Abstract
Combination therapy is a promising therapeutic strategy to enhance the efficacy of immune checkpoint blockade (ICB); however, predicting drugs for effective combination is challenging. Here we developed a general data-driven method called CM-Drug for screening compounds that can boost ICB treatment efficacy based on core and minor gene sets identified between responsive and nonresponsive samples in ICB therapy. The CM-Drug method was validated using melanoma and lung cancer mouse models, with combined therapeutic efficacy demonstrated in eight of nine predicted compounds. Among these compounds, taltirelin had the strongest synergistic effect. Mechanistic analysis and experimental verification demonstrated that taltirelin can stimulate CD8+ T cells and is mediated by the induction of thyroid-stimulating hormone. This study provides an effective and general method for predicting and evaluating drugs for combination therapy and identifies candidate compounds for future ICB combination therapy.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
Previously published bulk RNA-seq or microarray data that were reanalyzed here are available under the following accession codes: SRP070710 (ref. 4), SRP094781 (ref. 9), ERP105482 (ref. 10), SRP011540 (ref. 11), SRP128156 (ref. 12), SRP230414 (ref. 13), SRP250849 (ref. 14), ERP107734 (ref. 15), SRP217040 (ref. 16), SRP183455 (ref. 17), GSE130324 (ref. 37), GSE149825 (ref. 38), GSE126988 (ref. 39), PRJNA477854, GSE146834 (ref. 40), GSE148947 (ref. 23), GSE149479 (ref. 41), GSE132004 (ref. 21), GSE79485 (ref. 42), GSE218603 (ref. 56), GSE173107 (ref. 57), GSE186195 (ref. 58), GSE223110 (ref. 59) and GSE162935 (ref. 60). These datasets were from the ICBatlas36 and ICBcomb55 databases, which originally came from the NCBI GEO and dbGap databases. Our bulk RNA-seq data supporting the findings of this study have been deposited in the Genome Sequence Archive under accession code CRA011467. Source data are provided with this paper. All other data are available in the article and supplementary files or from the corresponding authors upon reasonable request.
Code availability
Code has been uploaded to GitHub at https://github.com/cloudsummer/CM-Drug (ref. 61).
References
Wei, S. C., Duffy, C. R. & Allison, J. P. Fundamental mechanisms of immune checkpoint blockade therapy. Cancer Discov. 8, 1069–1086 (2018).
Korman, A. J., Garrett-Thomson, S. C. & Lonberg, N. The foundations of immune checkpoint blockade and the ipilimumab approval decennial. Nat. Rev. Drug Discov. 21, 509–528 (2022).
Sharma, P. & Allison, J. P. The future of immune checkpoint therapy. Science 348, 56–61 (2015).
Hugo, W. et al. Genomic and transcriptomic features of response to anti-PD-1 therapy in metastatic melanoma. Cell 165, 35–44 (2016).
Kwon, M. et al. Determinants of response and intrinsic resistance to PD-1 blockade in microsatellite instability-high gastric cancer. Cancer Discov. 11, 2168–2185 (2021).
Meric-Bernstam, F., Larkin, J., Tabernero, J. & Bonini, C. Enhancing anti-tumour efficacy with immunotherapy combinations. Lancet 397, 1010–1022 (2021).
Yap, T. A. et al. Development of immunotherapy combination strategies in cancer. Cancer Discov. 11, 1368–1397 (2021).
Wang, Z. et al. Immunogenic camptothesome nanovesicles comprising sphingomyelin-derived camptothecin bilayers for safe and synergistic cancer immunochemotherapy. Nat. Nanotechnol. 16, 1130–1140 (2021).
Riaz, N. et al. Tumor and microenvironment evolution during immunotherapy with nivolumab. Cell 171, 934–949 (2017).
Gide, T. N. et al. Distinct immune cell populations define response to anti-PD-1 monotherapy and anti-PD-1/anti-CTLA-4 combined therapy. Cancer Cell 35, 238–255 (2019).
Auslander, N. et al. Robust prediction of response to immune checkpoint blockade therapy in metastatic melanoma. Nat. Med. 24, 1545–1549 (2018).
Miao, D. et al. Genomic correlates of response to immune checkpoint therapies in clear cell renal cell carcinoma. Science 359, 801–806 (2018).
Tumeh, P. C. et al. PD-1 blockade induces responses by inhibiting adaptive immune resistance. Nature 515, 568–571 (2014).
Amato, C. M. et al. Pre-treatment mutational and transcriptomic landscape of responding metastatic melanoma patients to anti-PD1 immunotherapy. Cancers 12, 1943 (2020).
Kim, S. T. et al. Comprehensive molecular characterization of clinical responses to PD-1 inhibition in metastatic gastric cancer. Nat. Med. 24, 1449–1458 (2018).
Jung, H. et al. DNA methylation loss promotes immune evasion of tumours with high mutation and copy number load. Nat. Commun. 10, 4278 (2019).
Cho, J.-W. et al. Genome-wide identification of differentially methylated promoters and enhancers associated with response to anti-PD-1 therapy in non-small cell lung cancer. Exp. Mol. Med. 52, 1550–1563 (2020).
Bhattacharya, S. et al. ImmPort: disseminating data to the public for the future of immunology. Immunol. Res. 58, 234–239 (2014).
Miao, Y.-R. et al. ImmuCellAI: a unique method for comprehensive T-cell subsets abundance prediction and its application in cancer immunotherapy. Adv. Sci. 7, 1902880 (2020).
Zemek, R. M. et al. Sensitization to immune checkpoint blockade through activation of a STAT1/NK axis in the tumor microenvironment. Sci. Transl. Med. 11, eaav7816 (2019).
Lu, W. et al. Reprogramming immunosuppressive myeloid cells facilitates immunotherapy for colorectal cancer. EMBO Mol. Med. 13, e12798 (2021).
Pelly, V. S. et al. Anti-inflammatory drugs remodel the tumor immune environment to enhance immune checkpoint blockade efficacy. Cancer Discov. 11, 2602–2619 (2021).
Ou, D.-L. et al. Regorafenib enhances antitumor immunity via inhibition of p38 kinase/Creb1/Klf4 axis in tumor-associated macrophages. J. Immunother. Cancer 9, e001657 (2021).
Subramanian, A. et al. A next generation connectivity map: L1000 platform and the first 1,000,000 profiles. Cell 171, 1437–1452 (2017).
Braverman, L. E. & Cooper, D. Werner & Ingbar’s the Thyroid: A Fundamental and Clinical Text (Lippincott Williams & Wilkins, 2012).
Zhu, S. et al. Combination strategies to maximize the benefits of cancer immunotherapy. J. Hematol. Oncol. 14, 156 (2021).
PD-1/PD-L1 landscape. Cancer Research Institute www.cancerresearch.org/pd-1-pd-l1-landscape (2022).
Galluzzi, L., Humeau, J., Buqué, A., Zitvogel, L. & Kroemer, G. Immunostimulation with chemotherapy in the era of immune checkpoint inhibitors. Nat. Rev. Clin. Oncol. 17, 725–741 (2020).
Beaver, J. A. & Pazdur, R. The Wild West of checkpoint inhibitor development. N. Engl. J. Med. 386, 1297–1301 (2022).
Muir, C. A. et al. Thyroid immune-related adverse events following immune checkpoint inhibitor treatment. J. Clin. Endocrinol. Metab. 106, e3704–e3713 (2021).
Brancatella, A. et al. Graves’ disease induced by immune checkpoint inhibitors: a case report and review of the literature. Eur. Thyroid J. 8, 192–195 (2019).
Kotwal, A., Kottschade, L. & Ryder, M. PD-L1 inhibitor-induced thyroiditis is associated with better overall survival in cancer patients. Thyroid 30, 177–184 (2020).
Muir, C. A., Tsang, V. H. M., Menzies, A. M. & Clifton-Bligh, R. J. Immune related adverse events of the thyroid—a narrative review. Front. Endocrinol. 13, 886930 (2022).
Khan, Z. et al. Genetic variation associated with thyroid autoimmunity shapes the systemic immune response to PD-1 checkpoint blockade. Nat. Commun. 12, 3355 (2021).
Dawidowska, A. et al. Immune-related thyroid adverse events predict response to PD-1 blockade in patients with melanoma. Cancers 14, 1248 (2022).
Yang, M. et al. ICBatlas: a comprehensive resource for depicting immune checkpoint blockade therapy characteristics from transcriptome profiles. Cancer Immunol. Res. 10, 1398–1406 (2022).
Di Tacchio, M. et al. Tumor vessel normalization, immunostimulatory reprogramming, and improved survival in glioblastoma with combined inhibition of PD-1, angiopoietin-2, and VEGF. Cancer Immunol. Res. 7, 1910–1927 (2019).
Gu, S. S. et al. Therapeutically increasing MHC-I expression potentiates immune checkpoint blockade. Cancer Discov. 11, 1524–1541 (2021).
Liu, P. et al. Crizotinib-induced immunogenic cell death in non-small cell lung cancer. Nat. Commun. 10, 1486 (2019).
Li, J. et al. Epigenetic and transcriptional control of the epidermal growth factor receptor regulates the tumor immune microenvironment in pancreatic cancer. Cancer Discov. 11, 736–753 (2021).
Sirait-Fischer, E. et al. Immune checkpoint blockade improves chemotherapy in the PyMT mammary carcinoma mouse model. Front. Oncol. 10, 1771 (2020).
Eissler, N. et al. Regulation of myeloid cells by activated T cells determines the efficacy of PD-1 blockade. Oncoimmunology 5, e1232222 (2016).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Leek, J. T., Johnson, W. E., Parker, H. S., Jaffe, A. E. & Storey, J. D. The sva package for removing batch effects and other unwanted variation in high-throughput experiments. Bioinformatics 28, 882–883 (2012).
Kim, D., Paggi, J. M., Park, C., Bennett, C. & Salzberg, S. L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat. Biotechnol. 37, 907–915 (2019).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Korotkevich, G. et al. Fast gene set enrichment analysis. Preprint at bioRxiv https://doi.org/10.1101/060012 (2021).
Fang, Z., Liu, X. & Peltz, G. GSEApy: a comprehensive package for performing gene set enrichment analysis in Python. Bioinformatics 39, btac757 (2023).
Gu, Z., Eils, R. & Schlesner, M. Complex heatmaps reveal patterns and correlations in multidimensional genomic data. Bioinformatics 32, 2847–2849 (2016).
Wu, T. et al. clusterProfiler 4.0: a universal enrichment tool for interpreting omics data. Innovation 2, 100141 (2021).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Miao, Y.-R. et al. ImmuCellAI-mouse: a tool for comprehensive prediction of mouse immune cell abundance and immune microenvironment depiction. Bioinformatics 38, 785–791 (2022).
Shumate, A., Wong, B., Pertea, G. & Pertea, M. Improved transcriptome assembly using a hybrid of long and short reads with StringTie. PLoS Comput. Biol. 18, e1009730 (2022).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Xia, Y. et al. ICBcomb: a comprehensive expression database for immune checkpoint blockade combination therapy. Brief. Bioinform. 25, bbad457 (2023).
Kadota, H. et al. Anti-programmed cell death-1 antibody and dasatinib combination therapy exhibits efficacy in metastatic colorectal cancer mouse models. Cancers 14, 6146 (2022).
Borchmann, S. et al. Tripartite antigen-agnostic combination immunotherapy cures established poorly immunogenic tumors. J. Immunother. Cancer 10, e004781 (2022).
Tilsed, C. M. et al. Retinoic acid induces an IFN-driven inflammatory tumour microenvironment, sensitizing to immune checkpoint therapy. Front. Oncol. 12, 849793 (2022).
Liu, N. et al. Supplementation with α-ketoglutarate improved the efficacy of anti-PD1 melanoma treatment through epigenetic modulation of PD-L1. Cell Death Dis. 14, 170 (2023).
Bruand, M. et al. Cell-autonomous inflammation of BRCA1-deficient ovarian cancers drives both tumor-intrinsic immunoreactivity and immune resistance via STING. Cell Rep. 36, 109412 (2021).
CM-Drug. GitHub https://github.com/cloudsummer/CM-Drug (2024).
Acknowledgements
This project was supported by the National Natural Science Foundation of China (31822030 and 32370717 to A.-Y.G., 82272844 to L.G.) and the National Key R&D Program of China (2021YFF0703704 to A.-Y.G.). We thank the Research Core Facilities for Life Science (Huazhong University of Science and Technology (HUST)), the Analytical and Testing Center of HUST, and the Optical Bioimaging Core Facility of Wuhan National Laboratory for Optoelectronics, HUST, for related analysis. We thank all the participants involved in the production of ICB data used in this project.
Author information
Authors and Affiliations
Contributions
A.-Y.G. and Y.X. conceived and provided the main idea of the project. Y.X. contributed to the bioinformatics and experimental data analyses, leading to critical findings. A.-Y.G., L.G., Y.X. and X.L. designed the main experiments. Y.X. created the figures and diagrams in the article. X.L. performed the main experiments. N.B. conducted a partial ICB booster efficacy experiment. Y.-R.M. assisted with a part of the methods. W.P. and M.Y. collected some public datasets for this project. W.P. assisted with some figure drawings. Y.G. assisted with literature research on the applicability of drugs. C.C. and H.L. performed experiments to detect TSH and TH levels. A.-Y.G. and Y.X. wrote the paper with input from the other authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Immunology thanks Eytan Ruppin and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: N. Bernard, in collaboration with the Nature Immunology team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Core & Minor gene sets expression pattern.
a, Expression pattern of Core & Minor gene sets in mouse validation data of renal carcinoma. b,c, The heatmaps depicting Gene Set Variation Analysis (GSVA) enrichments for mice in response and non-response groups following ICB treatment in the mesothelioma (b) and renal cancer (c) of datasets GSE117358. d-f, GSEA enrichment plot of the six Core & Minor gene sets among different comparison groups in datasets of GSE132004 (d), GSE148950 (e) and GSE160785 (f). P-values were calculated by two-tailed Kolmogorov Smirnov test, then FDR (BH) for correction of p-values.
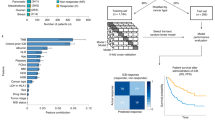
Extended Data Fig. 2 Predicting ICB booster by CM-Drug and validation in the melanoma mouse model.
a, Candidate compounds in human melanoma A375 cell line using LINCS L1000 data by CM-Drug and ranked them by CM-Score from high to low. The red and blue dots in the figure represent the NES score values of the Core and Minor gene sets, respectively. The dotted box represents the top 20 CM-Score compounds we focus. b, Individual tumor growth curves of B16-F10 tumor-bearing mice after treatment with PBS, miglitol, MG-132, taltirelin, anti-PD-1, a combination of anti-PD-1 with miglitol, MG-132 or taltirelin indicated in Fig. 4b. c, Histological observation of major organs in B16-F10 mice-bearing mice after new round taltirelin efficacy experiment by H&E staining. Images are representative of two biologically independent mice. Scale bar: 20 µm. d, Serological analysis in B16-F10 tumor-bearing mice after new round taltirelin efficacy experiment. Alanine aminotransferase (ALT), aspartate aminotransferase (AST), lactate dehydrogenase (LDH), creatine kinase (CK), blood urea nitrogen (BUN), and creatinine (CREA). Data are presented as means ± s.d. n = 9 mice per group. The red dashed lines represent the range of normal reference values.
Extended Data Fig. 3 Anti-tumor efficacy of TRH/TSH/TH alone or combined with anti-PD-1 in B16-F10 tumor-bearing mice.
a, Average tumor growth curves (full day 0 to day 13) of B16-F10 tumor-bearing mice after treatment indicated in Fig. 4e. Data are presented as the means ± s.d. b, Images of tumor tissues at the end of treatments. The cross symbols within rectangles represent no tumor due to tumor volume exceeding ethical limits or actual mouse death. c, Average tumor growth rate of B16-F10 tumor-bearing mice after treatment. Pairwise comparisons by two-sided Mann–Whitney U test, then FDR (BH) for correction of p-values. d, Kaplan-Meier survival plot of B16-F10 tumor-bearing mice after treatment. e, Images of B16-F10 tumor-bearing mice on day 7 and day 13 after the first treatment. f, Histological observation of major organs in B16-F10 mice-bearing mice at the end of treatments by H&E staining. Images are representative of two biologically independent mice. Scale bar: 20 µm. n = 9 for TSH and TH group, n = 8 for other groups (a, c, d).
Extended Data Fig. 4 Serological analysis.
a, Serological analysis in B16-F10 tumor-bearing mice at 14 days after treatment indicated in Fig. 4e. Alanine aminotransferase (ALT), aspartate aminotransferase (AST), lactate dehydrogenase (LDH), creatine kinase (CK), blood urea nitrogen (BUN), and creatinine (CREA). Data are presented as means ± s.d. n = 7 (PBS), 4 (TRH), 5 (TSH), 7 (TH), 7 (Anti-PD-1), 7 (TRH+anti-PD-1), 8 (TSH+anti-PD-1) and 6 (TH+anti-PD-1). The red dashed line represents the range of normal reference values. b, c, TSH (b) and Thyroxine (T4) (c) content in the serum of B16-F10 tumor-bearing mice at 14 days after treatment indicated in Fig. 4e. Data are presented as means ± s.d. n = 6 (PBS), 4 (TRH), 5 (TSH), 6 (TH), 6 (anti-PD-1), 6 (TRH+anti-PD-1), 6 (TSH+anti-PD-1) and 5 (TH+anti-PD-1) for b; n = 6 (PBS), 4 (TRH), 5 (TSH), 6 (TH), 6 (Anti-PD-1), 6 (TRH+anti-PD-1), 6 (TSH+anti-PD-1) and 6 (TH+anti-PD-1) for c.
Extended Data Fig. 5 Pattern validation of Core & Minor genes in mice treated with anti-PD-1 and taltirelin.
a, Schematic schedule for the anti-tumor experiment in B16-F10 tumor-bearing mice after intraperitoneal injection of PBS, taltirelin (0.5 mg kg−1) every 2 days for six times in the presence or absence of intraperitoneal injection of anti-PD-1 (100 μg per mouse) every 3 days for four times. The arrows indicate the drug injection time. b-e, Individual tumor growth curves of B16-F10 tumor-bearing mice after treatment as indicated. f,g, Average tumor growth curves (f) and tumor weights (g) of B16-F10 tumor-bearing mice after treatment indicated in a. Data are presented as means ± s.d. n = 8-10 mice per group as indicated in the figure. Due to the non-normal distribution of tumor volume data in both the anti-PD-1 and combination groups, non-parametric tests were employed: Kruskal-Wallis H test followed by two-sided Mann–Whitney U test pairwise comparisons, then FDR (BH) for correction of p-values. h, Average tumor growth rate of B16-F10 tumor-bearing mice in different groups. Data are presented as means ± s.d. n = 8-10 mice per group as indicated in the figure. Pairwise comparisons by two-sided Mann–Whitney U test, then FDR (BH) for correction of p-values. i, j, Expression pattern of the Core & Minor genes in tumor tissues. Images of tumor tissues at the end of tumor experiments (i). The tumor tissues circled in red are non-responders. Heatmap diagram representing the gene expression in six Core & Minor gene sets from RNA-seq analysis of tumor tissues after treatment indicated in a (j). Colors orange to blue represent gene expression from high to low. k-m, GSEA enrichment plot of six Core & Minor gene sets in four comparisons. Combination group vs. anti-PD-1 group (k), responders (R) vs. non-responders (NR) in the anti-PD-1 group and combination group (l). R vs. NR in taltirelin group (m).
Extended Data Fig. 6 Volcano plot of DEGs.
a-c, DEGs between response (R) and non-response (NR) samples of anti-PD-1 monotherapy (a), taltirelin monotherapy (b), and anti-PD-1 combined with taltirelin (c) groups in B16-F10 tumors on day 11 after treatment indicated in Extended Data Fig. 5a. Significantly (FDR < 0.05 and |log2FoldChange| > 1, P-values were calculated using a two-tailed Wald test, then FDR (BH) for correction of p-values. FDR values were used as criteria for significance filtering) activated and inhibited genes are highlighted in red and blue, respectively.
Extended Data Fig. 7 KEGG enrichment results (top 20).
The top 20 enriched pathways of the upregulated DEGs between responders (R) and non-responders (NR) for the single use of anti-PD-1 (a), single use of taltirelin (b), and combination treatment groups (c), respectively. d, KEGG enrichment results (q-value and hit count) of the ‘Autoimmune thyroid disease’ on DEGs between R and NR samples from monotherapy group or combination-treated group. e, Schematic diagram of genes in KEGG thyroid-related pathways overlapping with Core & Minor genes.
Extended Data Fig. 8 Anti-tumor immunity induced by anti-PD-1 combined with taltirelin therapy.
a, Gating strategies for identifying matured DCs (CD45+, F4/80−, CD11c+, MHC-II+, CD80+, CD86+), NK cells (CD45+, CD3−, NK1.1+), B cells (CD45+, CD3−, CD19+), NKT cells (CD45+, CD3+, NK1.1+), γδ T cells (CD45+, CD3+, TCR, γδ+), CD8+ T cells (CD45+, CD3+, CD8+), effector CD8+ T cells (CD45+, CD3+, CD8+, GzmB+), CD8+ Tcm cells (CD45+, CD3+, CD8+, CD44+, CD62L+), CD8+ Tem cells (CD45+, CD3+, CD8+, CD44+, CD62L−), CD4+ T cells (CD45+, CD3+, CD4+), CD4+ Tcm cells (CD45+, CD3+, CD4+, CD44+, CD62L+) and CD4+ Tem cells (CD45+, CD3+, CD4+, CD44+, CD62L−) in tumor tissues of B16-F10 tumor-bearing mice presented on Fig. 5f and b–d. b–d, The numbers of CD3+ T cells (b), NK cells (c) and B cells (d) in tumor tissues of B16-F10 tumor-bearing mice after treatment indicated in a. Data are presented as means ± s.d. n = 7 (PBS), 9 (taltirelin), 10 (anti-PD-1) and 10 (combination). One-way ANOVA followed by Bonferroni’s multiple comparisons post-test.
Extended Data Fig. 9 In vitro mechanism investigation of taltirelin sensitized ICB therapy.
a, Cell viability of B16-F10 cells after treatment with PBS, taltirelin (100 nM, 200 nM, 400 nM, 800 nM, 1600 nM) or TSH (10 nM, 20 nM, 40 nM, 80 nM) for 24 h. b, Flow cytometry gating strategy for the analysis of CD8+ T cells ex vivo. c,d, Representative flow cytometry images (c) and quantification (d) of the CFSE low cells of CD8+ T cells stimulated with PBS, taltirelin (1, 10, 100, 500, 1000 nM) or TSH (20 nM) in the presence or absence of 2.5 μg mL−1 ConA for 5 days. e,f, Percentages of IFN-γ+ (e) and GzmB+ cells (f) in CD8+ T cells after treatment as indicated in c by flow cytometry. One-way ANOVA followed by Tukey’s HSD post hoc test. g, Percentage of PD-1+ cells in CD8+ OT-I T cells after treatment indicated in Fig. 6i by flow cytometry. h, Cell viability of B16-F10 cells after treatment with PBS or L-thyroxine (10−8 M, 10−7 M, 10−6 M, 10−5 M) for 24 h. Data are presented as means ± s.d. i-k, CD86 (i), MHC-I SINFEKL (j) and MHC-II (k) MFI on BMDCs after treatment with PBS or L-thyroxine (10−8 M, 10−7 M, 10−6 M, 10−5 M) in the presence or absence of 10 μg mL−1 OVA for 24 h by flow cytometry. l, Percentages of IFN-γ+ in CD8+ T cells after treatment with PBS or L-thyroxine (10−8 M, 10−7 M, 10−6 M, 10−5 M) in the presence or absence of 2.5 μg mL−1 ConA for 5 days by flow cytometry. Data are presented as means ± s.d (a, d-l). n = 6 biologically independent samples (a,h). n = 3 biologically independent samples (d-g, i-l). One-way ANOVA followed by Bonferroni’s multiple comparisons post-test (a, g-l). One-way ANOVA followed by Tukey’s HSD post hoc test (d–f).
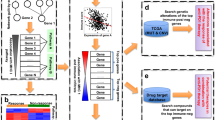
Extended Data Fig. 10 Diagram illustrating the method section.
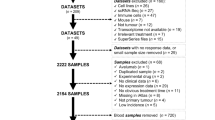
a, b, Dot plots of t-SNE dimension reduction of 10 ICB clinical treatment datasets. SRP128156 including two sub-datasets: G100556 (red circle) and G102647(blue circle), t-SNE dimension reduction shows SRP128156 has an obvious batch effect (a). t-SNE dimension reduction shows the results of batch effect removing by the ComBat function in SVA (R package) for SRP128156 dataset (b). c, Details on the identification of ICB-related gene sets. The Antigen processing and presentation pathway was taken as an example: In 7 of the 10 datasets, ‘Antigen processing and presentation pathway’ results are positively enriched significantly, and the corresponding 7 leading-edge subsets are obtained. If the gene appears in more than 70 % of leading-edge subsets (7 × 70% = 4.9, that is, more than or equal to 5 leading-edge subsets), the gene is retained. Otherwise, the gene is discarded. (P-values were calculated by two-tailed Kolmogorov Smirnov test, then FDR (BH) for correction of p-values.) d,e, Overview of the LASSO model test data. 23 positive samples for test data (samples from NCBI GEO database, GEO accession number: GSE218603, GSE173107, GSE186195, GSE223110, and GSE162935, detail in Table S7) (d). 2573 negative samples for test data (GSEA was conducted on the gene expression data perturbed by small-molecule compounds from the LINCS 2020 dataset (e). Samples exhibiting negatively significant enrichment (NES < 0, FDR < 0.05) in the Core & Minor gene sets following perturbation are considered ‘negative samples’, and their efficacy labels are set to ‘0’.).
Supplementary information
Supplementary Tables
Supplementary Tables 1–13.
Source data
Source Data Fig. 6
Unprocessed scans corresponding to Fig. 6a (PCR).
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Xia, Y., Li, X., Bie, N. et al. A method for predicting drugs that can boost the efficacy of immune checkpoint blockade. Nat Immunol 25, 659–670 (2024). https://doi.org/10.1038/s41590-024-01789-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-024-01789-x