1 to regulate mitochondrial quality control via acetyl-CoA metabolism
PKAN and PD are two distinct diseases with overlapping pathophysiology. Here, authors show that their pathogenic genes PANK2 and PINK1 interact. PANK2 regulates mitophagy via CoA metabolism, while PINK1 supervises PANK2 translation on mitochondria.
- Yunpeng Huang
- , Zhihui Wan
- & Bing Zhou
-
Article
| Open Access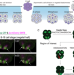 Vasa nucleates asymmetric translation along the mitotic spindle during unequal cell divisions
Vasa nucleates asymmetric translation along the mitotic spindle during unequal cell divisions
Association of mRNA translation with the mitotic spindle is thought to be involved in localized production of cell fate determinants. Here, the authors show Vasa facilitates asymmetric translation, which contributes to differential regulation during sea urchin embryogenesis.
- Ana Fernandez-Nicolas
- , Alicia Uchida
- & Mamiko Yajima
-
Article
| Open Access Structural basis for PoxtA-mediated resistance to phenicol and oxazolidinone antibiotics
Structural basis for PoxtA-mediated resistance to phenicol and oxazolidinone antibiotics
PoxtA confers resistance to ribosome-targeting oxazolidinone (linezolid) and chloramphenicol antibiotics. Here, Crowe-McAuliffe et al. provide structural insights into how binding of PoxtA to the ribosome indirectly promotes drug dissociation.
- Caillan Crowe-McAuliffe
- , Victoriia Murina
- & Vasili Hauryliuk

 Translational reprogramming as a driver of antimony-drug resistance in Leishmania
Translational reprogramming as a driver of antimony-drug resistance in Leishmania