Featured
-
-
Article
| Open Access Bone marrow stromal cells induce chromatin remodeling in multiple myeloma cells leading to transcriptional changes
Bone marrow stromal cells induce chromatin remodeling in multiple myeloma cells leading to transcriptional changes
Bone marrow stromal cells (BMSCs) are known to promote the development of drug resistance. Here, the authors investigate the chromatin remodeling and associated changes in gene expression in the multiple myeloma (MM) cells following their interactions with BMSCs, which are also observed in extramedullary disease (EMD).
- Moritz Binder
- , Raphael E. Szalat
- & Nikhil C. Munshi
-
Article
| Open Access Nuclear actin structure regulates chromatin accessibility
Nuclear actin structure regulates chromatin accessibility
Intranuclear actin contributes to nuclear structure. Inducing actin remodeling within the nucleus regulates chromatin accessibility, and is associated with phenotypic outcomes in mesenchymal stem cells. As such, dynamic actin remodeling may modulate gene expression.
- Buer Sen
- , Zhihui Xie
- & Janet Rubin
-
Article
| Open Access Multiscale modelling of chromatin 4D organization in SARS-CoV-2 infected cells
Multiscale modelling of chromatin 4D organization in SARS-CoV-2 infected cells
In this work, the authors apply polymer models to reconstruct the 3D structure of the genome during SARS-CoV-2 infection and examine how the virus impacts key mechanisms of chromatin organization.
- Andrea M. Chiariello
- , Alex Abraham
- & Mario Nicodemi
-
Article
| Open Access MeCP2 binds to methylated DNA independently of phase separation and heterochromatin organisation
MeCP2 binds to methylated DNA independently of phase separation and heterochromatin organisation
The heterochromatic 'condensates' may not be conserved across mammals. This study highlights the influence of host genome on nuclear architecture and challenges the hypothesis that heterochromatin and MeCP2 undergo phase separation.
- Raphaël Pantier
- , Megan Brown
- & Adrian Bird
-
Article
| Open Access The miR-144/Hmgn2 regulatory axis orchestrates chromatin organization during erythropoiesis
The miR-144/Hmgn2 regulatory axis orchestrates chromatin organization during erythropoiesis
Differentiation of stem and progenitor cells is a highly regulated process. Here, the authors uncover miR-144 and its target Hmgn2 as the backbone of the genetic regulatory circuit that controls the terminal differentiation of erythrocytes in vertebrates.
- Dmitry A. Kretov
- , Leighton Folkes
- & Daniel Cifuentes
-
Article
| Open Access A fine-scale Arabidopsis chromatin landscape reveals chromatin conformation-associated transcriptional dynamics
A fine-scale Arabidopsis chromatin landscape reveals chromatin conformation-associated transcriptional dynamics
Plants utilize transcriptional dynamics to adapt to cold stress. Here, Zhang et al. describe a network of chromatin interactions between gene promoters across the Arabidopsis genome that could facilitate co-regulation of gene expression during cold stress.
- Yueying Zhang
- , Qianli Dong
- & Huakun Zhang
-
Article
| Open Access The PTM profiling of CTCF reveals the regulation of 3D chromatin structure by O-GlcNAcylation
The PTM profiling of CTCF reveals the regulation of 3D chromatin structure by O-GlcNAcylation
CTCF, which is known to play critical role in chromatin structure, undergoes post-translational modifications (PTMs). In this research, O-GlcNAcylation was found to inhibit CTCF binding, impacting 3D chromatin structure, gene expression and cellular development.
- Xiuxiao Tang
- , Pengguihang Zeng
- & Junjun Ding
-
Article
| Open Access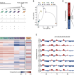 p53 rapidly restructures 3D chromatin organization to trigger a transcriptional response
p53 rapidly restructures 3D chromatin organization to trigger a transcriptional response
Here the authors uncover p53’s role as the master regulator of spatio-temporal genome organization. p53 controls the expression of 340 distal genes through newly formed and pre-existing loops between p53-bound enhancers and promoters.
- François Serra
- , Andrea Nieto-Aliseda
- & Biola M. Javierre
-
Article
| Open Access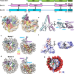 Structure of the human Bre1 complex bound to the nucleosome
Structure of the human Bre1 complex bound to the nucleosome
The structure of the nucleosome-bound human Bre1 complex reveals that its two RING domains bind the acidic patch and nucleosomal DNA, directing the E2 enzyme and ubiquitin for H2BK120-specific ubiquitination. The binding mode suggests a possible regulatory mechanism through nucleosomal DNA flexibility.
- Shuhei Onishi
- , Kotone Uchiyama
- & Toru Sengoku
-
Article
| Open Access BET inhibitors drive Natural Killer activation in non-small cell lung cancer via BRD4 and SMAD3
BET inhibitors drive Natural Killer activation in non-small cell lung cancer via BRD4 and SMAD3
Combination of BET inhibitors (BETi) with immunotherapy has been reported to be synergic for the treatment of non-small cell lung carcinoma (NSCLC). Here, the authors show that BETi-induced epigenetic reprogramming downregulates the expression of NK cell inhibitory receptors on NK cells, increasing their activation and cytotoxicity against NSCLC.
- Francesca Reggiani
- , Giovanna Talarico
- & Valentina Sancisi
-
Article
| Open Access Two DOT1 enzymes cooperatively mediate efficient ubiquitin-independent histone H3 lysine 76 tri-methylation in kinetoplastids
Two DOT1 enzymes cooperatively mediate efficient ubiquitin-independent histone H3 lysine 76 tri-methylation in kinetoplastids
Trypanosoma brucei DOT1A and DOT1B methylate H3K76 without H2B-ubiquitin. Based on structural and enzymatic data, Frisbie et al. reveal a mechanism of how these enzymes cooperatively and efficiently tri-methylate H3K76 in a ubiquitin-independent way.
- Victoria S. Frisbie
- , Hideharu Hashimoto
- & Erik W. Debler
-
Article
| Open Access Oncogenic enhancers prime quiescent metastatic cells to escape NK immune surveillance by eliciting transcriptional memory
Oncogenic enhancers prime quiescent metastatic cells to escape NK immune surveillance by eliciting transcriptional memory
Metastasis arises from disseminated tumour cells (DTCs) while the underlying mechanism of DTCs plasticity remains underexplored. Here, the authors show that spatially organized oncogenic enhancers on chromatin sustain the establishment of retinoic acid (RA)-stimulated transcriptional memory through activation of SOX9, supporting the escape of quiescent DTCs from NK-mediated immune surveillance.
- Daniela Michelatti
- , Sven Beyes
- & Alessio Zippo
-
Article
| Open Access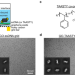 Functionalized graphene-oxide grids enable high-resolution cryo-EM structures of the SNF2h-nucleosome complex without crosslinking
Functionalized graphene-oxide grids enable high-resolution cryo-EM structures of the SNF2h-nucleosome complex without crosslinking
Nucleosome-protein complexes stick to the air-water interface and denature upon plunge freezing for cryoEM. Here, authors Chio and Palovcak et al. develop EM grids that protect such complexes and use these grids to study the ATP-dependent chromatin remodeler SNF2h.
- Un Seng Chio
- , Eugene Palovcak
- & Yifan Cheng
-
Article
| Open Access Kdm1a safeguards the topological boundaries of PRC2-repressed genes and prevents aging-related euchromatinization in neurons
Kdm1a safeguards the topological boundaries of PRC2-repressed genes and prevents aging-related euchromatinization in neurons
Kdm1a is a histone demethylase implicated in intellectual disability. Here, the authors show that removing Kdm1a in neurons of the adult mouse forebrain disrupts silencing of nonneuronal genes and chromatin organization, emphasizing its role in preserving neuronal genome integrity.
- Beatriz del Blanco
- , Sergio Niñerola
- & Ángel Barco
-
Article
| Open Access Implications of the three-dimensional chromatin organization for genome evolution in a fungal plant pathogen
Implications of the three-dimensional chromatin organization for genome evolution in a fungal plant pathogen
The spatial organization of eukaryotic genomes is linked to their biological functions. Here, the authors study the 3D genome organization of the phytopathogenic fungus Verticillium dahliae, revealing links to evolutionary features conserved throughout the Verticillium genus.
- David E. Torres
- , H. Martin Kramer
- & Bart P. H. J. Thomma
-
Article
| Open Access Cbp1 and Cren7 form chromatin-like structures that ensure efficient transcription of long CRISPR arrays
Cbp1 and Cren7 form chromatin-like structures that ensure efficient transcription of long CRISPR arrays
CRISPR arrays form the physical memory of prokaryotic adaptive immune systems by incorporating viral DNA sequences as spacers. Here, Blombach et al. show that transcription factor Cbp1 recruits chromatin protein Cren7 at CRISPR arrays, forming ‘chimeric’ chromatin-like structures that regulate expression of long CRISPR arrays in Sulfolobales archaea.
- Fabian Blombach
- , Michal Sýkora
- & Finn Werner
-
Article
| Open Access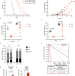 Caloric restriction leads to druggable LSD1-dependent cancer stem cells expansion
Caloric restriction leads to druggable LSD1-dependent cancer stem cells expansion
Caloric restriction (CR) has been demonstrated to have a role in tumour growth and therapy response but its effects on cancer stem cells are less known. Here, in Acute Promyelocytic Leukemia, the authors show that despite initial anti-tumour effect, CR drives the selection of leukaemia-initiating cells resulting in relapse which could be prevented by ablation of LSD1.
- Rani Pallavi
- , Elena Gatti
- & Pier Giuseppe Pelicci
-
Article
| Open Access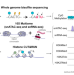 The chromatin landscape of healthy and injured cell types in the human kidney
The chromatin landscape of healthy and injured cell types in the human kidney
Comprehensive integration of gene expression with epigenetic features is needed to understand the transition of kidney cells from health to injury. Here, the authors integrate dual single nucleus RNA expression and chromatin accessibility, DNA methylation, and histone modifications to decipher the chromatin landscape of the kidney in reference and adaptive injury cell states, identifying a transcription factor network of ELF3, KLF6, and KLF10 which regulates adaptive repair and maladaptive failed repair.
- Debora L. Gisch
- , Michelle Brennan
- & Michael T. Eadon
-
Article
| Open Access Mapping nucleosome-resolution chromatin organization and enhancer-promoter loops in plants using Micro-C-XL
Mapping nucleosome-resolution chromatin organization and enhancer-promoter loops in plants using Micro-C-XL
The authors employ Micro-C-XL to investigate chromatin structures in plants, specifically focusing on nucleosome-resolution chromatin organizations and enhancer-promoter chromatin loops in Arabidopsis, rice, and soybean.
- Linhua Sun
- , Jingru Zhou
- & Hang He
-
Article
| Open Access Histone lactylation couples cellular metabolism with developmental gene regulatory networks
Histone lactylation couples cellular metabolism with developmental gene regulatory networks
While metabolic reprogramming has been shown to drive changes in cell identity, the link between cellular metabolism and gene expression remains poorly characterized. Here they show that histone lactylation couples metabolism and transcription during neural crest cell differentiation in the early embryo.
- Fjodor Merkuri
- , Megan Rothstein
- & Marcos Simoes-Costa
-
Article
| Open Access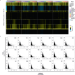 Cell-type differential targeting of SETDB1 prevents aberrant CTCF binding, chromatin looping, and cis-regulatory interactions
Cell-type differential targeting of SETDB1 prevents aberrant CTCF binding, chromatin looping, and cis-regulatory interactions
Here, the authors show how the histone methyltransferase SETDB1 is involved in cell-type specific regulation of chromatin landscape by catalyzing H3K9me3, which antagonizes CTCF binding. They further define the subsequent transcriptomic impact.
- Phoebe Lut Fei Tam
- , Ming Fung Cheung
- & Danny Leung
-
Article
| Open Access Phase-separated CCER1 coordinates the histone-to-protamine transition and male fertility
Phase-separated CCER1 coordinates the histone-to-protamine transition and male fertility
Here the authors reveal that phase‐separated nuclear CCER1 condensates are required for male fertility by mediating chromatin condensation and histone epigenetic modification, while loss‐of‐function variants of human CCER1 are pathogenic in patients with nonobstructive azoospermia (NOA).
- Dongdong Qin
- , Yayun Gu
- & Zhibin Hu
-
Article
| Open Access Loss of cohesin regulator PDS5A reveals repressive role of Polycomb loops
Loss of cohesin regulator PDS5A reveals repressive role of Polycomb loops
Through a genetic screen, the authors find that the cohesin regulator PDS5A is required for Polycomb target gene silencing. Derepression upon cohesin dysregulation is linked to loss of Polycomb loops without change in repressive chromatin domains.
- Daniel Bsteh
- , Hagar F. Moussa
- & Oliver Bell
-
Article
| Open Access Structure of the complete Saccharomyces cerevisiae Rpd3S-nucleosome complex
Structure of the complete Saccharomyces cerevisiae Rpd3S-nucleosome complex
In this study, the authors present the cryogenic electron microscopy reconstruction of the Rpd3S complex engaged with a nucleosome. The corresponding model describes the interactions that facilitate histone deacetylation within gene bodies by the Rpd3S complex.
- Jonathan W. Markert
- , Seychelle M. Vos
- & Lucas Farnung
-
Article
| Open Access The environmentally-regulated interplay between local three-dimensional chromatin organisation and transcription of proVWX in E. coli
The environmentally-regulated interplay between local three-dimensional chromatin organisation and transcription of proVWX in E. coli
Here, the authors use the proVWXoperon of Escherichia coli as a model system to show how the nucleoid associated protein H-NS regulates gene expression in vivo by local chromatin remodelling.
- Fatema-Zahra M. Rashid
- , Frédéric G. E. Crémazy
- & Remus T. Dame
-
Article
| Open Access CHEX-seq detects single-cell genomic single-stranded DNA with catalytical potential
CHEX-seq detects single-cell genomic single-stranded DNA with catalytical potential
The in situ single-stranded open chromatin landscape is dynamically regulated in single cells. In their efforts to understand brain cells’ functional dynamics and to complement the other single-cell chromatin approaches, the authors present a method named CHEX-seq (CHromatin EXposed).
- Youtao Lu
- , Jaehee Lee
- & James Eberwine
-
Article
| Open Access Release of Histone H3K4-reading transcription factors from chromosomes in mitosis is independent of adjacent H3 phosphorylation
Release of Histone H3K4-reading transcription factors from chromosomes in mitosis is independent of adjacent H3 phosphorylation
Methyl-phos switches on histones have been shown to regulate reader protein displacement from chromatin. However, in this study the authors find that H3T3ph is not required to remove transcription factors from H3K4me3 in mitosis. This might help to preserve promoter properties during cell division.
- Rebecca J. Harris
- , Maninder Heer
- & Jonathan M. G. Higgins
-
Article
| Open Access Chromatinization modulates topoisomerase II processivity
Chromatinization modulates topoisomerase II processivity
Here the authors discover that chromatin stimulates topoisomerase II function by enabling the enzyme to achieve exceptionally high processivity and efficient supercoiling relaxation, even under low torsional stress.
- Jaeyoon Lee
- , Meiling Wu
- & Michelle D. Wang
-
Article
| Open Access PAPγ associates with PAXT nuclear exosome to control the abundance of PROMPT ncRNAs
PAPγ associates with PAXT nuclear exosome to control the abundance of PROMPT ncRNAs
Pervasive transcription of the human genome generates an abundance of RNAs that must be processed and degraded by the nuclear RNA exosome. Here the authors show that polyA polymerase gamma (PAPγ) associates with PAXT providing key insights into the direct targeting of PROMPT ncRNAs by PAXT at their genomic sites.
- Xavier Contreras
- , David Depierre
- & Rosemary Kiernan
-
Article
| Open Access UV-induced G4 DNA structures recruit ZRF1 which prevents UV-induced senescence
UV-induced G4 DNA structures recruit ZRF1 which prevents UV-induced senescence
Senescence can be activated in a DNA damage-dependent and -independent manner. Here the authors reveal in cell lines that recruitment of the factor ZRF1 at G4 DNA structures can prevents UV-induced senescence.
- Alessio De Magis
- , Michaela Limmer
- & Katrin Paeschke
-
Article
| Open Access HMGA2 directly mediates chromatin condensation in association with neuronal fate regulation
HMGA2 directly mediates chromatin condensation in association with neuronal fate regulation
High-mobility group AT-hook (HMGA) proteins 1 and 2 are nonhistone chromatin proteins involved in different biological processes. Here the authors reveal that HMGA2 is a bona fide chromatin condensation factor that undergoes liquid–liquid phase separation, and that its chromatin condensation activity is important for the maintenance of mouse neural progenitor cells.
- Naohiro Kuwayama
- , Tomoya Kujirai
- & Yukiko Gotoh
-
Article
| Open Access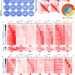 Regulation of CTCF loop formation during pancreatic cell differentiation
Regulation of CTCF loop formation during pancreatic cell differentiation
Here the authors show that differentiation of hESCs into pancreatic cells involves changes in CTCF loops by alteration of pioneer factor recruitment, histone modifications, and DNA methylation, leading to changes in enhancer-promoter interactions and gene expression.
- Xiaowen Lyu
- , M. Jordan Rowley
- & Victor G. Corces
-
Article
| Open Access An autoinhibited state of 53BP1 revealed by small molecule antagonists and protein engineering
An autoinhibited state of 53BP1 revealed by small molecule antagonists and protein engineering
Here, using small molecule antagonists and protein engineering, the authors identify an autoinhibited state of 53BP1 leading to its chromatin binding surface being obstructed. Such small molecule ligands present a potential avenue for the development of cancer therapy drugs.
- Gaofeng Cui
- , Maria Victoria Botuyan
- & Georges Mer
-
Article
| Open Access Context-dependent perturbations in chromatin folding and the transcriptome by cohesin and related factors
Context-dependent perturbations in chromatin folding and the transcriptome by cohesin and related factors
Enhancer–promoter looping and topologically associating domain are at the base of chromatin structures. Here the authors present a computational workflow in which multi-omics datasets are compared systematically to explore how three-dimensional (3D) structure and gene expression are regulated by cohesin and related factors.
- Ryuichiro Nakato
- , Toyonori Sakata
- & Katsuhiko Shirahige
-
Article
| Open Access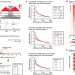 Multi-feature clustering of CTCF binding creates robustness for loop extrusion blocking and Topologically Associating Domain boundaries
Multi-feature clustering of CTCF binding creates robustness for loop extrusion blocking and Topologically Associating Domain boundaries
Most mammalian TAD boundaries, which separate functional chromosomal domains, bind the CTCF protein. Here, the authors identify multi-level clustering of CTCF binding sites at TAD boundaries and confirm their individual contribution to TAD formation.
- Li-Hsin Chang
- , Sourav Ghosh
- & Daan Noordermeer
-
Article
| Open Access SHIELD: a platform for high-throughput screening of barrier-type DNA elements in human cells
SHIELD: a platform for high-throughput screening of barrier-type DNA elements in human cells
Chromatin boundary elements are hard to define and characterize. Here the authors report Site-specific Heterochromatin Insertion of Elements at Lamina-associated Domains (SHIELD) for high-throughput screening of barrier-type DNA elements in human cells.
- Meng Zhang
- , Mary Elisabeth Ehmann
- & Huimin Zhao
-
Article
| Open Access NEUROD1 reinforces endocrine cell fate acquisition in pancreatic development
NEUROD1 reinforces endocrine cell fate acquisition in pancreatic development
Errors during pancreas development and specification of the endocrine lineage can result in severe neonatal diabetes. Here they show that loss of NEUROD1 leads to disturbances in endocrine cell identity acquisition during pancreas development, with cellular reprogramming occurring at the single-cell level within both α- and β-cell populations.
- Romana Bohuslavova
- , Valeria Fabriciova
- & Gabriela Pavlinkova
-
Article
| Open Access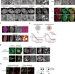 Chemical-induced phase transition and global conformational reorganization of chromatin
Chemical-induced phase transition and global conformational reorganization of chromatin
The authors show that adriamycin induces global changes on chromatin conformation associated with phase transitions mediated through Histone H1.
- Tengfei Wang
- , Shuxiang Shi
- & Chao-Po Lin
-
Article
| Open Access The COMPASS subunit Spp1 protects nascent DNA at the Tus/Ter replication fork barrier by limiting DNA availability to nucleases
The COMPASS subunit Spp1 protects nascent DNA at the Tus/Ter replication fork barrier by limiting DNA availability to nucleases
The Spp1 subunit of Set1C is recruited to the Tus/Ter replication fork barrier via its PHD domain and protects the replication fork by preventing excessive ssDNA formation, stressing the importance of chromatin structure in the replication stress response.
- Nagham Ghaddar
- , Yves Corda
- & Vincent Géli
-
Article
| Open Access TAZ2 truncation confers overactivation of p300 and cellular vulnerability to HDAC inhibition
TAZ2 truncation confers overactivation of p300 and cellular vulnerability to HDAC inhibition
The E1A binding protein p300 and its close paralogue CREB-binding protein are transcriptional coactivators with intrinsic histone acetyltransferase activity. Here, the authors show that the TAZ2 domain of p300 has a HAT autoinhibitory function that is relieved upon binding of transcription factors.
- Longxia Xu
- , Hongwen Xuan
- & Xiaobing Shi
-
Article
| Open Access MLL-AF4 cooperates with PAF1 and FACT to drive high-density enhancer interactions in leukemia
MLL-AF4 cooperates with PAF1 and FACT to drive high-density enhancer interactions in leukemia
Previous studies have reported MLL-AF4 binding at intragenic and intergenic enhancers, however, the role of MLL-AF4 in enhancer function remains to be investigated. Here, the authors show that MLL-AF4 cooperates with PAF1 and FACT at enhancers to promote high-density interactions with oncogene promoters in leukemia.
- Nicholas T. Crump
- , Alastair L. Smith
- & Thomas A. Milne
-
Article
| Open Access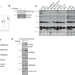 Histone H3 serine-57 is a CHK1 substrate whose phosphorylation affects DNA repair
Histone H3 serine-57 is a CHK1 substrate whose phosphorylation affects DNA repair
Histone post-translational modifications control several fundamental processes on DNA. Here, the authors describe a conserved phosphorylation of histone H3 on the globular core and show that it loosens the nucleosome and regulates DNA repair.
- Nikolaos Parisis
- , Pablo D. Dans
- & Daniel Fisher
-
Article
| Open Access Epigenetic and molecular coordination between HDAC2 and SMAD3-SKI regulates essential brain tumour stem cell characteristics
Epigenetic and molecular coordination between HDAC2 and SMAD3-SKI regulates essential brain tumour stem cell characteristics
The role of histone deacetylases (HDACs) in glioblastoma brain tumour stem cells (BTSCs) remains to be explored. Here, pharmacological inhibition and genetic loss of function approaches show that HDAC2 leads to the maintenance of BTSC growth and self-renewal through its association with the components of the TGF-β signalling pathway.
- Ravinder K. Bahia
- , Xiaoguang Hao
- & Samuel Weiss
-
Article
| Open Access Hdac1 and Hdac2 regulate the quiescent state and survival of hair-follicle mesenchymal niche
Hdac1 and Hdac2 regulate the quiescent state and survival of hair-follicle mesenchymal niche
Cell division and stem cell maintenance are tightly linked. Here they show that in the hair follicle an epigenetic mechanism maintains mesenchymal niche dormancy in a highly proliferative microenvironment while repurposing mitogenic signaling to orchestrate the hair cycle clock.
- Hadas Sibony-Benyamini
- , Emil Aamar
- & David Enshell-Seijffers
-
Article
| Open Access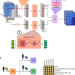 Getting personal with epigenetics: towards individual-specific epigenomic imputation with machine learning
Getting personal with epigenetics: towards individual-specific epigenomic imputation with machine learning
The authors present eDICE, an attention-based model that enables accurate imputation of missing portions of the observed epigenetic landscape, and show that eDICE can be used to predict individualspecific epigenomic variation in the EN-TEx dataset.
- Alex Hawkins-Hooker
- , Giovanni Visonà
- & Gabriele Schweikert
-
Article
| Open Access The AT-hook is an evolutionarily conserved auto-regulatory domain of SWI/SNF required for cell lineage priming
The AT-hook is an evolutionarily conserved auto-regulatory domain of SWI/SNF required for cell lineage priming
This study demonstrates that an evolutionary conserved, autoregulatory ‘AT-hook’ domain of SWI/SNF regulates gene transcription and enhancer activation by modulating SWI/SNF intrinsic catalytic activity and is critical for cell lineage priming.
- Dhurjhoti Saha
- , Solomon Hailu
- & Blaine Bartholomew
-
Article
| Open Access Molecular insights into Spindlin1-HBx interplay and its impact on HBV transcription from cccDNA minichromosome
Molecular insights into Spindlin1-HBx interplay and its impact on HBV transcription from cccDNA minichromosome
The mechanism by which the chromatinized HBV genome is transcribed remains poorly understood. In this study, Liu et al. demonstrate how HBx exploits Spindlin1, a histone methylation reader, to overcome heterochromatin barriers and enhance HBV transcription from the cccDNA minichromosome.
- Wei Liu
- , Qiyan Yao
- & Haitao Li
-
Article
| Open Access Ezh2 emerges as an epigenetic checkpoint regulator during monocyte differentiation limiting cardiac dysfunction post-MI
Ezh2 emerges as an epigenetic checkpoint regulator during monocyte differentiation limiting cardiac dysfunction post-MI
Modulating pro-inflammatory immune cell kinetics after myocardial infarction is a critical step to prevent heart dysfunction. In this study, the authors show that Ezh2 pharmacological inhibition, acting as an epigenetic checkpoint in monocytes and macrophages, prevents myocardial infarction-induced cardiac dysfunction.
- Julie Rondeaux
- , Déborah Groussard
- & Sylvain Fraineau
-
Article
| Open Access MOF-mediated histone H4 Lysine 16 acetylation governs mitochondrial and ciliary functions by controlling gene promoters
MOF-mediated histone H4 Lysine 16 acetylation governs mitochondrial and ciliary functions by controlling gene promoters
Here the authors show that epigenetic regulation through the histone acetyltransferase MOF and the acetylation of histone H4 lysine 16 affect essential functions in the mitochondria and primary cilia. This regulation is important to promote epidermal differentiation and hair follicle formation.
- Dongmei Wang
- , Haimin Li
- & Rui Yi

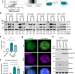 The nucleic acid binding protein SFPQ represses EBV lytic reactivation by promoting histone H1 expression
The nucleic acid binding protein SFPQ represses EBV lytic reactivation by promoting histone H1 expression