Article
|
Open Access
Featured
-
-
Article
| Open Access TAD border deletion at the Kit locus causes tissue-specific ectopic activation of a neighboring gene
TAD border deletion at the Kit locus causes tissue-specific ectopic activation of a neighboring gene
Research on the Kit locus shows TAD boundary deletion may or may not trigger ectopic gene activation in different cell types, influenced by active enhancers’ position relative gene promoters.
- Evelyn Kabirova
- , Anastasiya Ryzhkova
- & Nariman Battulin
-
Article
| Open Access DNA methylation-based high-resolution mapping of long-distance chromosomal interactions in nucleosome-depleted regions
DNA methylation-based high-resolution mapping of long-distance chromosomal interactions in nucleosome-depleted regions
Here, the authors present MTAC, a method to map chromosomal interactions in budding yeast. By applying MTAC to various viewpoints, they find that most of the long-distance chromosomal interactions detected by MTAC reflect tethering by the nuclear pore complexes.
- Yi Li
- , James Lee
- & Lu Bai
-
Article
| Open Access Multiscale modelling of chromatin 4D organization in SARS-CoV-2 infected cells
Multiscale modelling of chromatin 4D organization in SARS-CoV-2 infected cells
In this work, the authors apply polymer models to reconstruct the 3D structure of the genome during SARS-CoV-2 infection and examine how the virus impacts key mechanisms of chromatin organization.
- Andrea M. Chiariello
- , Alex Abraham
- & Mario Nicodemi
-
Article
| Open Access MeCP2 binds to methylated DNA independently of phase separation and heterochromatin organisation
MeCP2 binds to methylated DNA independently of phase separation and heterochromatin organisation
The heterochromatic 'condensates' may not be conserved across mammals. This study highlights the influence of host genome on nuclear architecture and challenges the hypothesis that heterochromatin and MeCP2 undergo phase separation.
- Raphaël Pantier
- , Megan Brown
- & Adrian Bird
-
Article
| Open Access A fine-scale Arabidopsis chromatin landscape reveals chromatin conformation-associated transcriptional dynamics
A fine-scale Arabidopsis chromatin landscape reveals chromatin conformation-associated transcriptional dynamics
Plants utilize transcriptional dynamics to adapt to cold stress. Here, Zhang et al. describe a network of chromatin interactions between gene promoters across the Arabidopsis genome that could facilitate co-regulation of gene expression during cold stress.
- Yueying Zhang
- , Qianli Dong
- & Huakun Zhang
-
Article
| Open Access The PTM profiling of CTCF reveals the regulation of 3D chromatin structure by O-GlcNAcylation
The PTM profiling of CTCF reveals the regulation of 3D chromatin structure by O-GlcNAcylation
CTCF, which is known to play critical role in chromatin structure, undergoes post-translational modifications (PTMs). In this research, O-GlcNAcylation was found to inhibit CTCF binding, impacting 3D chromatin structure, gene expression and cellular development.
- Xiuxiao Tang
- , Pengguihang Zeng
- & Junjun Ding
-
Article
| Open Access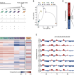 p53 rapidly restructures 3D chromatin organization to trigger a transcriptional response
p53 rapidly restructures 3D chromatin organization to trigger a transcriptional response
Here the authors uncover p53’s role as the master regulator of spatio-temporal genome organization. p53 controls the expression of 340 distal genes through newly formed and pre-existing loops between p53-bound enhancers and promoters.
- François Serra
- , Andrea Nieto-Aliseda
- & Biola M. Javierre
-
Article
| Open Access Kdm1a safeguards the topological boundaries of PRC2-repressed genes and prevents aging-related euchromatinization in neurons
Kdm1a safeguards the topological boundaries of PRC2-repressed genes and prevents aging-related euchromatinization in neurons
Kdm1a is a histone demethylase implicated in intellectual disability. Here, the authors show that removing Kdm1a in neurons of the adult mouse forebrain disrupts silencing of nonneuronal genes and chromatin organization, emphasizing its role in preserving neuronal genome integrity.
- Beatriz del Blanco
- , Sergio Niñerola
- & Ángel Barco
-
Article
| Open Access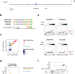 Cbp1 and Cren7 form chromatin-like structures that ensure efficient transcription of long CRISPR arrays
Cbp1 and Cren7 form chromatin-like structures that ensure efficient transcription of long CRISPR arrays
CRISPR arrays form the physical memory of prokaryotic adaptive immune systems by incorporating viral DNA sequences as spacers. Here, Blombach et al. show that transcription factor Cbp1 recruits chromatin protein Cren7 at CRISPR arrays, forming ‘chimeric’ chromatin-like structures that regulate expression of long CRISPR arrays in Sulfolobales archaea.
- Fabian Blombach
- , Michal Sýkora
- & Finn Werner
-
Article
| Open Access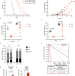 Caloric restriction leads to druggable LSD1-dependent cancer stem cells expansion
Caloric restriction leads to druggable LSD1-dependent cancer stem cells expansion
Caloric restriction (CR) has been demonstrated to have a role in tumour growth and therapy response but its effects on cancer stem cells are less known. Here, in Acute Promyelocytic Leukemia, the authors show that despite initial anti-tumour effect, CR drives the selection of leukaemia-initiating cells resulting in relapse which could be prevented by ablation of LSD1.
- Rani Pallavi
- , Elena Gatti
- & Pier Giuseppe Pelicci
-
Article
| Open Access Mapping nucleosome-resolution chromatin organization and enhancer-promoter loops in plants using Micro-C-XL
Mapping nucleosome-resolution chromatin organization and enhancer-promoter loops in plants using Micro-C-XL
The authors employ Micro-C-XL to investigate chromatin structures in plants, specifically focusing on nucleosome-resolution chromatin organizations and enhancer-promoter chromatin loops in Arabidopsis, rice, and soybean.
- Linhua Sun
- , Jingru Zhou
- & Hang He
-
Article
| Open Access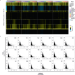 Cell-type differential targeting of SETDB1 prevents aberrant CTCF binding, chromatin looping, and cis-regulatory interactions
Cell-type differential targeting of SETDB1 prevents aberrant CTCF binding, chromatin looping, and cis-regulatory interactions
Here, the authors show how the histone methyltransferase SETDB1 is involved in cell-type specific regulation of chromatin landscape by catalyzing H3K9me3, which antagonizes CTCF binding. They further define the subsequent transcriptomic impact.
- Phoebe Lut Fei Tam
- , Ming Fung Cheung
- & Danny Leung
-
Article
| Open Access Loss of cohesin regulator PDS5A reveals repressive role of Polycomb loops
Loss of cohesin regulator PDS5A reveals repressive role of Polycomb loops
Through a genetic screen, the authors find that the cohesin regulator PDS5A is required for Polycomb target gene silencing. Derepression upon cohesin dysregulation is linked to loss of Polycomb loops without change in repressive chromatin domains.
- Daniel Bsteh
- , Hagar F. Moussa
- & Oliver Bell
-
Article
| Open Access The environmentally-regulated interplay between local three-dimensional chromatin organisation and transcription of proVWX in E. coli
The environmentally-regulated interplay between local three-dimensional chromatin organisation and transcription of proVWX in E. coli
Here, the authors use the proVWXoperon of Escherichia coli as a model system to show how the nucleoid associated protein H-NS regulates gene expression in vivo by local chromatin remodelling.
- Fatema-Zahra M. Rashid
- , Frédéric G. E. Crémazy
- & Remus T. Dame
-
Article
| Open Access CHEX-seq detects single-cell genomic single-stranded DNA with catalytical potential
CHEX-seq detects single-cell genomic single-stranded DNA with catalytical potential
The in situ single-stranded open chromatin landscape is dynamically regulated in single cells. In their efforts to understand brain cells’ functional dynamics and to complement the other single-cell chromatin approaches, the authors present a method named CHEX-seq (CHromatin EXposed).
- Youtao Lu
- , Jaehee Lee
- & James Eberwine
-
Article
| Open Access UV-induced G4 DNA structures recruit ZRF1 which prevents UV-induced senescence
UV-induced G4 DNA structures recruit ZRF1 which prevents UV-induced senescence
Senescence can be activated in a DNA damage-dependent and -independent manner. Here the authors reveal in cell lines that recruitment of the factor ZRF1 at G4 DNA structures can prevents UV-induced senescence.
- Alessio De Magis
- , Michaela Limmer
- & Katrin Paeschke
-
Article
| Open Access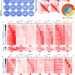 Regulation of CTCF loop formation during pancreatic cell differentiation
Regulation of CTCF loop formation during pancreatic cell differentiation
Here the authors show that differentiation of hESCs into pancreatic cells involves changes in CTCF loops by alteration of pioneer factor recruitment, histone modifications, and DNA methylation, leading to changes in enhancer-promoter interactions and gene expression.
- Xiaowen Lyu
- , M. Jordan Rowley
- & Victor G. Corces
-
Article
| Open Access Context-dependent perturbations in chromatin folding and the transcriptome by cohesin and related factors
Context-dependent perturbations in chromatin folding and the transcriptome by cohesin and related factors
Enhancer–promoter looping and topologically associating domain are at the base of chromatin structures. Here the authors present a computational workflow in which multi-omics datasets are compared systematically to explore how three-dimensional (3D) structure and gene expression are regulated by cohesin and related factors.
- Ryuichiro Nakato
- , Toyonori Sakata
- & Katsuhiko Shirahige
-
Article
| Open Access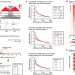 Multi-feature clustering of CTCF binding creates robustness for loop extrusion blocking and Topologically Associating Domain boundaries
Multi-feature clustering of CTCF binding creates robustness for loop extrusion blocking and Topologically Associating Domain boundaries
Most mammalian TAD boundaries, which separate functional chromosomal domains, bind the CTCF protein. Here, the authors identify multi-level clustering of CTCF binding sites at TAD boundaries and confirm their individual contribution to TAD formation.
- Li-Hsin Chang
- , Sourav Ghosh
- & Daan Noordermeer
-
Article
| Open Access SHIELD: a platform for high-throughput screening of barrier-type DNA elements in human cells
SHIELD: a platform for high-throughput screening of barrier-type DNA elements in human cells
Chromatin boundary elements are hard to define and characterize. Here the authors report Site-specific Heterochromatin Insertion of Elements at Lamina-associated Domains (SHIELD) for high-throughput screening of barrier-type DNA elements in human cells.
- Meng Zhang
- , Mary Elisabeth Ehmann
- & Huimin Zhao
-
Article
| Open Access MLL-AF4 cooperates with PAF1 and FACT to drive high-density enhancer interactions in leukemia
MLL-AF4 cooperates with PAF1 and FACT to drive high-density enhancer interactions in leukemia
Previous studies have reported MLL-AF4 binding at intragenic and intergenic enhancers, however, the role of MLL-AF4 in enhancer function remains to be investigated. Here, the authors show that MLL-AF4 cooperates with PAF1 and FACT at enhancers to promote high-density interactions with oncogene promoters in leukemia.
- Nicholas T. Crump
- , Alastair L. Smith
- & Thomas A. Milne
-
Article
| Open Access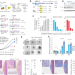 Engineered MED12 mutations drive leiomyoma-like transcriptional and metabolic programs by altering the 3D genome compartmentalization
Engineered MED12 mutations drive leiomyoma-like transcriptional and metabolic programs by altering the 3D genome compartmentalization
There are currently a lack of genetic models to study the biology of Uterine fibroids (UFs) tumours. Here the authors precisely engineer cells with mutant MED12 Gly-44 and generate myometrial smooth muscle cells (SMCs) that recapitulate major UFs-like cellular, transcriptional, and metabolic alterations.
- Kadir Buyukcelebi
- , Xintong Chen
- & Mazhar Adli
-
Article
| Open Access Single cohesin molecules generate force by two distinct mechanisms
Single cohesin molecules generate force by two distinct mechanisms
Cohesin protein complex is a molecular machine that extrudes DNA loops. Here, authors show that bending of the molecule’s long coiled coils is driven by thermal fluctuations and engagement of ATPase domains uses ATP energy to generate strong force.
- Georgii Pobegalov
- , Lee-Ya Chu
- & Maxim I. Molodtsov
-
Article
| Open Access Single-cell transcriptomics and epigenomics unravel the role of monocytes in neuroblastoma bone marrow metastasis
Single-cell transcriptomics and epigenomics unravel the role of monocytes in neuroblastoma bone marrow metastasis
The bone marrow is a common site of metastasis for neuroblastoma patients. Here, the authors perform single cell RNA-seq and ATAC-seq of bone marrow aspirates from 16 subjects and show conservation of tumor cell plasticity in metastases and identify tumor-to-bone marrow cell signals that trigger tumor promoting monocytes.
- Irfete S. Fetahu
- , Wolfgang Esser-Skala
- & Sabine Taschner-Mandl
-
Article
| Open Access Enhancer hijacking at the ARHGAP36 locus is associated with connective tissue to bone transformation
Enhancer hijacking at the ARHGAP36 locus is associated with connective tissue to bone transformation
Heterotopic ossification is a disorder characterized by the transformation of soft tissue to bone. In this study the authors report heterotopic ossification in a child caused by enhancer hijacking at the ARHGAP36 gene, the ectopic activation by an enhancer that it not its own.
- Uirá Souto Melo
- , Jerome Jatzlau
- & Malte Spielmann
-
Article
| Open Access MYC reshapes CTCF-mediated chromatin architecture in prostate cancer
MYC reshapes CTCF-mediated chromatin architecture in prostate cancer
The functional link between MYC and CTCF in prostate cancer remains to be investigated. Here, the authors highlight the role of MYC in rewiring chromatin architecture by interacting with CTCF protein.
- Zhao Wei
- , Song Wang
- & Haiyang Guo
-
Article
| Open Access Combinatorial effects on gene expression at the Lbx1/Fgf8 locus resolve split-hand/foot malformation type 3
Combinatorial effects on gene expression at the Lbx1/Fgf8 locus resolve split-hand/foot malformation type 3
Congenital limb defects are often associated with genomic rearrangements. Here they provide insights into the molecular mechanism underlying SHFM3-associated structural variations, offering a conceptual framework for how genomic rearrangements can alter gene expression and cause disease.
- Giulia Cova
- , Juliane Glaser
- & Stefan Mundlos
-
Article
| Open Access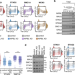 Different NIPBL requirements of cohesin-STAG1 and cohesin-STAG2
Different NIPBL requirements of cohesin-STAG1 and cohesin-STAG2
NIPBL is considered the cohesin loader. Here, the authors report that a drastic reduction of NIPBL levels reduces chromatin-bound cohesin-STAG2 genome wide while cohesin-STAG1 increases and can still be found at CTCF-bound sites but cannot form loops.
- Dácil Alonso-Gil
- , Ana Cuadrado
- & Ana Losada
-
Article
| Open Access HSFA1a modulates plant heat stress responses and alters the 3D chromatin organization of enhancer-promoter interactions
HSFA1a modulates plant heat stress responses and alters the 3D chromatin organization of enhancer-promoter interactions
Here the authors show genome-wide chromatin changes and transcriptional reprogramming occur in response to heat stress in tomato, leading to the transient formation of promoter-enhancer contacts to drive the expression of heat-stress responsive genes. Furthermore, they show chromatin spatial reorganization requires HSFA1a, a transcription factor (TF) essential for heat stress tolerance in tomato.
- Ying Huang
- , Jing An
- & Moussa Benhamed
-
Article
| Open Access Low input capture Hi-C (liCHi-C) identifies promoter-enhancer interactions at high-resolution
Low input capture Hi-C (liCHi-C) identifies promoter-enhancer interactions at high-resolution
Here the authors present the low input capture Hi-C (liCHi-C) method, a cost-effective, flexible method to map and robustly compare promoter interactomes at high resolution. liCHi-C identifies new disease-associated genes and structural variants to ultimately illuminate their pathogenic effects.
- Laureano Tomás-Daza
- , Llorenç Rovirosa
- & Biola M. Javierre
-
Article
| Open Access ChIATAC is an efficient strategy for multi-omics mapping of 3D epigenomes from low-cell inputs
ChIATAC is an efficient strategy for multi-omics mapping of 3D epigenomes from low-cell inputs
Methods connecting genes to their cis-regulatory elements usually requires millions of input cells. Here the authors combine proximity ligation in ChIA-PET and transposase accessibility in ATAC-seq, reporting ChIATAC, to map interactions between open chromatin loci in low numbers of input cells.
- Haoxi Chai
- , Harianto Tjong
- & Yijun Ruan
-
Article
| Open Access Reference panel guided topological structure annotation of Hi-C data
Reference panel guided topological structure annotation of Hi-C data
Predicting topological structures from Hi-C data provides insight into comprehending gene expression and regulation. Here, the authors present RefHiC, an attention-based deep learning framework that leverages a reference panel of Hi-C datasets to assist topological structure annotation from a given study sample.
- Yanlin Zhang
- & Mathieu Blanchette
-
Article
| Open Access Structural basis of RNA polymerase II transcription on the chromatosome containing linker histone H1
Structural basis of RNA polymerase II transcription on the chromatosome containing linker histone H1
The mechanism by which RNAPII transcribes the DNA in the chromatosome with H1 has remained enigmatic. Here the authors present the cryo-EM structures of the RNAPII-chromatosome complexes, and explain how RNAPII is regulated by H1 in chromatin.
- Rina Hirano
- , Haruhiko Ehara
- & Hitoshi Kurumizaka
-
Article
| Open Access Hi-TrAC reveals division of labor of transcription factors in organizing chromatin loops
Hi-TrAC reveals division of labor of transcription factors in organizing chromatin loops
It is currently not clear how architectural proteins orchestrate chromatin looping at different scales of genome organisation. Here the authors report Hi-TrAC as a proximity ligation-free method to profile genome-wide chromatin interactions at single nucleosome resolution among regulatory elements.
- Shuai Liu
- , Yaqiang Cao
- & Keji Zhao
-
Article
| Open Access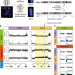 Human centromere repositioning activates transcription and opens chromatin fibre structure
Human centromere repositioning activates transcription and opens chromatin fibre structure
In this study, using a human neocentromere as a model, the authors show that centromeres have a special chromatin structure. Centromere repositioning triggers transcriptional activation, epigenetic remodelling and chromatin fibre decompaction.
- Catherine Naughton
- , Covadonga Huidobro
- & Nick Gilbert
-
Article
| Open Access Deciphering multi-way interactions in the human genome
Deciphering multi-way interactions in the human genome
Mapping higher order chromatin architecture is important. Here the authors use long sequencing reads to map genome-wide multi-way contacts and investigate higher order chromatin organisation; they use hypergraph theory for data representation and analysis, and apply this to different cell types.
- Gabrielle A. Dotson
- , Can Chen
- & Indika Rajapakse
-
Article
| Open Access Multimodal single cell sequencing implicates chromatin accessibility and genetic background in diabetic kidney disease progression
Multimodal single cell sequencing implicates chromatin accessibility and genetic background in diabetic kidney disease progression
Diabetic kidney disease leads to changes in glucose metabolism and inflammation. Here the authors use multimodal single cell sequencing to show that this disease leads to reduced accessibility of glucocorticoid receptor binding sites in the proximal tubule and increased gluconeogenesis.
- Parker C. Wilson
- , Yoshiharu Muto
- & Benjamin D. Humphreys
-
Article
| Open Access NODULIN HOMEOBOX is required for heterochromatin homeostasis in Arabidopsis
NODULIN HOMEOBOX is required for heterochromatin homeostasis in Arabidopsis
Arabidopsis NDX was previously reported as a regulator of euchromatic gene expression. Here the authors show that NDX functions at pericentromeric regions and regulates heterochromatin homeostasis by controlling siRNA production and non-CG methylation.
- Zsolt Karányi
- , Ágnes Mosolygó-L
- & Lóránt Székvölgyi
-
Article
| Open Access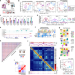 Postmitotic differentiation of human monocytes requires cohesin-structured chromatin
Postmitotic differentiation of human monocytes requires cohesin-structured chromatin
How chromatin structure and gene accessibility changes during monocyte differentiation is not clearly defined. Here the authors characterize the chromatin changes during macrophage or dendritic cell maturation from monocytes and the dependence of this upon cohesin and CTCF.
- Julia Minderjahn
- , Alexander Fischer
- & Michael Rehli
-
Article
| Open Access Loop-extrusion and polymer phase-separation can co-exist at the single-molecule level to shape chromatin folding
Loop-extrusion and polymer phase-separation can co-exist at the single-molecule level to shape chromatin folding
Two main mechanisms have been proposed to shape 3D genome architecture - loop extrusion and phase separation. Here the authors combine these mechanisms in polymer models in a manner that best fits 3D genome, based on both Hi-C and super-resolution locus imaging data, proposing that these two physical processes can indeed coexist simultaneously within cells to define loops and TADs.
- Mattia Conte
- , Ehsan Irani
- & Mario Nicodemi
-
Article
| Open Access Large-scale multi-omics analysis suggests specific roles for intragenic cohesin in transcriptional regulation
Large-scale multi-omics analysis suggests specific roles for intragenic cohesin in transcriptional regulation
Cohesin complex is essential for transcriptional regulation and chromosome folding. Here the authors provide a large-scale multi-omics analysis suggesting that cohesin bound on gene bodies has a specific role in transcriptional regulation.
- Jiankang Wang
- , Masashige Bando
- & Ryuichiro Nakato
-
Article
| Open Access JMJD3 intrinsically disordered region links the 3D-genome structure to TGFβ-dependent transcription activation
JMJD3 intrinsically disordered region links the 3D-genome structure to TGFβ-dependent transcription activation
Here the authors demonstrate that TGFβ drives multi-enhancer contacts and ultimately gene activation during neuronal commitment, and that this requires the intrinsically disordered region (IDR) of the histone demethylase JMJD3 likely through its role in promoting phase-separated biomolecular condensates.
- Marta Vicioso-Mantis
- , Raquel Fueyo
- & Marian A. Martínez-Balbás
-
Article
| Open Access Reconstruct high-resolution 3D genome structures for diverse cell-types using FLAMINGO
Reconstruct high-resolution 3D genome structures for diverse cell-types using FLAMINGO
High-resolution reconstruction of spatial chromosome organisation is in demand. Here the authors report FLAMINGO, for reconstructing high-resolution 3D Genome Organisation from HiC data which they use to generate both 5 kb and 1 kb-resolution 3D chromosomal structures for the human genome.
- Hao Wang
- , Jiaxin Yang
- & Jianrong Wang
-
Article
| Open Access Analysis of sub-kilobase chromatin topology reveals nano-scale regulatory interactions with variable dependence on cohesin and CTCF
Analysis of sub-kilobase chromatin topology reveals nano-scale regulatory interactions with variable dependence on cohesin and CTCF
Chromosome conformation capture (3 C) techniques have captured largescale 3D genome architecture. Here the authors present their “Tiled-MCC” approach for generation of 3 C data across megabase-scale loci at very high (up to 20 bp) resolution, which allowed them to observe nano-scale chromatin structures and investigate how these structures depend on cohesin and CTCF.
- Abrar Aljahani
- , Peng Hua
- & A. Marieke Oudelaar
-
Article
| Open Access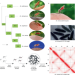 Anopheles mosquitoes reveal new principles of 3D genome organization in insects
Anopheles mosquitoes reveal new principles of 3D genome organization in insects
Anopheles mosquitoes are vectors of human malaria, and better understanding of them has implications for public health. Here, the authors apply Hi-C, FISH, RNA-seq, and ChIP-seq techniques to comprehensively characterize chromatin architecture and its evolutionary dynamics in five Anopheles species.
- Varvara Lukyanchikova
- , Miroslav Nuriddinov
- & Veniamin Fishman
-
Article
| Open Access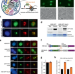 Genome-wide maps of nucleolus interactions reveal distinct layers of repressive chromatin domains
Genome-wide maps of nucleolus interactions reveal distinct layers of repressive chromatin domains
Mapping nucleolar-associated domains (NADs) is challenging as the nucleolus is membrane-less. Here the authors adapted the DNA adenine methyltransferase identification (DamID) method by engineering a “nucleolar histone” that can mark genomic contacts with the nucleolus through DNA methylation (m6A) and characterized NADs in mouse embryonic stem cells (ESCs) and derived neural progenitors (NPCs).
- Cristiana Bersaglieri
- , Jelena Kresoja-Rakic
- & Raffaella Santoro
-
Article
| Open Access Dynamic Runx1 chromatin boundaries affect gene expression in hematopoietic development
Dynamic Runx1 chromatin boundaries affect gene expression in hematopoietic development
RUNX1 is a large and complex gene with two alternative promoters and multiple hematopoietic enhancers. Here the authors show that unlike smaller genes Runx1 becomes sub-compartmentalized during differentiation and gene activation. This subcompartmentalization partially depends on CTCF binding at promoter-proximal CTCF sites and transcription.
- Dominic D. G. Owens
- , Giorgio Anselmi
- & Marella F. T. R. de Bruijn
-
Article
| Open Access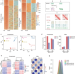 MyoD is a 3D genome structure organizer for muscle cell identity
MyoD is a 3D genome structure organizer for muscle cell identity
Pioneer transcription factors (TFs) have been proposed to act as protein anchors to orchestrate cell type-specific 3D genome architecture. MyoD is a pioneer TF for myogenic lineage specification. Here the authors provide further support for the role of MyoD in 3D genome architecture in muscle stem cells by comparing MyoD knockout and wild-type mice.
- Ruiting Wang
- , Fengling Chen
- & Dahai Zhu
-
Article
| Open Access Heterochromatin is a quantitative trait associated with spontaneous epiallele formation
Heterochromatin is a quantitative trait associated with spontaneous epiallele formation
The molecular origins of epialleles remain unknown. Here, the authors show that a positive feedback loop between H3K9me2 and CMT2/3 is a major contributing factor to the origins of spontaneous epialleles and that heterochromatin is a quantitative trait associated with spontaneous epiallele formation.
- Yinwen Zhang
- , Hosung Jang
- & Robert J. Schmitz

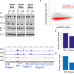 Acute depletion of BRG1 reveals its primary function as an activator of transcription
Acute depletion of BRG1 reveals its primary function as an activator of transcription