Abstract
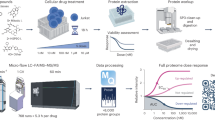
High-content cell profiling has proven invaluable for single-cell phenotyping in response to chemical perturbations. However, methods with improved throughput, information content and affordability are still needed. We present a new high-content spectral profiling method named vibrational painting (VIBRANT), integrating mid-infrared vibrational imaging, multiplexed vibrational probes and an optimized data analysis pipeline for measuring single-cell drug responses. Three infrared-active vibrational probes were designed to measure distinct essential metabolic activities in human cancer cells. More than 20,000 single-cell drug responses were collected, corresponding to 23 drug treatments. The resulting spectral profile is highly sensitive to phenotypic changes under drug perturbation. Using this property, we built a machine learning classifier to accurately predict drug mechanism of action at single-cell level with minimal batch effects. We further designed an algorithm to discover drug candidates with new mechanisms of action and evaluate drug combinations. Overall, VIBRANT has demonstrated great potential across multiple areas of phenotypic screening.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The raw FTIR imaging data generated in this work are available from the corresponding author on reasonable request. The processed single-cell FTIR spectrum data are available at https://github.com/MinLabColumbia/VIBRANT.
Code availability
The codes are available at https://github.com/MinLabColumbia/VIBRANT.
References
Kaddurah-Daouk, R., Kristal, B. S. & Weinshilboum, R. M. Metabolomics: a global biochemical approach to drug response and disease. Annu. Rev. Pharmacol. Toxicol. 48, 653–683 (2008).
Chandrasekaran, S. N., Ceulemans, H., Boyd, J. D. & Carpenter, A. E. Image-based profiling for drug discovery: due for a machine-learning upgrade? Nat. Rev. Drug Discov. 20, 145–159 (2021).
Wang, Y.-Y. et al. CeDR Atlas: a knowledgebase of cellular drug response. Nucleic Acids Res. 50, D1164–D1171 (2022).
Zhao, Y. et al. Measurement methods of single cell drug response. Talanta 239, 123035 (2022).
Fritzsch, F. S. O., Dusny, C., Frick, O. & Schmid, A. Single-cell analysis in biotechnology, systems biology, and biocatalysis. Annu. Rev. Chem. Biomol. Eng. 3, 129–155 (2012).
Walsh, A. J. et al. Quantitative optical imaging of primary tumor organoid metabolism predicts drug response in breast cancer. Cancer Res. 74, 5184–5194 (2014).
Loo, L.-H., Wu, L. F. & Altschuler, S. J. Image-based multivariate profiling of drug responses from single cells. Nat. Methods 4, 445–453 (2007).
Bray, M.-A. et al. Cell Painting, a high-content image-based assay for morphological profiling using multiplexed fluorescent dyes. Nat. Protoc. 11, 1757–1774 (2016).
Haghighi, M., Caicedo, J. C., Cimini, B. A., Carpenter, A. E. & Singh, S. High-dimensional gene expression and morphology profiles of cells across 28,000 genetic and chemical perturbations. Nat. Methods 19, 1550–1557 (2022).
Spitzer, M. H. & Nolan, G. P. Mass cytometry: single cells, many features. Cell 165, 780–791 (2016).
Bendall, S. C. et al. Single-cell mass cytometry of differential immune and drug responses across a human hematopoietic continuum. Science 332, 687–696 (2011).
Heath, J. R., Ribas, A. & Mischel, P. S. Single-cell analysis tools for drug discovery and development. Nat. Rev. Drug Discov. 15, 204–216 (2016).
Jamieson, L. E. & Byrne, H. J. Vibrational spectroscopy as a tool for studying drug-cell interaction: could high throughput vibrational spectroscopic screening improve drug development? Vib. Spectrosc. 91, 16–30 (2017).
Zhao, Z., Chen, C., Xiong, H., Ji, J. & Min, W. Metabolic activity phenotyping of single cells with multiplexed vibrational probes. Anal. Chem. 92, 9603–9612 (2020).
Tipping, W. J., Lee, M., Serrels, A., Brunton, V. G. & Hulme, A. N. Stimulated Raman scattering microscopy: an emerging tool for drug discovery. Chem. Soc. Rev. 45, 2075–2089 (2016).
Ma, J., Pazos, I. M., Zhang, W., Culik, R. M. & Gai, F. Site-specific infrared probes of proteins. Annu. Rev. Phys. Chem. 66, 357–377 (2015).
Mignolet, A. & Goormaghtigh, E. High throughput absorbance spectra of cancerous cells: a microscopic investigation of spectral artifacts. Analyst 140, 2393–2401 (2015).
Mignolet, A., Mathieu, V. & Goormaghtigh, E. HTS-FTIR spectroscopy allows the classification of polyphenols according to their differential effects on the MDA-MB-231 breast cancer cell line. Analyst 142, 1244–1257 (2017).
Flower, K. R. et al. Synchrotron FTIR analysis of drug treated ovarian A2780 cells: an ability to differentiate cell response to different drugs? Analyst 136, 498–507 (2011).
Denbigh, J. L. et al. Probing the action of a novel anti-leukaemic drug therapy at the single cell level using modern vibrational spectroscopy techniques. Sci. Rep. 7, 2649 (2017).
Shi, L. et al. Mid-infrared metabolic imaging with vibrational probes. Nat. Methods 17, 844–851 (2020).
Liu, X. et al. Towards mapping mouse metabolic tissue atlas by mid-infrared imaging with heavy water labeling. Adv. Sci. 9, e2105437 (2022).
Büttner, F. H. Cell-based assays for high-throughput screening. Expert Opin. Drug Discov. 1, 373–378 (2006).
Costa-Mattioli, M. & Walter, P. The integrated stress response: From mechanism to disease. Science 368, eaat5314 (2020).
Blay, V., Tolani, B., Ho, S. P. & Arkin, M. R. High-throughput screening: today’s biochemical and cell-based approaches. Drug Discov. Today 25, 1807–1821 (2020).
Stirling, D. R. et al. CellProfiler 4: improvements in speed, utility and usability. BMC Bioinf. 22, 433 (2021).
Perlman, Z. E. et al. Multidimensional drug profiling by automated microscopy. Science 306, 1194–1198 (2004).
Taymaz-Nikerel, H., Karabekmez, M. E., Eraslan, S. & Kırdar, B. Doxorubicin induces an extensive transcriptional and metabolic rewiring in yeast cells. Sci. Rep. 8, 13672 (2018).
Jamdade, V. S. et al. Therapeutic targets of triple-negative breast cancer: a review. Br. J. Pharmacol. 172, 4228–4237 (2015).
Kummar, S. et al. Advances in using PARP inhibitors to treat cancer. BMC Med. 10, 25 (2012).
Caulfield, S. E., Davis, C. C. & Byers, K. F. Olaparib: a novel therapy for metastatic breast cancer in patients with a BRCA1/2 mutation. J. Adv. Pract. Oncol. 10, 167–174 (2019).
Johannessen, C. M., Clemons, P. A. & Wagner, B. K. Integrating phenotypic small-molecule profiling and human genetics: the next phase in drug discovery. Trends Genet. 31, 16–23 (2015).
Moffat, J. G., Rudolph, J. & Bailey, D. Phenotypic screening in cancer drug discovery - past, present and future. Nat. Rev. Drug Discov. 13, 588–602 (2014).
Rana, S. et al. A multichannel nanosensor for instantaneous readout of cancer drug mechanisms. Nat. Nanotechnol. 10, 65–69 (2015).
Vincent, F. et al. Phenotypic drug discovery: recent successes, lessons learned and new directions. Nat. Rev. Drug Discov. 21, 899–914 (2022).
Liu, F. T., Ting, K. M. & Zhou, Z.-H. Isolation forest. In Proc. 2008 Eighth IEEE International Conference on Data Mining (eds Giannotti, F. et al.) (IEEE, 2008).
Liang, H. & Tan, A. R. Iniparib, a PARP1 inhibitor for the potential treatment of cancer, including triple-negative breast cancer. IDrugs 13, 646–656 (2010).
Mateo, J., Ong, M., Tan, D. S. P., Gonzalez, M. A. & de Bono, J. S. Appraising iniparib, the PARP inhibitor that never was—what must we learn? Nat. Rev. Clin. Oncol. 10, 688–696 (2013).
Bayat Mokhtari, R. et al. Combination therapy in combating cancer. Oncotarget 8, 38022–38043 (2017).
Chalakur-Ramireddy, N. K. R. & Pakala, S. B. Combined drug therapeutic strategies for the effective treatment of triple negative breast cancer. Biosci. Rep. 38, BSR20171357 (2018).
Ianevski, A., Giri, A. K. & Aittokallio, T. SynergyFinder 2.0: visual analytics of multi-drug combination synergies. Nucleic Acids Res. 48, W488–W493 (2020).
Seydel, C. Single-cell metabolomics hits its stride. Nat. Methods 18, 1452–1456 (2021).
Qian, W. W. et al. Batch equalization with a generative adversarial network. Bioinformatics 36, i875–i883 (2020).
Ando, D. M., McLean, C. Y. & Berndl, M. Improving phenotypic measurements in high-content imaging screens. Preprint at bioRxiv https://doi.org/10.1101/161422 (2017).
Yeh, K., Kenkel, S., Liu, J.-N. & Bhargava, R. Fast infrared chemical imaging with a quantum cascade laser. Anal. Chem. 87, 485–493 (2015).
Zhang, Y. et al. Single-shot volumetric chemical imaging by mid-infrared photothermal Fourier light field microscopy. Preprint at bioRxiv https://doi.org/10.1101/2022.06.24.497516 (2022).
Zhang, L., Zhang, Y. D., Zhao, P. & Huang, S.-M. Predicting drug-drug interactions: an FDA perspective. AAPS J. 11, 300–306 (2009).
Vlachogiannis, G. et al. Patient-derived organoids model treatment response of metastatic gastrointestinal cancers. Science 359, 920–926 (2018).
Hannon, G. J. RNA interference. Nature 418, 244–251 (2002).
Chen, S. et al. Genome-wide CRISPR screen in a mouse model of tumor growth and metastasis. Cell 160, 1246–1260 (2015).
Acknowledgements
W.M. acknowledges support from National Institutes of Health (grant no. R01 EB029523) and Chan Zuckerberg Initiative (Dynamic Imaging grant no. 2023-321166). We also thank L. Sun for help with the partial data analysis.
Author information
Authors and Affiliations
Contributions
X.L. performed the cell culture and drug treatment experiments, FTIR imaging collection and data analysis. L.S., Z.Z. and J.S. provided advice in experiments and data analysis. X.L. and W.M. conceived the concept and wrote the paper with input from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests. Columbia University has filed a provisional patent application based on this work.
Peer review
Peer review information
Nature Methods thanks Malgorzata Baranska, Erik Goormaghtigh and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available. Primary Handling Editor: Rita Strack, in collaboration with the Nature Methods team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Average single-cell IR spectrum of individual vibrational probe or control.
(a) azido-PA. (b) 13C-AA. (c) d34-OA. (d) control without any labeling.
Extended Data Fig. 2
Single-cell image segmentation of MDA-MB-231 cells using CellProfiler.
Extended Data Fig. 3 Batch effects evaluation of VIBRANT with multiplexed vibrational probes.
3D scatter plots on three metabolic activities of cells treated by drugs were used for visualization. (a) Dactolisib, (b) Doxorubicin, and (c) Triacsin-C were selected as representatives. It can be observed that the shifts of the three metabolic activities between different batches are minimal. For (a) dactolisib, batch1 contains 495 cells, batch2 contains 693 cells and batch3 contains 700 cells. For (b) doxorubicin, batch1 contains 509 cells, batch2 contains 154 cells and batch 3 contains 443 cells. For (c) triacsin-C, batch 1 contains 534 cells, batch2 contains 427 cells and batch 3 contains 953 cells.
Extended Data Fig. 4 Dose responses of the vibrational probes of MDA-MB-231 cells after cycloheximide and daunorubicin treatments.
Error bar represents mean ± SD in 3 groups. For cycloheximide that specifically inhibits protein synthesis, only 13C-AA showed dose-response curve. The IC50 value from 13C-AA is 0.51 μM, which is around 2.5 times smaller than the IC50 value from the cell viability assay. For daunorubicin, the drug that intercalates DNA, signals from all three vibrational probes demonstrated dose responses. This is consistent with its mechanism to interfere with macromolecule metabolism. For the IC50 values, the IC50 from 13C-AA is 4.79 μM; from azido-PA is 1.97 μM; from d34-OA is 5.5 μM. Compared to the one measured in cell viability assay (1.28 μM), these IC50 values are slightly larger but on the same magnitude.
Extended Data Fig. 5 Feature importance ranking from Random Forest classifier.
The ranking was projected on IR spectrum with color gradient coding, where red color indicates more important features. This result further supports that features from metabolic labeling are highly important in predicting drug-perturbed cell phenotypes.
Extended Data Fig. 6 Prediction performance of drug MoAs using label-free FTIR imaging data.
Drugs belong to 6 MoAs were tested. MDA-MB-231 cells were treated by drugs (anisomycin, cycloheximide, MG-132, bortezomib, dactolisib, everolimus, doxorubicin, epirubicin, triacsin-c) at their IC50 concentrations without any labeling. The confusion matrix of the LDA classifier performance based on (a) data from the same batch (5,749 cells) and (b) data from different batches (2,982 cells) are presented. It can be observed that the accuracy dropped dramatically after including data from different batches using label-free measurements.
Extended Data Fig. 7
Test compounds’ Mahalanobis distance of annotated referenced drug MoA groups.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Liu, X., Shi, L., Zhao, Z. et al. VIBRANT: spectral profiling for single-cell drug responses. Nat Methods 21, 501–511 (2024). https://doi.org/10.1038/s41592-024-02185-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-024-02185-x