Featured
-
-
Article
| Open Access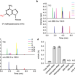 C2-methyladenosine in tRNA promotes protein translation by facilitating the decoding of tandem m2A-tRNA-dependent codons
C2-methyladenosine in tRNA promotes protein translation by facilitating the decoding of tandem m2A-tRNA-dependent codons
Duan et al. demonstrate that the m2A modification is ubiquitous in plants and tRNA m2A37 promotes a relaxed conformation of tRNA, enhancing translation efficiency by facilitating decoding of tandem m2A-tRNA-dependent codons.
- Hong-Chao Duan
- , Chi Zhang
- & Guifang Jia
-
Article
| Open Access Antibiotic hyper-resistance in a class I aminoacyl-tRNA synthetase with altered active site signature motif
Antibiotic hyper-resistance in a class I aminoacyl-tRNA synthetase with altered active site signature motif
Aminoacyl-tRNA synthetases translate the genetic code. These enzymes harbor signature catalytic motifs dating from their ancient ancestors. A natural variation of one of the stated motifs was discovered and linked to antibiotic hyper-resistance.
- A. Brkic
- , M. Leibundgut
- & I. Gruic-Sovulj
-
Article
| Open Access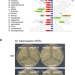 MenT nucleotidyltransferase toxins extend tRNA acceptor stems and can be inhibited by asymmetrical antitoxin binding
MenT nucleotidyltransferase toxins extend tRNA acceptor stems and can be inhibited by asymmetrical antitoxin binding
Bacteria can control their growth using internal toxins regulated by antitoxins. Here, the authors show MenT toxins from Mycobacterium tuberculosis work by modifying tRNAs, and asymmetric antitoxin binding blocks toxin activity.
- Xibing Xu
- , Ben Usher
- & Pierre Genevaux
-
Article
| Open Access Extensive breaking of genetic code degeneracy with non-canonical amino acids
Extensive breaking of genetic code degeneracy with non-canonical amino acids
Genetic code expansion is limited by the degeneracy of the 61 sense codons which encode for only 20 amino acids. Here, the authors show that by combining hyperaccurate ribosomes and in vitro transcribed tRNAs, dramatic and extensive breaking of sense codon degeneracy can be achieved.
- Clinton A. L. McFeely
- , Bipasana Shakya
- & Matthew C. T. Hartman
-
Article
| Open Access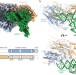 Structural basis for a degenerate tRNA identity code and the evolution of bimodal specificity in human mitochondrial tRNA recognition
Structural basis for a degenerate tRNA identity code and the evolution of bimodal specificity in human mitochondrial tRNA recognition
Aminoacyl-tRNA synthetases catalyze the ligation of amino acids to their cognate tRNAs. Here the authors report the cryo-EM structure of a human mitochondrial seryl-tRNA synthetase•mtRNASer complex showing how strong mutation pressure on mtRNA genes drove a rewiring of intermolecular recognition rules.
- Bernhard Kuhle
- , Marscha Hirschi
- & Paul Schimmel
-
Article
| Open Access Quality control of protein synthesis in the early elongation stage
Quality control of protein synthesis in the early elongation stage
Peptidyl-tRNAs (pep-tRNAs) frequently dissociate from ribosome, called as pep-tRNA drop-off. But, its function remained unclear. The authors proposed a new quality control mechanism of protein synthesis by active rejection of miscoded pep-tRNAs in the early stage of translation.
- Asuteka Nagao
- , Yui Nakanishi
- & Tsutomu Suzuki
-
Article
| Open Access Copy number variation in tRNA isodecoder genes impairs mammalian development and balanced translation
Copy number variation in tRNA isodecoder genes impairs mammalian development and balanced translation
Enigmatically tRNA genes exist in several hundred copies in mammalian genomes. Here the authors find a precipitous failure of development and increased amino acid misincorporation upon systematic elimination of tRNA-Phe genes in mice using CRISPR.
- Laetitia A. Hughes
- , Danielle L. Rudler
- & Aleksandra Filipovska
-
Article
| Open Access Regulation of BRCA1 stability through the tandem UBX domains of isoleucyl-tRNA synthetase 1
Regulation of BRCA1 stability through the tandem UBX domains of isoleucyl-tRNA synthetase 1
Aminoacyl-tRNA synthetases possess unique domains. In this study the structure of the vertebrate IARS1 and EARS1 complex reveals that vertebrate IARS1 protects the DNA repair factor BRCA1 from proteolytic degradation via its UBX-fold domain.
- Scisung Chung
- , Mi-Sun Kang
- & Yunje Cho
-
Article
| Open Access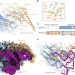 Structural basis for shape-selective recognition and aminoacylation of a D-armless human mitochondrial tRNA
Structural basis for shape-selective recognition and aminoacylation of a D-armless human mitochondrial tRNA
Mitochondrial tRNAs are indispensable and yet underwent an extreme mutational erosion. The authors report the structures of a mitochondrial aaRS-tRNA complex and show how the most degenerated of all human mtRNAs is recognized by its cognate synthetase to maintain mitochondrial gene expression.
- Bernhard Kuhle
- , Marscha Hirschi
- & Paul Schimmel
-
Article
| Open Access Oligodendrocyte differentiation alters tRNA modifications and codon optimality-mediated mRNA decay
Oligodendrocyte differentiation alters tRNA modifications and codon optimality-mediated mRNA decay
Mutations in tRNA processing factors can lead to myelin disorders. This study shows that differentiated oligodendrocytes, cells that make the myelin, are characterized by different tRNA modifications and mRNA decay compared to their precursor cells.
- Sophie Martin
- , Kevin C. Allan
- & Jeff Coller
-
Article
| Open Access CHIKV infection reprograms codon optimality to favor viral RNA translation by altering the tRNA epitranscriptome
CHIKV infection reprograms codon optimality to favor viral RNA translation by altering the tRNA epitranscriptome
Viruses completely depend on the host translational machinery, but their genomes are often enriched in rare codons and therefore should be translated with poor efficiency. Here, Jungfleisch et al. apply Ribo-Seq and RNASeq to provide a global view on the translational changes occurring during Chikungunya virus (CHIKV) infection. CHIKV infection induces a codon-specific reprogramming of the host translation machinery to favor the translation of viral RNA genomes over host mRNAs via tRNA modification.
- Jennifer Jungfleisch
- , René Böttcher
- & Juana Díez
-
Article
| Open Access Compact IF2 allows initiator tRNA accommodation into the P site and gates the ribosome to elongation
Compact IF2 allows initiator tRNA accommodation into the P site and gates the ribosome to elongation
Initiation factor 2 (IF2) guides the ribosome to the elongation phase of protein synthesis. Here, Basu et al. provide structural insights into how compact IF2-GDP makes way for initiator tRNA accommodation into the peptidyl (P) site of the ribosome.
- Ritwika S. Basu
- , Michael B. Sherman
- & Matthieu G. Gagnon
-
Article
| Open Access Regulation of A-to-I RNA editing and stop codon recoding to control selenoprotein expression during skeletal myogenesis
Regulation of A-to-I RNA editing and stop codon recoding to control selenoprotein expression during skeletal myogenesis
Stop codon recoding is a unique mechanism for selenoprotein biogenesis. Here, the authors report regulatory systems for selenoprotein expression during skeletal muscle differentiation by regulating A-to-I RNA editing and stop codon recoding.
- Yuta Noda
- , Shunpei Okada
- & Tsutomu Suzuki
-
Article
| Open Access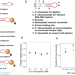 A multiplex platform for small RNA sequencing elucidates multifaceted tRNA stress response and translational regulation
A multiplex platform for small RNA sequencing elucidates multifaceted tRNA stress response and translational regulation
This paper develops a multiplex RNA-seq method that reports tRNA abundance, modification, charging, and fragmentation. Results show stress-induced regulation in translational elongation and association of modification and tRNA fragment biogenesis.
- Christopher P. Watkins
- , Wen Zhang
- & Tao Pan
-
Article
| Open Access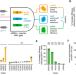 Multiplex suppression of four quadruplet codons via tRNA directed evolution
Multiplex suppression of four quadruplet codons via tRNA directed evolution
Genetic code expansion strategies are limited to specific codons that can be reassigned to new amino acids. Here the authors show that quadruplet-decoding tRNAs (qtRNAs) can be rapidly discovered and evolved to decode new quadruplet codons, enabling four independent decoding events in a single protein in living cells.
- Erika A. DeBenedictis
- , Gavriela D. Carver
- & Ahmed H. Badran
-
Article
| Open Access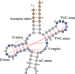 Repurposing tRNAs for nonsense suppression
Repurposing tRNAs for nonsense suppression
Here, the authors report de novo design, optimization and characterization of tRNAs that decode UGA stop codons in E. coli. The structure of the ribosome in a complex with the designed tRNA bound to a UGA stop codon suggests that distinct A-site ligands (tRNAs versus release factors) induce distinct conformation of the stop codon within the mRNA in the decoding center.
- Suki Albers
- , Bertrand Beckert
- & Zoya Ignatova
-
Article
| Open Access The genomic loci of specific human tRNA genes exhibit ageing-related DNA hypermethylation
The genomic loci of specific human tRNA genes exhibit ageing-related DNA hypermethylation
The epigenome has been shown to change with age, potentially impacting on ageing-related disease. Here the authors investigate the DNA methylation state of the genomic loci of human tRNA and observe enrichment for age-related DNA hypermethylation at tRNA loci.
- Richard J. Acton
- , Wei Yuan
- & Christopher G. Bell
-
Article
| Open Access Wobble tRNA modification and hydrophilic amino acid patterns dictate protein fate
Wobble tRNA modification and hydrophilic amino acid patterns dictate protein fate
Wobble uridine (U34) tRNA modifications are important for the decoding of AA-ending codons. Here the authors show that while the U34-codon content of mRNAs are predictive of changes in ribosome translation elongation, the resulting outcome in protein expression also relies on specific hydrophilic motifs-dependent protein aggregation and clearance.
- Francesca Rapino
- , Zhaoli Zhou
- & Pierre Close
-
Article
| Open Access Inhibitory mechanism of reveromycin A at the tRNA binding site of a class I synthetase
Inhibitory mechanism of reveromycin A at the tRNA binding site of a class I synthetase
Reveromycin A (RM-A) selectively inhibits eukaryotic cytoplasmic isoleucyltRNA synthetase (IleRS). Herein, the authors show that RM-A molecule occupies the tRNAIle binding site of Saccharomyces cerevisiae IleRS, and that RM-A cooperates with isoleucine or isoleucyl-adenylate for IleRS binding.
- Bingyi Chen
- , Siting Luo
- & Huihao Zhou
-
Article
| Open Access A substrate binding model for the KEOPS tRNA modifying complex
A substrate binding model for the KEOPS tRNA modifying complex
KEOPS is an evolutionary conserved complex with a core of four core subunits—Pcc1, Kae1, Bud32 and Cgi121—that catalyzes the universal and essential tRNA modification N6-threonylcarbamoyl adenosine (t6A). Here the authors describe a Cgi121-tRNA crystal structure and new composite model of the KEOPS holo-enzyme-substrate complex that shed light on the t6A catalytic cycle and its regulation.
- Jonah Beenstock
- , Samara Mishelle Ona
- & Frank Sicheri
-
Article
| Open Access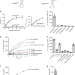 Mechanism of aminoacyl-tRNA acetylation by an aminoacyl-tRNA acetyltransferase AtaT from enterohemorrhagic E. coli
Mechanism of aminoacyl-tRNA acetylation by an aminoacyl-tRNA acetyltransferase AtaT from enterohemorrhagic E. coli
AtaT is a type-II toxin from enterohemorrhagic E. coli, reported to acetylate the aminoacyl-moiety of initiator Met-tRNAfMet, thus inhibiting translation initiation. Biochemical analysis suggests that AtaT has a broader specificity for aminoacyl-tRNAs and inhibits global translation. Structure of AtaT in complex with acetylated Met-tRNAfMet offers insight into the substrate selection by the enzyme.
- Yuka Yashiro
- , Yuriko Sakaguchi
- & Kozo Tomita
-
Article
| Open Access Structural basis for the transition from translation initiation to elongation by an 80S-eIF5B complex
Structural basis for the transition from translation initiation to elongation by an 80S-eIF5B complex
The GTPase eIF5B is involved in the correct positioning of the initiator Met-tRNAiMet on the ribosome during the late stages of translation initiation. Here the authors present a cryo-EM structure of the ribosome in complex with eIF5B and Met-tRNAiMet immediately before transition into elongation, providing insight on Met-tRNAiMetdelivery and how the correct reading frame is established.
- Jinfan Wang
- , Jing Wang
- & Israel S. Fernández
-
Article
| Open Access Quantitative tRNA-sequencing uncovers metazoan tissue-specific tRNA regulation
Quantitative tRNA-sequencing uncovers metazoan tissue-specific tRNA regulation
The relative abundance of specific tRNA can impact protein production rate, folding, and messenger RNA stability. Here the authors describe QuantM-tRNA seq — a method to monitor tRNA abundance and sequence variants — and uncover distinctions in isodecoder expression between tissues that are independent of the anticodon pool of each tRNA family.
- Otis Pinkard
- , Sean McFarland
- & Jeff Coller
-
Article
| Open Access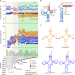 Relaxed sequence constraints favor mutational freedom in idiosyncratic metazoan mitochondrial tRNAs
Relaxed sequence constraints favor mutational freedom in idiosyncratic metazoan mitochondrial tRNAs
Bilaterian mitochondria-encoded tRNA genes accumulate mutations at higher rates than their cytoplasmic tRNA counterparts, resulting in idiosyncratic structures. Here the authors suggest an evolutionary basis for the observed mutational freedom of mitochondrial (mt) tRNAs and reveal the associated co-adaptive structural and functional changes in mt aminoacyl-tRNA synthetases.
- Bernhard Kuhle
- , Joseph Chihade
- & Paul Schimmel
-
Article
| Open Access Biogenesis and functions of aminocarboxypropyluridine in tRNA
Biogenesis and functions of aminocarboxypropyluridine in tRNA
E. coli and human tRNAs contain 3-(3-amino-3-carboxypropyl)uridine (acp3U) modification. Here the authors identify E. coli TapT and human DTWD1/2 as tRNA aminocarboxypropyltransferases responsible for acp3U formation. Inhibition of acp3U modification results in genome instability in heat-stressed E. coli and growth defects in human cells.
- Mayuko Takakura
- , Kensuke Ishiguro
- & Tsutomu Suzuki
-
Article
| Open Access Defects in t6A tRNA modification due to GON7 and YRDC mutations lead to Galloway-Mowat syndrome
Defects in t6A tRNA modification due to GON7 and YRDC mutations lead to Galloway-Mowat syndrome
The biosynthesis of N6-threonylcarbamoylated adenosine 37 in tRNA (t6A) involves the YRDC enzyme and the KEOPS complex. Here, the authors report mutations in YRDC and the KEOPS component GON7 in Galloway-Mowat syndrome and determine the crystal structure of a GON7-containg subcomplex that suggests a role in KEOPS complex stability.
- Christelle Arrondel
- , Sophia Missoury
- & Géraldine Mollet
-
Article
| Open Access Toxin-mediated ribosome stalling reprograms the Mycobacterium tuberculosis proteome
Toxin-mediated ribosome stalling reprograms the Mycobacterium tuberculosis proteome
MazF endoribonucleases are thought to arrest growth of Mycobacterium tuberculosis by global translation inhibition. Here, Barth et al. show that MazF-mt9 cleaves a specific tRNA, leading to ribosome stalling at AAA codons and thus selective mRNA degradation and changes in transcriptome and proteome.
- Valdir C. Barth
- , Ju-Mei Zeng
- & Nancy A. Woychik
-
Article
| Open Access Crystal structure of an adenovirus virus-associated RNA
Crystal structure of an adenovirus virus-associated RNA
Adenovirus Virus-Associated (VA) noncoding RNAs interfere with the host system by mimicking double stranded RNAs. Here, the authors report a 2.7 Å crystal structure of VA-I RNA providing an understanding of its function.
- Iris V. Hood
- , Jackson M. Gordon
- & Jinwei Zhang
-
Article
| Open Access Dual pathways of tRNA hydroxylation ensure efficient translation by expanding decoding capability
Dual pathways of tRNA hydroxylation ensure efficient translation by expanding decoding capability
5-carboxymethoxyuridine (cmo5U) is one of the RNA modifications found in bacterial tRNA anticodons. Here the authors show that the first step of cmo5U biosynthesis from uridine is mediated by either one of two parallel factors, TrhP or TrhO, and that cmo5U modification is required for efficient translation.
- Yusuke Sakai
- , Satoshi Kimura
- & Tsutomu Suzuki
-
Article
| Open Access Microbiome characterization by high-throughput transfer RNA sequencing and modification analysis
Microbiome characterization by high-throughput transfer RNA sequencing and modification analysis
The authors present a molecular approach and a software suite for high-throughput sequencing and characterization of microbial tRNA transcripts, which provide unique physiological insights into complex environmental microbes.
- Michael H. Schwartz
- , Haipeng Wang
- & A. Murat Eren
-
Article
| Open Access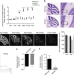 Elongator mutation in mice induces neurodegeneration and ataxia-like behavior
Elongator mutation in mice induces neurodegeneration and ataxia-like behavior
Elp6 is a component of the Elongator complex that regulates tRNAs and translation. Here the authors identify a mutation in the Elp6 gene that contributes to the cerebellar ataxia-like phenotype in a mutant mouse.
- Marija Kojic
- , Monika Gaik
- & Brandon J. Wainwright
-
Article
| Open Access Evolutionary instability of CUG-Leu in the genetic code of budding yeasts
Evolutionary instability of CUG-Leu in the genetic code of budding yeasts
The genetic code for amino acids is nearly universal, and among eukaryotic nuclear genomes the only known reassignments are of codon CUG in yeasts. Here, the authors identify a third independent CUG transition in budding yeasts that is still ongoing with alternative tRNAs present in the genome.
- Tadeusz Krassowski
- , Aisling Y. Coughlan
- & Kenneth H. Wolfe
-
Article
| Open Access A chiral selectivity relaxed paralog of DTD for proofreading tRNA mischarging in Animalia
A chiral selectivity relaxed paralog of DTD for proofreading tRNA mischarging in Animalia
The number of tRNA isodecoders has expanded significantly during evolution, which has resulted in ambiguity in tRNA selection by synthetases. Here the authors identify and characterize a dedicated proofreading factor that eliminates errors caused by ambiguity in tRNA selection by eukaryotic tRNA synthetases.
- Santosh Kumar Kuncha
- , Mohd Mazeed
- & Rajan Sankaranarayanan
-
Article
| Open Access Double mimicry evades tRNA synthetase editing by toxic vegetable-sourced non-proteinogenic amino acid
Double mimicry evades tRNA synthetase editing by toxic vegetable-sourced non-proteinogenic amino acid
Non-proteinogenic (np) amino acids in the food chain present challenges for the human translation machinery. Here the authors show that, while AlaRS and ProRS activate toxic np azetidine-2-carboxylic acid (Aze) present in sugar beets and lilies, only the AlaRS editing system rejects Aze.
- Youngzee Song
- , Huihao Zhou
- & Paul Schimmel
-
Article
| Open Access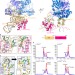 Structural basis for tRNA-dependent cysteine biosynthesis
Structural basis for tRNA-dependent cysteine biosynthesis
tRNA-dependent cysteine biosynthesis is catalyzed by the transsulfursome protein complex. Here, the authors use a multidisciplinary approach to structurally characterize the archaeal transsulfursome and propose a model for tRNA channeling in the complex.
- Meirong Chen
- , Koji Kato
- & Min Yao
-
Article
| Open Access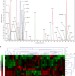 tRNA-mediated codon-biased translation in mycobacterial hypoxic persistence
tRNA-mediated codon-biased translation in mycobacterial hypoxic persistence
Mycobacteria can adapt to the stress of human infection by entering a dormant state. Here the authors show that hypoxia-induced dormancy in M. bovisBCG involves the reprogramming of tRNA wobble modifications and copy numbers, coupled with biased use of synonymous codons in survival genes.
- Yok Hian Chionh
- , Megan McBee
- & Peter C. Dedon
-
Article
| Open Access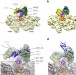 Cryo-EM study of start codon selection during archaeal translation initiation
Cryo-EM study of start codon selection during archaeal translation initiation
Initiation factor eIF2, common to eukaryotes and archaea, is a central actor in translation initiation. Here the authors describe two cryo-EM structures of archaeal 30S initiation complexes that provide a novel view of the central role that e/aIF2 plays in start codon selection.
- Pierre-Damien Coureux
- , Christine Lazennec-Schurdevin
- & Yves Mechulam
-
Article
| Open Access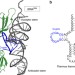 Essential structural elements in tRNAPro for EF-P-mediated alleviation of translation stalling
Essential structural elements in tRNAPro for EF-P-mediated alleviation of translation stalling
Ribosomes tend to stall during the translation of consecutive proline residues, which can be rescued by the co-translational factor EF-P. Here the authors identify a structural element of tRNAProresponsible for specific recognition by EF-P and stimulation of Pro-Pro peptide bond formation.
- Takayuki Katoh
- , Ingo Wohlgemuth
- & Hiroaki Suga
-
Article
| Open Access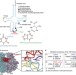 Novel base-pairing interactions at the tRNA wobble position crucial for accurate reading of the genetic code
Novel base-pairing interactions at the tRNA wobble position crucial for accurate reading of the genetic code
The anticodon loops of almost all tRNAs contain modifications known to be important for their function. Here the authors use crystallography to provide new mechanistic insights into how the modification at the wobble position of the E. coli tRNALysUUUassists in discrimination between cognate and near-cognate codons.
- Alexey Rozov
- , Natalia Demeshkina
- & Gulnara Yusupova
-
Article |
 Growth-regulating Mycobacterium tuberculosis VapC-mt4 toxin is an isoacceptor-specific tRNase
Growth-regulating Mycobacterium tuberculosis VapC-mt4 toxin is an isoacceptor-specific tRNase
Toxin–antitoxin systems of the Vap class regulate the growth of several bacterial pathogens including Mycobacterium tuberculosis. Here, the authors show that toxin VapC-mt4 arrests M. tuberculosis growth by specifically cleaving three tRNAs at a single site in their anticodon stem loop, leading to translation inhibition.
- Jonathan W. Cruz
- , Jared D. Sharp
- & Nancy A. Woychik
-
Article
| Open Access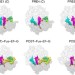 Fluctuations between multiple EF-G-induced chimeric tRNA states during translocation on the ribosome
Fluctuations between multiple EF-G-induced chimeric tRNA states during translocation on the ribosome
EF-G enhances the rate of tRNA–mRNA translocation on the ribosome. Here the authors use single-molecule FRET to follow tRNA translocation in real time, identifying new chimeric intermediates and suggesting how EF-G binding and GTP hydrolysis change the energetic landscape of translocation to accelerate forward tRNA movement.
- Sarah Adio
- , Tamara Senyushkina
- & Marina V. Rodnina
-
Article
| Open Access Structural basis for full-spectrum inhibition of translational functions on a tRNA synthetase
Structural basis for full-spectrum inhibition of translational functions on a tRNA synthetase
Borrelidin is an antibiotic with antimicrobial, antifungal, antimalarial and immunosuppressive activity that targets threonyl-tRNA synthetase. Here the authors show that borrelidin functions by preventing binding of all three ThrRS substrates and inducing a distinct, non-productive, conformation of the enzyme.
- Pengfei Fang
- , Xue Yu
- & Min Guo
-
Article |
 Plant tumour biocontrol agent employs a tRNA-dependent mechanism to inhibit leucyl-tRNA synthetase
Plant tumour biocontrol agent employs a tRNA-dependent mechanism to inhibit leucyl-tRNA synthetase
Agrobacterium radiobacter strain K84 generates an antibiotic targeting pathogenic strains of Agrobacterium tumefaciens, enabling its use as a biocontrol to prevent infection of crops. Here the authors show that this antibiotic inhibits leucyl-tRNA synthetases via an unusual mechanism that depends on binding of tRNALeu.
- Shaileja Chopra
- , Andrés Palencia
- & John S. Reader
-
Article
| Open Access Structural insights into protein-only RNase P complexed with tRNA
Structural insights into protein-only RNase P complexed with tRNA
RNase P is a key enzyme implicated in transfer RNA maturation that removes the 5′-leader sequences from transfer RNA precursors. In this study, a biophysical characterization of a novel protein-only variant of RNase P, known as PRORP (PROteinaceous RNase P), reveals that transfer RNA recognition by PRORP is similar to that by ribonucleoprotein RNase P.
- Anthony Gobert
- , Franziska Pinker
- & Philippe Giegé
-
Article |
 Reprogramming of tRNA modifications controls the oxidative stress response by codon-biased translation of proteins
Reprogramming of tRNA modifications controls the oxidative stress response by codon-biased translation of proteins
In response to stress, yeast cells selectively translate proteins that can enhance cell survival. In this study, the authors find that tRNALEU(CAA)in yeast cells is modified in response to oxidative stress by a methyltransferase and this results in the selective translation of the mRNA for these genes.
- Clement T.Y. Chan
- , Yan Ling Joy Pang
- & Peter C. Dedon
-
Article |
 Potential for interdependent development of tRNA determinants for aminoacylation and ribosome decoding
Potential for interdependent development of tRNA determinants for aminoacylation and ribosome decoding
Aminoacyl-transfer RNA synthetases are conserved between bacteria and eukaryotes; however, bacterial enzymes cannot acylate eukaryote tRNAs. Now, fusion of a human and bacterial enzyme is shown to overcome the species barrier and confer tRNA specificity during both codon selection and proofreading on the ribosome.
- Cuiping Liu
- , Howard Gamper
- & Ya-Ming Hou
-
Article |
 ALKBH8-mediated formation of a novel diastereomeric pair of wobble nucleosides in mammalian tRNA
ALKBH8-mediated formation of a novel diastereomeric pair of wobble nucleosides in mammalian tRNA
Uridines at the wobble position of transfer RNA anticodons are usually modified to allow efficient decoding of messenger RNA codons. In this study, ALKBH8 is shown to be a bifunctional transfer RNA modification enzyme required for the formation of a novel diastereomeric pair of modified wobble uridines.
- Erwin van den Born
- , Cathrine B. Vågbø
- & Pål Ø. Falnes

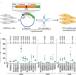 Translation velocity determines the efficacy of engineered suppressor tRNAs on pathogenic nonsense mutations
Translation velocity determines the efficacy of engineered suppressor tRNAs on pathogenic nonsense mutations