Abstract
Myriad policy, ethical and legal considerations underpin the sharing of biological resources, implying the need for standardised and yet flexible ways to digitally represent diverse ‘use conditions’. We report a core lexicon of terms that are atomic, non-directional ‘concepts of use’, called Common Conditions of use Elements. This work engaged biobanks and registries relevant to the European Joint Programme for Rare Diseases and aimed to produce a lexicon that would have generalised utility. Seventy-six concepts were initially identified from diverse real-world settings, and via iterative rounds of deliberation and user-testing these were optimised and condensed down to 20 items. To validate utility, support software and training information was provided to biobanks and registries who were asked to create Sharing Policy Profiles. This succeeded and involved adding standardised directionality and scope annotations to the employed terms. The addition of free-text parameters was also explored. The approach is now being adopted by several real-world projects, enabling this standard to evolve progressively into a universal basis for representing and managing conditions of use.
Similar content being viewed by others
Introduction
There is a widespread desire to maximise the use and re-use of biomedical data and samples. This ambition is, however, plagued by many complex and diverse issues pertaining to information governance. To be conducted in a responsible manner, the sharing and access of data and samples must take account of many constraints, not least individual consent, requirements set by custodians (researchers, institutions) and funders, policies set by ethics committees, and legal considerations. Beyond straightforward cataloguing of shareable datasets in archives, such as the National Centre for Biotechnology Information’s database of Genotypes and Phenotypes (dbGaP) and the European Genome-phenome Archive (EGA), the wider and more challenging world of data governance operates on an inconsistent and poorly structured basis.
Numerous general policy documents and guidelines have been created in recent years, which do a great job of establishing general principles for data and sample sharing. But these resources typically fail to provide a sufficiently fine-grained and concrete basis for operationalising the recommended principles. Custodian organisations currently have not universally adopted a standardised or consistent way to structure or express the wide array of information that might be part of their sharing and access policies. Consequently, the field is challenged by considerable diversity regarding consent form design and content, data sharing agreement clauses, choices over licensing models, institutional data sharing rules, and data management plans. This is not to say that specific elements of such tools should always or often be identical, as there will always be a need for context-specific variation on such matters. Nevertheless, more consistency and standardisation are needed at the level of specific concept structuring. That is, below the overarching general policy level, and above the detailed clause and textual level, there needs to be more uniformity and agreement over what would be a useful (non-redundant, unambiguous, sufficiently comprehensive) list of governance considerations and parameters.
Creating a fully comprehensive and universally applicable ‘conditions of use’ concept list for the whole biomedical domain would be unrealistic. But to make progress one could break down this semantic challenge to address distinct governance use cases, and then concentrate on the most widely used/needed concepts and parameters. This would bring substantial and rapid impact. The most significant previous efforts in this direction would be the Consent Codes list of permitted and conditional forms of data use1, and the further development and improvement of this as represented by the Data Use Ontology2 (DUO). These approaches focus primarily on capturing headline conditions for acceptable sharing of genomics datasets and reflect the concepts and combinations of concepts that emerged organically from the field. DUO and similar ontologies are intended to capture common permissions inherent in biomedical research data. Ontologies of this nature are especially useful where research consortia prospectively design informed consent materials and data governance strategies to be compatible with selected ontology terms. However, there remains a demonstrated need for flexible systems that can capture complex and conditional permissions in data, in a manner that enables logical computer-based reasoning. This class of systems may be necessary to represent and to compare permissions inherent in numerous categories of datasets, such as legacy datasets for which data governance rules have already been generated, or those that are subject to regulatory requirements that can only be described using contingent and conditional language.
Another related initiative is the “Automatable Discovery and Access Matrix” (ADA-M)3. This approach emphasised the design of a data structure into which conditions of use information could be placed. To that end it included a list of concepts of use, but these were largely based on the Consent Codes and DUO terms, and these were not rigorously validated by community testing. In addition, much of the substantive information conveyed in an ADA-M profile regarding permissible and impermissible data uses is articulated through its free text fields. Consequently, ADA-M cannot enable the machine-actionable discovery of data according to its data governance conditions.
Given the above, and with the European Joint Programme for Rare Diseases (EJP-RD) stated aim: “…. to create an effective rare diseases research ecosystem for progress, innovation and for the benefit of everyone with a rare disease. We support rare diseases stakeholders by funding research, bringing together data resources & tools, providing dedicated training courses, and translating high quality research into effective treatments” (https://www.ejprarediseases.org/), we worked to devise a rational lexicon for “conditions of use” information. Our aim was to meet the need within the programme to convey the use conditions associated with the various bioresources, in a convenient and automatable way. Further, it should be easy to deploy requiring only minimal training, with profiles that are human readable as well as being able to be represented in a computer readable format. To this end the resultant system needed to be able to encode use conditions that were applicable to data and biosamples.
The resulting design is called “Common Conditions of use Elements (CCE)”, representing a highly validated set of atomic (non-overlapping) concepts of use that offer a robust and consistent basis to many data governance tools and activities. CCE is explicitly not seeking to be an ontology (though it could inform future extensions to existing ontologies), but to be a categorisation framework for conditions of use concepts. As a version 1.0 product, the hope is that various groups in different settings will adopt and provide feedback on its utility, enabling it to be progressively improved.
Results
To produce a useful CCE list, efforts started on the domain of rare disease related registries, biobanks, and data repositories - especially as related to European Reference Networks (ERNs)4.
We identified four principles that are necessary for the construction of the CCE concepts:
-
(1)
be atomic, i.e., represent an operationally pure and singular concept, (elemental).
-
(2)
have no directionality, i.e., not convey any indication about whether that mode of use is allowed, forbidden, or obligatory, (neutrality).
-
(3)
be generalised, i.e., be a modular category of use that states no customisation details, conditionalities or dependencies, (generic).
-
(4)
be widely applicable and relevant, (relevance).
By way of example, consider a potential CCE category such as “Regulatory Jurisdiction”. This meets design principle #1 because it is not conflated with any other concept, such as “Geographical Area”. It meets design principle #2 because it does not attempt to covey whether data/sample use is permitted or forbidden in certain Regulatory Jurisdictions. It meets design principle #3 because it does not name any specific Regulatory Jurisdictions or imply the presence or absence of any modifying considerations (e.g., the degree of anonymity of the data). And it meets design principle #4 because it is a widely relevant consideration. Hence, it is simply a pure, generalised, categorisation concept pertaining to conditions of use of some un-named artifact.
The initial set of suggestions for CCE list items contained 76 preliminary concepts, produced after the independent extraction of all possible concepts from a series of domain consent forms, and material/Data transfer agreements (MTAs/DTAs) and after the elimination of duplicates. Each of the 76 concepts was characterised as being primarily a ‘criterion’ or ‘boundary conditions’ of use (‘who’, ‘what’, ‘why’, ‘when’, ‘where’ – 46 such elements), versus mainly a ‘process’ or ‘method’ of use (‘how’ – 19 such elements), or others not fitting into either group and so not further evaluated as a possible CCE item (11 such elements). After this categorisation, we debated and refined the list, according to the four principles above.
An overview of the analysis process, with the initial set of CCE terms can be found at the supplementary Table 1, that also summarises the process of refinement and criteria.
Directional items were made non-directional. For example, the Opt out form for Genomics England has the concept of “Patient has requested NOT to receive additional findings” in part 7B,was deemed to have the core concept of “return of incidental findings”. This concept can then be forbidden using an appropriate rule. Similarly, to meet the requirement of being atomic non-atomic items were split into their component parts. As an example, one of the ERN registries forms, (that relate to how the resources can be used) has the use condition of “recontacting the participant to return results or incidental findings” within the conditions that a participant must agree to. This term was split into the 3 CCE terms: (Re-)Identification, Return Of Results and Return Of Incidental Findings.
To make the resulting system more broadly applicable, the group also considered that some of the resources may condition use upon (i) the commercial or non-commercial character of the entities that use the data, and (ii) whether or not the intended use is profit-motivated. These concepts are typically conflated and or one is omitted (with conflation being implied/inferred) in alternative approaches.
The resulting 14 core concepts from this review and refinement process, were given a definition and a priority ranking based on anticipated utility and degree of relevance to the targeted use case (i.e., factors relevant to the secondary user of data or samples).
Based on community suggestions we added in 4 extra concepts that had not been surfaced by the starting set of consent forms and MTAs. Two of the original CCE terms were interpreted in different ways by the testers (Clinical Use and (Re-)Identification) indicating that they were not truly atomic in nature. Hence, these terms were split as discussed below, which resulted in 6 new terms in total.
By rounds of alpha-testing and consultation with the intended user community we progressively distilled the list down to what was felt to be a practically useful and usable size. The final version 1.0 CCE list comprises 20 items, as presented in Table 1 (criterion type terms) and Table 2 (process type terms). The modifications to the initial list of CCE terms are stated in the subsequent paragraph.
Some considerations that went into producing the full list of CCEs in consultation with the alpha test group, were as follows:
CCE Terms: Geographical area and regulatory jurisdiction
Initially only the term geographical area was proposed, as it was thought that this could encompass both concepts, however with jurisdictions such as the EU spanning complex geographies, we concluded that to maintain the atomic nature of CCE these two concepts needed to be separated.
CCE Terms: Clinical care use and clinical research use
The initially suggested term of “Clinical Use” was highlighted by the alpha testers as being too vague, in that it covered both clinical research use AND clinical care of the patient. This led to the splitting of the term into “Clinical Research Use” and “Clinical Care Use”.
CCE Terms: (Re-)Identification of individuals without the involvement of the resource provider and (Re-)identification of individuals mediated by the resource provider
A single (Re-)Identification term was initially introduced to cover any activity where a resource was used in a manner that allowed the reversing of anonymisation/pseudonymisation. It was assumed that this would always be forbidden, and “safeguards” would apply during use to prevent this occurring. However, we found that alpha testers interpreted this term in two distinctly diverse ways. One was to forbid the re-identification of participants as expected, while the other related to ways that the participant could be re-contacted (via the supplying institution) for some legitimate purpose such as recruitment to further studies. Hence the two CCE terms were devised to capture these two separate concepts of use.
Terms added based on feedback from alpha testers
The following CCE terms were added to cover the full range of concepts deemed important by alpha testers: Publication Moratorium, Publication, User Authentication, and Ethics Approval.
Validation of the final 20 cce terms by creation of regularised sharing/access policies
As a final validation exercise for CCEs against our intended use case, four biobanks, three patient registries and a data platform were asked to try to use the CCE lexicon as a basis for structuring an overview (called a Policy Profile) of their main sharing policy items. Guidance was prepared to support this task, in which it was made very clear that the goal was:
-
To base these Policy Profiles upon the 20 CCE concepts, each of which could be used zero or more times according to what they wanted to express in their policy.
-
To represent only those Policy Profile items applicable to the secondary user of the data or samples, not those that apply to the custodian biobank/registry themselves.
-
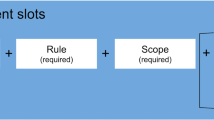
To accomplish the exercise by merely adding a directionality statement (“Permitted”, “Obligated”, “Forbidden” or ‘‘No Requirements’’) and a “scope” indicator (“Whole of resource” or “Part of resource”) to the employed CCE items. This approach leverages the Digital Use Conditions (DUC)5 Schema for expressing conditions of use information which was recently developed by an IRDiRC Task Force. The DUC schema employs the term “asset” to refer to both a collection of items or a singular item, being made available for some form of use. In contrast, since the CCEs in this current work are designed to support the creation of Policy Profiles for whole organisational structures (e.g., biobanks or RD registries) we instead use the term “resource”.
Supplementary Table 2 contains the feedback from validators on all CCE Terms. Overall, the process was very straightforward, with two major take-home lessons emerging. First, organisations can have very different objectives in mind when creating such Policies Profiles – ranging from wanting to be quite comprehensive in stating what can and cannot be done with their data or samples, through to others that merely wanted to provide a minimal list of headline “showstopper” categories of use that could never be allowed. Second, most groups were rather cautious about any public release or exposure of their Policy Profiles (we even had to adapt our support tools because of this concern, so that no profile was ever left on a public server) - the implications of which probably merit further investigation. Nevertheless, despite the diversity of objectives, CCEs were able to accommodate all these approaches. Additionally, there was no request for any additional CCE concepts to be added.
Policy Profiles such as these demonstrate how well CCEs can provide the basis for a consistently clear and structured representation of access policies across many organisations. However, they may not provide sufficient details for some more-demanding applications (e.g., Data Access Committee deliberations). Therefore, as one last final test of CCEs, the alpha-tester organisations kindly generated some additional free-text parameters to supplement and elaborate a number of the term + rule + scope triads previously provided in their Policy Profiles. These have been collated into a set, that gives an example for each CCE term from the free texts (Table 3 for criterion type terms and Table 4 for Process type terms) where provided.
Discussion
In this work we have developed, validated, and recommend the Common Conditions of use Elements (CCEs): a set of conditions of use concepts that are atomic, non-directional, and which should be widely useful as a foundational layer in support of many aspects of data and sample sharing. The CCE list was devised by extensive discussion and alpha testing with many community members, followed by real-world testing in terms of creating general and detailed Policy Profiles. This latter work employed online software to utilise CCEs in the context of the DUC Schema for structuring information governance metadata.
We took a different approach to previous initiatives. Consent Codes aimed to provide a set of controlled terms that could be used to represent participant consent for uses in specific types of research. DUO expanded this to include general use conditions but focused on data only. Further the designers of DUO constructed it as a series of interconnected terms that formed a formal ontology. This means that care must be taken to ensure that a desirable term is present in the correct context within the DUO ontology.
ADA-M took an approach wherein each term was specific and separate. The ADA-M terms were split into two main categories: those that related to “Permissions” for specific uses and those that related to “Terms” of use. Like the current work these were paired with a rule base that either permitted or forbid the specific permission term or stating whether a given Term was true or not. CCEs (when used within a rule base such as the Digital Use Conditions DUC Schema) seek to simplify the approach taken by ADA-M. The idea of the two classes of terms is maintained but the rule base is simplified, and an optional free text field is added to allow elaboration of the full statement (CCE term, rule and scope) as required. While the free text is not currently able to be used as a filter, it does provide context to the end user when selected profiles are read.
The directional neutrality of CCEs ensures that uses covered by individual CCE terms could be permitted as well as forbidden using the appropriate rules when used in a data structure such as the DUC schema. While ADA-M had previously tried to take a similar approach, its’ complexity made it difficult to use, hampering its adoption. CCEs aimed to avoid this complexity by using simple independent atomic terms. The terms were also explained in plain English, with the aim of increasing their adoption by making them more accessible than some of the alternative approaches taken in this area.
Clearly, a series of such CCEs based on these principles would not, in and of themselves, be sufficient to describe a complete sharing or access policy. But that is not the objective of CCEs. Instead, the aim is to establish a widely relevant standardised set of atomic, non-directional, generic conditions of use concepts. These can then be employed as the basis for designing forms, contracts, tools and infrastructures that would be more innately inter-operable, and one would only need to elaborate the CCEs with directionality and other specifics to produce data sharing policies etc., that would all be structurally very similar and hence compatible and comparable.
Work is now underway to use the CCE model within the EJP-RD project, in particular to use them to gather Policy Profiles (that provide a summary of the use conditions, in a similar fashion to how the nutritional information for food is displayed in Europe) for many registries, biobanks and other online resources, and use these metadata to underpin data and sample discovery services and sharing activities. Additionally, other international initiatives have begun exploring ways to employ the CCE concept, not least BBMRI-ERIC6, GA4GH, the IMI EPND project7, and the FAIR community8. Some of this work entails extending the list of CCEs to suit specialised use cases, such as dynamic consent and support for GDPR considerations9,10. In the case of the FAIR community, CCE concepts are being tested for compatibility with semantic web models, such as Open Digital Rights Language (ODRL)11, to afford increased machine-readability. We anticipate versioning CCEs in a public manner, as these real-world activities progress. As this occurs, increasingly validated CCE concepts will ideally become included in formal ontologies, such as DUO or the Informed Consent Ontology12 (ICO). Table 5 illustrates the current overlap between CCEs (version 1.0) and DUO, in order to highlight gaps that could usefully be filled. All CCE terms are shown, along with their direct or indirect mapping to DUO.
Table 5 illustrates that whilst some DUO terms reflect composite data use conditions that bring together multiple related conditions of use as one ontology term, CCE would express similar conditions through the combination of multiple separate atomic terms. For example, in DUO, use by a commercial entity is commingled with for-profit use; jurisdiction and geographical location are likewise linked together. Further, select DUO terms are explicitly directional, and are intended to indicate that a certain behaviour is obligated in the use of data. Examples thereof include “collaboration required” and “time limit on use.” Conversely, CCEs would express these terms without implying directionality, which would enable users thereof to indicate the presence, absence, or explicit preclusion of each condition. The use of a lexicon composed of atomic and non-directional terms can enable communities to express the full range of permissions in their data using one common system. This can be leveraged to help communities with case-specific data governance needs to develop and tailor bespoke systems that are suitable to their needs, building upon a wider library of CCE terms. It also could enable organisations to use CCEs to enable interoperability across distinct, context-specific ontologies, allowing researchers to leverage context-specific ontologies of their choosing, whilst still enabling for the interoperable comparison of data governance conditions that have been expressed using multiple distinct ontologies, absent prior coordination.
Methods
The system needed to be suitable for use by all the project partners involved, including those from research and healthcare sectors, as well as non-profit organisations. Given the broad range of activities undertaken by the various partners, the resulting use conditions needed to consider use cases outside of the traditional research community in which other systems had been developed.
Defining CCE terms
CCE development work was undertaken within the project team and in conjunction with members of the rare disease community, to produce the version 1.0 model. This work involved many rounds of iterative testing and improvement of items in the CCE list and drafting and refining definitions for each. A reductionist approach was taken to devising the CCE terms.
This started with the capturing of concepts from various types of documents including Informed Consent Forms (ICF); data access policies (DAP); data/material transfer agreements (DTA/MTA) that were either accessible via the EJP-RD project (e.g., ICFs from ERNs and Biobanking community),or publicly available consents, DTA, and MTA, namely the Manchester Tissue Bank Material Transfer agreement; the UKRI consent form; Genomic England Cancer Research Consent Form; Cancer UK generic systemic anti-cancer treatment consent form; the Cancer research UK Immunotherapy consent form; the UKRI generic consent template; the UK Data services consent; the BMA/Law society - Consent template and Genomics England opt out ADDITONAL FINDINGS Q7 as well as ICF from the ERN and Biobanking community. Concepts were extracted and assessed against the CCE criteria detailed in Results. When terms did not meet these criteria, they were initially reviewed to determine if they could be adapted to meet the criteria (i.e., by breaking down into simpler atomic concepts and or removing the directionality), or otherwise rejected. The list was also checked to ensure that it contained elements that were identified as being of importance to rare disease patients based on a survey by McCormack et al.13. These concepts are listed in the Supplementary Table 1 together with the CCE terms to which they map to or the reason why they were not considered for integration in the final set of CCE terms.
In this way, we identified and optimised many facets of the CCE design, not least: revealing and addressing aspects that were likely to create misunderstanding; highlighting elements that needed to be split further into truly atomic concepts; bringing forward suggestions of frequently-needed concepts that were missing from the evolving CCE list; and, guiding subjective decisions about what was sufficiently ‘common’ and hence useful to go into a version 1.0 CCE list, versus what concepts were too specialised or infrequently used.
Progress was facilitated by means of a shared Excel document, and the use and stepwise development of custom software that included help texts and instructional videos, combined with regular feedback discussions with testers.
Using CCE terms to produce CCE statements using the DUC schema and alpha testers assessment of their utility
The final set of CCE terms were evaluated for their utility by employing them as “condition terms” in the DUC schema. This schema allows each CCE term to be converted to a “statement” by the addition of a suitable rule (from DUC’s set of Obligatory, Permitted, Forbidden or No Requirement). To complete a CCE statement a “scope” of the rule was assigned specifying whether the statement applied to the “whole of the resource” or “part of the resource”.
Web based tool for constructing DUC profiles using CCE statements
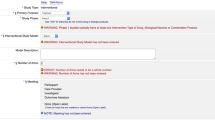
An online tool was developed14 (https://ducejprd.le.ac.uk) that includes a web based “wizard” interface that enabled alpha testers to select CCEs, and then enter Rules and Scope values. Alpha testers used the tool to make their resource level Policy Profiles. The tool also includes sections to provide details of the resource to which the use conditions apply to. Users could add as many or as few CCE statements to a profile as they wish. CCE statements could reuse or omit CCE terms as needed. CCE statements are independent of each other, and so the tool does not enable users to enter inter-statement dependencies. Profiles were reviewed, and their meaning was clarified with the alpha testers where required.
Data availability
The original datasets were provided in confidence and to respect that confidentiality, we show the generated profiles as aggregated data in the manuscript (supplementary Table 2).
Code availability
The code for the web-based tool for construction of a DUC/CCE profile (https://ducejprd.le.ac.uk) can be found at this public repository https://github.com/Cafe-Variome/DucCCE with read only access.
Change history
15 May 2024
The ORCiD for Maria del Carmen Sanchez Gonzalez in the PDF version of this article has been amended; the HTML version was correct at the time of publication.
References
Dyke, S. O. M. et al. Consent Codes: Upholding Standard Data Use Conditions. Plos Genet 12, e1005772 6 (2016).
Lawson, J. et al. The Data Use Ontology to streamline responsible access to human biomedical datasets. Cell Genom 1, 1–9 (2021).
Woolley, J. P. et al. Responsible sharing of biomedical data and biospecimens via the “Automatable Discovery and Access Matrix” (ADA-M). Npj Genom Medicine 3, 1–6 (2018).
Tumiene, B. et al. European Reference Networks: challenges and opportunities. J Community Genet. 12(2), 217–229 (2021).
Jeanson, F. et al. Getting your DUCs in a row - standardising the representation of Digital Use Conditions. Sci. Data https://doi.org/10.1038/s41597-024-03280-6 (2024).
Holub, P. et al. BBMRI-ERIC Directory: 515 Biobanks with Over 60 Million Biological Samples. Biopreserv Biobank. 14(6), 559–562 (2016).
European Platform for Neurodegenerative Diseases. https://epnd.org (2023).
Wilkinson, M. D. et al. The FAIR Guiding Principles for scientific data management and stewardship. Sci Data. 3, 160018 (2016).
Teare, H. J. A., Prictor, M. & Kaye, J. Reflections on dynamic consent in biomedical research: the story so far. Eur. J. Hum. Genet. 29, 649–656 (2021).
Mondschein, C.F. & Monda, C. The EU’s General Data Protection Regulation (GDPR) in a Research Context. In Fundamentals of Clinical Data Science (eds Kubben, P., Dumontier, M., & Dekker, A.) 55–71, https://doi.org/10.1007/978-3-319-99713-1_5 (Springer, Boston, MA, 2019).
Iannella, R. The Open Digital Rights Language: XML for Digital Rights Management. Information Secur. Technical Rep. 9, 47–55 (2004).
Lin, Y. et al. Development of a BFO-based Informed Consent Ontology (ICO). Proceedings of the International Conference on Biomedical Ontology (2014).
McCormack, P. et al. ‘You should at least ask’. The expectations, hopes and fears of rare disease patients on large-scale data and biomaterial sharing for genomics research. Eur J Hum Genet 24, 1403–1408 (2016).
Riaz, U., Veal, C. D., Gibson, S. J., Maini, M. & Brookes, A. J. DUC Profile Creator Using CCEs. https://ducejprd.le.ac.uk (2023).
Acknowledgements
The authors wish to thank Lotte Boormans (ERN eUROGEN) and Nawel Lalout (World Duchenne Organization) for piloting CCE terms along with DUC software. We also thank the developers of the Digital Use Conditions (DUC) structure and members of the IRDiRC ‘Machine Readable Consent and Use Conditions’ Task Force (https://irdirc.org/machine-readable-consent-and-use-conditions/) for providing project oversight and utility testing of CCEs. Finally, we acknowledge and thank the ‘European Joint Programme on Rare Diseases’ for funding this work as part of the EU Horizon 2020 programme, Grant Agreement N°825575, which contributed to the development work, supported publication costs, and resourced the IRDiRC Scientific Secretariat which is hosted at INSERM in Paris, France.
Author information
Authors and Affiliations
Contributions
Maria del Carmen Sanchez Gonzalez: Original CCE Design and refinement (based on tester feedback), review and clarification of alpha testers profiles, manuscript preparation, suggestions for the Digital Use Condition schema based on review of tester feedback. Pim Kamerling: Original CCE design and refinement (based on tester feedback), manuscript preparation, suggestions for the Digital Use Condition schema based on review of tester feedback, liaison with the ERKReg community testers (primary contact) via his position in VASCERN the European Reference Network for Rare Multisystemic Vascular Diseases, who’s time was provided as a good will gesture to progress the work described. Mariapia Iermito: Original CCE design. Sara Casati: Original CCE design, BBMRI-Eric CCE advocate, and liaison with the BBMRI-Eric community testers (primary contact). Umar Riaz: Design and coding of the DUC/CCE web-based profile generation tool. Colin D. Veal: CCE refinement (based on tester feedback), design of the DUC/CCE web-based profile generation tool and suggestions for the Digital Use Condition schema based on review of tester feedback. Monika Maini: CCE refinement (based on tester feedback) and design of the DUC/CCE web-based profile generation tool. Francis Jeanson: DUC lead developer and liaison for the CCE integration including adaption of DUC schema and rule base to accommodate the CCE application. Oussama Mohammed Benhamed: Assessment of the Open Digital Rights Language (ODRL) for use with CCE terms to produce a use conditions profile for a bioresource. Esther van Enckevort: Acquisition of project’s grant funding, development, and coding of the CCE profile generation tool using the Molgenis platform. Annalisa Landi: Manuscript preparation and assessment of CCE terms for their utility to represent responses to questions in the EJP-RD integrated consent form. Yanis Mimouni: Assessment of CCE terms for their utility to represent responses to questions in the EJP-RD integrated consent form and testing of the web-based DUC/CCE profile generation tool to construct a use conditions profile for their resource. Clèmence Le Cornec: Testing of the web-based DUC/CCE profile generation tool to construct a use conditions profile for their resource and engagement with the CCE design team to explain and or clarify their choice of CCE terms and statement construction. Domenico A. Coviello: Testing of the web-based DUC/CCE profile generation tool to construct a use conditions profile for their resource and engagement with the CCE design team to explain and or clarify their choice of CCE terms and statement construction. Tiziana Franchin: Testing of the web-based DUC/CCE profile generation tool to construct a use conditions profile for their resource and engagement with the CCE design team to explain and or clarify their choice of CCE terms and statement construction. Francesca Fusco: Testing of the web-based DUC/CCE profile generation tool to construct a use conditions profile for their resource and engagement with the CCE design team to explain and or clarify their choice of CCE terms and statement construction. Jose Antonio Ramírez García: Testing of the web-based DUC/CCE profile generation tool to construct a use conditions profile for their resource and engagement with the CCE design team to explain and or clarify their choice of CCE terms and statement construction. Loes F.M. van der Zanden: Testing of the web-based DUC/CCE profile generation tool to construct a use conditions profile for their resource and engagement with the CCE design team to explain and or clarify their choice of CCE terms and statement construction. Alexander Bernier: Adaption of DUC schema and rule base to accommodate CCE application. Mark D. Wilkinson: Assessment of the Open Digital Rights Language (ODRL) for use with CCE terms to produce a use conditions profile for a bioresource. Heimo Mueller: BBMRI Eric CCE advocate and liaison. Spencer J. Gibson: Original CCE Design and refinement (based on tester feedback), review and clarification of alpha testers profiles, manuscript preparation, design of the DUC/CCE web-based profile generation tool and associated training material, suggestions for the Digital Use Condition schema based on review of tester feedback and general coordination of the work relating to the CCE work in the presented article. Anthony J. Brookes: Original CCE design and refinement (based on tester feedback), manuscript preparation and final (pre-submission) reviewer/editor, suggestions for the Digital Use Condition schema based on review of tester feedback, oversight of the work presented in this manuscript (as lead PI) and acquisition of project’s grant funding.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Sanchez Gonzalez, M.d.C., Kamerling, P., Iermito, M. et al. Common conditions of use elements. Atomic concepts for consistent and effective information governance. Sci Data 11, 465 (2024). https://doi.org/10.1038/s41597-024-03279-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41597-024-03279-z