Abstract
Macrophages have long been considered as particularly plastic cells. However, recent work combining fate mapping, single-cell transcriptomics and epigenetics has undermined the macrophage plasticity dogma. Here, we discuss recent studies that have carefully dissected the response of individual macrophage subsets to pulmonary insults and call for an adjustment of the macrophage plasticity concept. We hypothesize that prolonged tissue residency shuts down much of the plasticity of macrophages and propose that the restricted plasticity of resident macrophages has been favored by evolution to safeguard tissue homeostasis. Recruited monocytes are more plastic and their differentiation into resident macrophages during inflammation can result in a dual imprinting from both the ongoing inflammation and the macrophage niche. This results in inflammation-imprinted resident macrophages, and we speculate that rewired niche circuits could maintain this inflammatory state. We believe that this revisited plasticity model offers opportunities to reset the macrophage pool after a severe inflammatory episode.
Similar content being viewed by others
Main
Macrophages are thought to be particularly plastic1. This plasticity is considered central to their ability to switch from a basal state, in which they perform tissue-specific homeostatic functions2,3,4,5, to an inflammatory state, in which they are capable of eliminating invading pathogens. In this Perspective, we will focus on the distinct subsets of lung macrophages (Fig. 1) and discuss recent reports that have carefully studied the functional plasticity of these cells. Before we zoom in on the specific subsets of pulmonary macrophages, it is important to distinguish resident macrophages (ResMacs) that were present before the inflammation from macrophages that developed during the inflammation. Most ResMacs resident in tissue self-maintain throughout life with minimal input from circulating monocytes on a day-to-day basis (Fig. 2a). Loss of ResMacs allows circulating monocytes to engraft in the ResMac pool6,7,8, but not all recruited monocytes differentiate into long-lived ResMacs. In some cases, monocytes recruited to inflamed tissues differentiate into short-lived macrophages that are only present during the inflammatory window and disappear during the resolution phase9,10. We propose to define these short-lived cells as transient macrophages (TransMacs; Fig. 2b). Additionally, rather than subdividing macrophages into embryonic and recruited/monocyte-derived macrophages, we propose to distinguish TransMacs from ResMacs regardless of their cellular origin, as most ResMacs contain some monocyte-derived cells that were recruited under steady-state conditions11,12. We also need to define a third group of macrophages that were recently described as functionally distinct. Namely, ResMacs that developed during inflammation and acquired an epigenetic memory of that event13, which we refer to as inflammation-imprinted ResMacs (InfResMacs; Fig. 2b). While fate-mapping tools have helped to distinguish ResMacs from InfResMacs, we often lack a clear profile of TransMacs. As such, in this Perspective, we will focus on the ontogeny, functional characteristics, plasticity and epigenetic memory of ResMacs and InfResMacs.
Three subsets of macrophages reside within the lung. The most abundant are SiglecFhiCD11chi AMs located in the alveoli. AMs are found in proximity to type I/II epithelial cells and perialveolar interstitial fibroblasts. Together, these cells probably comprise the AM niche. Two main populations of IMs have been identified: Lyve1hiCD206hi IMs and CX3CR1hiMHC-IIhi IMs. Their precise localization within the parenchyma and the composition of their niche has not been determined. However, reports have described Lyve1hiCD206hi and CX3CR1hiMHC-IIhi IMs to be preferentially associated with blood vessels and nerve cells proximal to the bronchioles, respectively. We refer to these cells as perivascular and peribronchial IMs. The ability to determine the localization of these cells suggests the possibility that endothelial cells and pericytes may represent the niche for perivascular IMs, while nerve cells, peribronchial fibroblasts and smooth muscle cells could represent the niche for peribronchial IMs. BACH2, BTB domain and CNC homolog 2; CCR2, C–C chemokine receptor type 2; CEBPβ, CCAAT/enhancer-binding protein β; MHC-II, major histocompatibility complex II; PPARγ, peroxisome proliferator-activated receptor γ.
a, Pulmonary macrophages develop alongside the lung from multiple sequential and overlapping waves of embryonic precursors derived from distinct locations of embryonic hematopoiesis. The lung develops in overlapping stages from the pseudoglandular stage (E12.5–E15.5) over the canalicular stage (E15.5–E17.5) and the saccular stage (E17.5 until birth) to the alveolar stage (postnatal week 1 until postnatal week 3). The magnifications show the sequential increase in branching and the formation of the alveoli. b, The onset of inflammation and subsequent resolution reshape the composition of the pulmonary macrophage pool. An inflammatory episode results in the recruitment of monocytes. Some monocytes differentiate into short-lived TransMacs, which will disappear as the inflammation is resolved. Some monocytes can differentiate into InfResMacs and persist in the lung.
Pulmonary ResMac subsets
Three distinct pulmonary ResMac subsets have been described in the murine lung (Fig. 1): SiglecFhiCD11chi alveolar macrophages (AMs), Lyve1hiCD206hi interstitial macrophages (IMs) and CX3CR1hiMHC-IIhi IMs14,15,16. It has been proposed that each tissue contains a restricted number of niches17, but the precise composition of each macrophage niche remains to be determined. AMs reside in the alveoli, where type II epithelial cells produce colony-stimulating factor 2 (CSF2; also known as granulocyte-macrophage colony-stimulating factor), which, together with autocrine transforming growth factor-β signaling, is critical for the induction of peroxisome proliferator-activated receptor-γ in AMs and the expression of a gene module that is essential for surfactant recycling18,19,20,21,22. Based on reports of cross-talk between liver Kupffer cells and liver mesenchymal cells called stellate cells23, we consider the possibility that there could be similar cross-talk between AMs and mesenchymal perialveolar interstitial fibroblasts (Fig. 1).
The precise localization of the two main IM populations and the composition of their respective niche is unclear. Some have proposed that both Lyve1hiCD206hi IMs and CX3CR1hiMHC-IIhi IMs are mainly found in the peribronchial region15. Others have proposed that the Lyve1hiCD206hi IMs are preferentially found in the peribronchial region, while CX3CR1hiMHC-IIhi IMs are found in the alveolar interstitium16. CX3CR1hiMHC-IIhi IMs expressing higher amounts of CD169 (ref. 24) were reported to be closely associated with nerves14,24, while the Lyve1hiCD206hi IMs were reported to associate with blood vessels14 and to have a role in blood vessel integrity14, which may be the result of cross-talk with endothelial cells and pericytes. Lyve1hiCD206hi IMs produce high amounts of interleukin (IL)-10 (ref. 14), implying that they may not only regulate the infiltration of immune cells, but also participate in the immune suppression of extravasating cells, including dendritic cells14,25,26. The association of CX3CR1hiMHC-IIhi IMs with nerves14,24 is reminiscent of the macrophage–nerve crosstalk described in the intestinal muscularis externa and the dermis27,28,29. However, it should be noted that the distinction between blood- and nerve-associated macrophages in the bronchioles is not always clear cut16. Peribronchial fibroblasts and lymphatic vessels are also found in the bronchioles (Fig. 1) and may also comprise the peribroncheal IM niche. More work is needed to identify robust markers to distinguish the IM subsets, unravel their precise localization, determine which cells constitute their respective niches and understand the cell–cell interactions driving their distinct transcriptome and unique functions.
Ontogeny of pulmonary ResMacs
There are multiple sequential and overlapping waves of embryonic precursors originating from distinct sites of embryonic hematopoiesis30,31,32 (Fig. 2a). The precise contribution of these distinct waves to the final pool of ResMacs remains highly debated. AMs develop in the first week of life18 and then self-maintain. Experiments using parabiotic mice10,18, shielded chimeras33 and Cx3cr1- or Pdzk1ip1-based fate mapping12,34 found that bone marrow precursors contribute only minimally to the adult AM pool. However, studies using the Ms4a3-based fate-mapping mouse found a greater contribution of bone marrow-derived monocytes to the AM pool than was previously reported11. It is currently unclear why this system predicts greater monocyte contribution. Additionally, fate mapping of early yolk sac precursors using tamoxifen-inducible Csf1r-CRE or Runx1-CRE mice efficiently labeled microglia but not AMs35. The identification of what looks like a monocyte waterfall continuum between fetal monocytes and pre-AMs before birth led to the hypothesis that AMs would primarily be derived from fetal liver monocytes18. Therefore, AMs could be primarily derived from late erythro-myeloid progenitor (EMP)-derived fetal liver progenitors35. Applying trajectory inference algorithms to single-cell sequencing data from lung development has also linked AMs to a monocytic intermediate36. Transfers of distinct precursors into Csf2ra−/− mice revealed that yolk sac macrophages, fetal liver monocytes and bone marrow-derived monocytes could generate self-maintaining and functional AMs that protect mice from developing pulmonary alveolar proteinosis, although in competition settings, fetal liver monocytes generated more AMs than the other precursors8. Therefore, most AMs probably develop from fetal liver monocytes, with some small contributions from yolk sac macrophages and bone marrow-derived monocytes (Fig. 2a).
The ontogeny of IMs is less well understood. Fate mapping of yolk sac-derived macrophages reveals that many IMs are present in the lung interstitium before birth37, and are later diluted by an influx of bone marrow-derived monocytes14. IMs were identified in tamoxifen-inducible Csf1r-CRE fate-mapping studies38, while parabiosis and fate-mapping experiments reveal a continuous contribution of bone marrow precursors to the IM pool14,37. Altogether, this indicates that while the contribution of yolk sac macrophages to IMs is higher than for AMs, these cells are first diluted by fetal liver monocytes and then by a slow and continuous input from bone marrow-derived monocytes after birth (Fig. 2a).
Potential link between macrophage residency and plasticity
Ontogeny studies have revealed that Lyve1hiCD206hi IMs, CX3CR1hiMHC-IIhi IMs and SiglecFhiCD11chi AMs contain a mix of macrophages from distinct cellular origins. Despite this, single-cell analysis reveals relatively homogeneous populations14,36,39,40. It is of course possible that some transcriptomic differences may have been missed by single-cell RNA sequencing. However, when distinct precursors were transferred into the lungs of neonatal mice and the fully differentiated AMs that developed from these precursors were later profiled by bulk transcriptomic profiling, only few transcriptomic differences linked to ontogeny were found8. Although the possibility of epigenetic differences cannot be excluded, these findings highlight a dominant role for tissue imprinting in the cellular identity of steady-state macrophages, as this imprinting seems to overwrite most of the transcriptomic differences of the distinct precursors4,8,17,33,41.
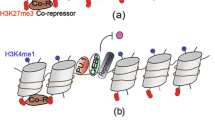
To explain why AMs developing from monocytes during pulmonary inflammation differ functionally13,42,43 from pre-existing AMs, we propose that the steady-state niche is not only able to impose the identity of ResMacs resident in tissue, but is also able to restrict the plasticity of these cells (Fig. 3). This would allow steady-state tissues to imprint a relatively consistent transcriptomic identity on different types of precursors and shape these cells into similar ResMacs regardless of their ontogeny (Fig. 3a). The timing of AM development may also influence their functional specialization. Neonatal lungs display a strong type 2 bias44. The production of IL-33 and CSF2 by epithelial cells is thought to synergize with the production of IL-13 and CSF2 by basophils and group 2 innate lymphoid cells to drive AM development36. However, the precise contribution of CSF2 from epithelial cells versus immune cells is still unknown. Studies across multiple organs have focused on the transcriptomic identity gained during the development into ResMacs23,45,46 and have tried to identify the tissue-specific signals that induce the expression of particular tissue-specific gene modules. Much less is known about the signals that repress genes that were expressed in the precursors. The mechanisms underpinning the ability of the niche to epigenetically silence monocyte- and inflammation-associated genes remain unknown (Fig. 3a). Furthermore, it is not clear whether these mechanisms are tissue specific.
a, At steady state, the niche is able to imprint tissue-resident identity on macrophages and restrict their plasticity. As a result, distinct precursors can be shaped into ResMacs with a similar transcriptomic profile, irrespective of their ontogeny. The steady-state circuits that maintain the ResMac profile are also responsible for silencing the monocyte signature and establishing the ResMac profile. b, Monocytes that differentiate into InfResMacs during inflammation possess a mixed identity and express both a tissue-specific signature, akin to their ResMac counterparts, and an inflammatory signature resulting from the inflammatory event during which they differentiated. c, ResMacs present before the onset of inflammation regain their steady-state transcriptomic profile rapidly. However, InfResMacs are more strongly imprinted and, as a result, retain a proportion of their inflammatory signature, resulting in a stronger reaction to a subsequent insult. d, Once the tissue returns to homeostasis, steady-state circuits begin to convert InfResMacs into ResMacs. With time, these cells would again become restricted in their plasticity and closely resemble ResMacs.
We hypothesize that some genes encoding pro-inflammatory cytokines, which are poised or lowly expressed in circulating monocytes, are epigenetically silenced in ResMacs to ensure that terminally differentiated ResMacs do not over-react and do not cause too much tissue damage. Monocytes that differentiate during inflammation and can access the ResMac niche would differentiate into InfResMacs with a mixed identity: they would express part of the main tissue-specific signature of ResMacs but also some genes that have been induced by inflammatory signals (Fig. 3b). ResMacs present in the tissue before the inflammatory event would recover their steady-state transcriptomic profile more rapidly during the resolution phase and show only minor differences when restimulated months later. In contrast, InfResMacs would have undergone stronger imprinting from the inflammation and would maintain a proportion of their inflammatory profile during the resolution phase, yielding a stronger reaction upon restimulation (Fig. 3c). Once the tissue returns to homeostasis, InfResMacs would convert into ResMacs with restricted plasticity. The epigenetic profile of InfResMacs would be converted to a ResMac profile—a state in which these cells would be restricted in their plasticity and in their capacity to produce pro-inflammatory cytokines (Fig. 3d). The experimental data supporting this theoretical model are discussed below.
Macrophage origin and plasticity during pulmonary fibrosis
Fibrotic lung diseases are associated with fibroblast activation and extracellular matrix deposition47. Monocytes and macrophages have been implicated in both the initiation and resolution of fibrosis48 and both AMs and IMs are found in fibrotic lesions49,50. Treatment with bleomycin results in a decrease in AM numbers, but the exact signals leading to the AM disappearance are yet to be elucidated42. Changes to the alveolar epithelium and pulmonary fibroblasts, which may constitute the AM niche (Fig. 1), could render the niche unable to support homeostatic AMs. Bleomycin can induce changes to the alveolar epithelium, driving cellular senescence and apoptosis, as well as initiating epithelial–mesenchymal transition51,52. Indeed, repetitive injury of the alveolar epithelium is thought to be a key trigger in the development of idiopathic pulmonary fibrosis53. Following the loss of resident AMs (ResAMs), the niche is repopulated by a massive engraftment of bone marrow-derived monocytes that give rise to the monocyte-derived AMs (and therefore represent InfResMacs; Fig. 2b) that appear to be key drivers of bleomycin-induced pulmonary fibrosis42. The differentiation of these pro-fibrotic monocyte-derived cells into AMs is supported by single-cell sequencing49. SiglecF+CD11c+MHC-IIhi InfResAMs were found at sites of Pdgfra+ and Pdgfrb+ fibroblast accumulation and were identified as the main source of Pdgf-aa—a growth factor that drives fibroblast proliferation49. In this setting, depletion of CX3CR1+ cells using the tamoxifen-inducible CX3CR1-Cre-ERT2 diphtheria toxin fragment A (DTA) mouse model resulted in decreased fibrosis and reduced collagen deposition49; however, this approach lacks specificity, as both monocytes and peribronchial IMs express CX3CR1.
InfResAMs that developed during fibrosis and ResAMs present before the onset of fibrosis have been shown by bulk RNA sequencing to share a transcriptional signature of 1,842 genes42. In addition, InfResAMs expressed 1,937 genes that were not acquired by ResAMs isolated from the same fibrotic lungs42. Part of the InfResMacs-specific fibrosis-induced signature may be imprinted in monocytes before they differentiate into InfResAMs. Fibrosis was shown to induce the generation of SatM monocytes with pro-fibrotic activity54. Repeated lung injury could also induce trained immunity55,56, rewiring the bone marrow to produce these pro-fibrotic monocytes. InfResAMs are long-lived and persist for at least 10 months in the lungs after the resolution of fibrosis—a time point at which they differentially expressed 330 genes compared with ResAMs isolated from the same lungs42, indicating that differentiation during fibrosis leads to the acquisition of a fibrotic signature not induced in macrophages that were terminally differentiated at the time of the inflammatory event.
The more restricted response of ResAMs during fibrosis may be linked to the environmental regulation of AM metabolism, which has been shown to limit their responses in type 2 settings57. The finding that ResMacs respond less aggressively than monocytes recruited during the inflammatory window is not limited to the lung. ResMacs from the peripheral nerves and liver respond only moderately to inflammation58,59. The limited responsiveness of ResAMs in the healthy lung may be seen as a protective strategy against fibrosis. Minor insults would only result in a small loss of ResAMs, and their proliferative capacity would ensure that the AM pool is mainly reconstituted through the proliferation of the remaining ResAMs without input from recruited monocytes. In contrast, strong pulmonary insults resulting in a marked drop in the number of ResAMs would allow the development of more reactive InfResAMs. The return to pulmonary homeostasis gradually silences the fibrosis-induced genes in InfResAMs and converts them into ResMacs (Fig. 3d), although a small part of this signature is maintained for at least 10 months42. When mice were treated with bleomycin every 2 months and the profile of InfResMacs was analysed 3 weeks after the first and second bleomycin treatments60, the monocyte-derived AMs that had developed during the first bleomycin injury responded similarly to the second bleomycin injury as the ResAMs that were present in the tissue before the first injury. This indicates that the 2 months between the bleomycin treatments was sufficient to convert the InfResMacs that developed during the first bleomycin injury into ResMacs, However, the monocytes recruited to the lung during the second injury gave rise to InfResMacs with a stronger pro-fibrotic profile than the monocytes that gave rise to InfResMacs during the first bleomycin injury60. This suggests that inflammatory memory is not linked to ontogeny itself and that a return to homeostasis for 2 months is sufficient to erase most of the inflammatory signature, but that the environmental cues imprinting the InfResMacs profile during the second injury must be stronger. Therefore, it will be important to study the inflammation-induced epigenetic memory in the cells that represent the AM niche, such as the alveolar epithelial cells, perialveolar interstitial fibroblasts or resident lymphocytes, as these cells may be involved in the imprinting of a stronger fibrosis-induced signature of InfResAMs during the second bleomycin injury.
While the AM pool contracts across a range of fibrotic settings, the IM population as a whole expands61,62. In chimeric mice reconstituted with bone marrow cells from Lyve1Cre/GFPSlco2b1fl/DTR mice, the human diphtheria toxin receptor (DTR) is expressed on perivascular Lyve1hiCD206hi IMs and the administration of diphtheria toxin resulted in an 80% reduction in Lyve1hiCD206hi IMs14. When challenged with bleomycin, these mice suffered greater weight loss, collagen deposition and inflammatory cell infiltration than Lyve1hiCD206hi IM-sufficient mice, indicating that Lyve1hiCD206hi IMs may perform a protective role by repressing the infiltration of immune cells into the fibrotic lung14. However, it should be noted that macrophage depletion itself can lead to fibroblast activation and, as such, may contribute to the observed phenotype23. The field still lacks a system to target CX3CR1hiMHC-Ihi IMs and, as such, the specific role of these cells cannot yet be fully addressed.
Macrophage origin and plasticity during pulmonary infections
AMs represent the first line of defense and play a central role against a range of respiratory pathogens, including influenza and Streptococcus pneumoniae19,63. Yet, pulmonary immune responses need to be contained to avoid excessive tissue damage and safeguard gas exchange. Resident AMs are less responsive than recruited monocyte-derived macrophages in the context of allergy or viral infection64,65. Ccr2−/− mice, which have fewer circulating monocytes66, are less susceptible to influenza infection and display a decreased morbidity and mortality67, even if the viral load is slightly elevated in these mice68. Conversely, mice lacking AMs mount stronger inflammatory responses during Brucella infection69. The anti-inflammatory role of resident AMs may in part be linked to their production of type I interferon (IFN)70 as type I IFNs decrease IL-1 and increase IL-10 production in monocyte-derived macrophages71. However, excessive type I and III IFNs aggravate pulmonary influenza infections by impairing epithelial regeneration72. In addition, while early type I IFN reduces the viral titer and protects mice from severe acute respiratory syndrome coronavirus 1 (SARS-CoV-1)-associated pathology, delayed type I IFN responses lead to higher viral loads, but also to exacerbated activation and recruitment of inflammatory monocytes73. The primary source of the excessive type I IFN response in SARS-CoV-1 infections has been proposed to be plasmacytoid dendritic cells, not AMs. AMs may also have a tolerogenic activity on lung epithelial cells via cell–cell delivery of suppressor of cytokine signaling proteins74 and connexin-43-dependent intercellular communication75. This conditioning may be reciprocal, as surfactant proteins produced by epithelial cells can in turn suppress the immune response of AMs to Toll-like receptor ligands76,77,78. However, the role of surfactant proteins can be varied and context dependent. Surfactant protein A was described as a local amplifier of various aspects of a type 2 response in AMs79, while surfactant protein D seems to have no role in regulating IL-4-dependent responses and proliferation57.
AMs have low expression of complement-associated genes such as C1qa, C1qb, C1qc, C2, C4b and C3ar1 (ref. 15), while macrophages isolated from other tissues have high expression of these genes across multiple tissues (unpublished data and www.immgen.org). Signals from the AM niche may be responsible for the active suppression of these genes. Interestingly, epigenetic profiling of AMs during inflammation has shown that while the C1q locus is inaccessible in ResAMs and remains closed upon influenza infection, complement genes are highly accessible in InfResAMs that have engrafted in the AM pool during influenza infection13. This could have important clinical implications as, during human SARS-Cov-2 infection, recruited macrophages were C1Qhi, compared with the C1Qlo ResAMs80, suggesting that the suppression of the complement-associated genes in ResAMs is conserved in humans. Importantly, the overt activation of complement has been linked to coronavirus 2019 (COVID-19) severity, prompting the launch of clinical trials aiming to block components of the complement pathway81,82. It is tempting to speculate that the sharp drop in ResAMs and their replacement by recruited macrophages expressing high levels of the complement-associated genes, which occurs in patients with severe COVID-19, may fuel the complement cascade and actively participate in the COVID-19 pathology. In fact, it may even be that the suppression of these complement genes in ResAMs is the result of evolutionary pressure to protect the lungs from complement-mediated collateral damage during viral infections.
Bacteria and viruses have developed strategies to co-opt AMs, taking advantage of their relative hyporesponsiveness and enabling their persistence and dissemination within the tissue19,63. The relocation of Mycobacterium tuberculosis-infected AMs into the pulmonary interstitium was shown to be a key step preceding bacterial dissemination83. Following viral infection, the AM population is depleted and then restored by monocyte engraftment into the AM niche and their differentiation into long-lived InfResAMs42,43. The extent of monocyte engraftment is determined by the level of AM depletion, which in turn is linked to the dose and/or virulence of the pathogen. This viral-induced reduction in AM numbers renders mice temporarily more susceptible to bacterial infections84. If only very few AMs are lost as a result of an infection, the proliferation of resident cells will outcompete the engraftment of monocytes. These variables, as well as differences between animal housing, may explain why in some studies monocytes did not contribute to the AM pool following viral infection10,85, while in other studies they were shown to contribute to the AM pool13,42,43. InfResAMs that developed during an infection have been shown to differ functionally from the ResAMs that were present before the infection13,43. Pulmonary infection with murid herpesvirus 4 resulted in the replacement of 90% of ResAMs with InfResAMs, which persisted after the infection had been cleared and protected the mice from house dust mite-driven allergic asthma43. Adoptive transfer studies showed that the InfResAMs from murid herpesvirus 4-infected mice could transfer protection against asthma to uninfected recipients, underlining the AM-intrinsic reprogramming. The fact that these InfResAMs developed during inflammation may not be the only explanation for their altered phenotype. Viral infection also affects bone marrow-derived monocyte production and is associated with the production of MHC-II+Sca1+ regulatory monocytes that produce high amounts of IL-10 (ref. 43). Unfortunately, the remaining ResAMs that survived the herpes virus infection (10% of total) were not compared with the InfResMacs in this study43, and it remains unknown whether these cells were equally impacted by the infection. A comparison of ResAMs that were present before influenza infection with InfResAMs that developed during influenza infection13 revealed InfResAMs to be more responsive to Toll-like receptor ligands or subsequent S. pneumoniae infection than ResAMs. The increased IL-6 production by InfResAMs represents an innate memory that provides protection to subsequent S. pneumonia infection. The epigenetic landscape of these ResAMs and InfResAMs indicated that monocytes entering the influenza virus-infected lungs gained an epigenetic profile associated with tissue residency, but also acquired an inflammatory signature imprinted by the antiviral cues. As a result, the IL-6 enhancer regions were more accessible in InfResAMs that had developed during the influenza infection than in ResAMs that were present before the onset of infection in the same lungs (Fig. 3b). This altered epigenetic landscape was still evident 1 month after the clearance of the infection but was dampened at 2 months post-infection. Although we have focused here on the differences in responsiveness between InfResMacs and ResMacs, it is important to note that ResMacs can respond to inflammatory stimuli. In fact, both InfResAMs and ResAMs produce comparable amounts of tumor necrosis factor after in vitro stimulation with Pam3CSK4 (ref. 13) (Fig. 3b). Long-term memory characterized by altered gene expression, metabolism and antimicrobial responsiveness has been proposed to also occur in ResAMs, as was reported in an adenovirus infection model85. The priming of this phenotype is dependent on CD8+ T cells in an IFN-γ- and cell–cell contact-dependent manner85. The phagocytic capacity of ResAMs can also be influenced by a previous infection, which dampens their phagocytic capacity through the overexpression of signal regulatory protein α86. ResAMs with impaired phagocytosis recovered their capabilities when transferred into naive mice, and naive ResAMs became paralyzed when transferred into the lungs of recently infected mice, pointing towards a dominant role of the niche in shaping macrophage function.
Altogether, these findings indicate that both ResAMs that were present before an infection and InfResAMs that develop during an infection can acquire an innate memory, altering their response to subsequent infections. However, the InfResMacs that develop during an infection are epigenetically more plastic and, as such, infection has a greater impact on them. An assessment of AM and IM responses to in vivo lipopolysaccharide (LPS) stimulation40 revealed that IMs upregulated around 1,000 genes, while AMs upregulated 250 genes. Only 50 upregulated genes were common to both populations, including Tnf and Il1b. Analysis of the epigenetic profile of these genes revealed that some of the differences in expression could be linked to differing chromatin accessibility between IMs and AMs. However, for an important fraction of these genes, AMs and IMs had comparable patterns of open chromatin landscapes. These data suggest the presence of distinct transcriptional activators and repressors in IMs and AMs40.
Much less is known about the effect of residency on the plasticity of IMs during infections. It should also be noted that IMs have a shorter half-life compared with AMs14. A robust IM fate-mapping system will be required to conclusively distinguish IMs that were present before the infection from IMs that developed during the infection in order to address these questions.
Resetting the tissue memory clock by manipulating InfResMacs
The ability to distinguish pulmonary ResMacs and InfResMacs revealed InfResMacs to be more strongly imprinted across a range of inflammatory events. This inflammatory imprinting has the capacity to generate a long-lasting memory that will alter the macrophage function during subsequent challenges. Multiple pulmonary infections during life may therefore allow the engraftment of successive waves of InfResMacs. If sufficient time passes between insults, the lung will return to homeostasis and the InfResMac signature will be progressively lost. However, recurrent infections or exposure to toxic agents may fuel the memory of InfResMacs. As such, the duration of residence in a homeostatic environment becomes a key parameter of ResMac biology, as others have recently independently proposed87,88. Innate memory in InfResMacs may be beneficial in some cases and may confer resistance to subsequent infections, but this may also lead to exacerbated tissue damage89. Here, we propose that the increased reactivity of InfResMacs to inflammation is not due to their monocytic origin, but rather to their differentiation during inflammation.
Although currently speculative, this revisited plasticity model offers therapeutic opportunities (Fig. 4). Depletion of ResMacs after a profound inflammatory episode would allow a new wave of monocytes to engraft in the pool of ResMacs resident in tissue. As these cells would develop in the absence of overt inflammation, this approach should yield ResMacs that acquire a normal steady-state profile (Fig. 4a). Treatment with CSF1 receptor antibodies has been shown to deplete macrophages in patients90. It remains to be seen if treatment with CFS1 receptor antibodies in patients who have recovered from severe fibrosis would result in the elimination of long-lived InfResMacs and protect these patients from relapsing. To be successful, such a strategy would require that the niche cells have also recovered sufficiently to imprint a steady-state signature on the repopulating monocytes.
a, Depletion of InfResMacs may allow a fresh wave of monocytes to engraft in the ResMac pool. The development of these monocytes in the absence of overt inflammation could yield ResMacs that acquire the normal steady-state profile. b, Blocking the inflammatory niche circuits that maintain the InfResMac identity could allow the conversion of InfResMacs into ResMacs under the influence of steady-state circuits.
This revisited plasticity model also has implications for cancer therapy where CSF1 receptor blocking agents are currently undergoing clinical trials91. In some patients, the global immunosuppressive effects of cancer could imprint a suppressed state on circulating monocytes. In this setting, CSF1 receptor blockade could empty macrophage niches throughout the body, allowing the immune-suppressed monocytes to colonize the ResMac pools. If the immune-suppressed state is maintained in these macrophages for a prolonged period of time, it could influence the risk of metastasis or opportunistic infections in these organs.
A safer option may be to manipulate the cell–cell circuits that control the steady-state imprinting, rather than depleting macrophages (Fig. 4b). Macrophages constantly sense the environmental signals in their niche and rapidly lose their tissue-specific gene signature when put in culture3,92,93,94. Therefore, silencing of particular inflammatory genes may also require constant tissue signals. These are probably the same signals that convert InfResMacs into ResMacs during the resolution phase and could form the basis for therapeutic strategies seeking to accelerate a return to homeostasis. It may also be that the long-lived InfResMac profile is actively maintained by signals produced by activated niche cells. One could easily imagine pathogenic circuits fueling the reciprocal activation between fibroblasts and InfResMacs. Understanding the nature of these long-lived pathogenic circuits may provide therapeutic strategies that could breakdown these circuits and allow a quicker restoration of the steady-state profile for the macrophages and their niche. This could be one of the mechanisms underlying the beneficial effects of prolonged treatment with anti-IL-1β antibodies (canakinumab) observed during the Canakinumab Anti-Inflammatory Thrombosis Outcome Study (CANTOS) trial. The trial showed that long-term anti-inflammatory intervention in patients with myocardial infarction not only resulted in a reduced risk for cardiovascular disease but was also beneficial for autoimmune arthritis and cancer95. It is possible that such a prolonged anti-inflammatory treatment allows the InfResMacs to revert to the homeostatic ResMac profile.
The revised plasticity model could also have clinical implications for COVID-19. Patients severely affected by SARS-CoV-2 undergo a massive loss of ResAMs80. If monocytes were to engraft as InfResAMs during the SARS-CoV-2 infection, it is conceivable that these cells would have a strong epigenetic memory of the SARS-CoV-2-associated cytokine storm and, as a result, this may put these patients at risk of developing secondary inflammatory diseases. Finding a way to accelerate the return of InfResAMs and their niche to homeostasis could potentially provide a promising therapeutic avenue to protect these patients from secondary complications.
Concluding remarks
Altogether, the recent findings obtained through the combination of single-cell technologies, epigenetics and fate mapping call for a thorough adjustment of the macrophage plasticity concept. The available data suggest that prolonged residency in tissues shuts down much of the plasticity of macrophages. Together with the fact that tissues endow macrophages with strong self-maintenance capacity, this could represent an important mechanism by which tissues avoid excessive immune reactions and protect themselves from collateral damage. It appears that severe inflammatory episodes induce the generation of InfResMacs that maintain a more inflammatory profile for months. It is hoped that a better understanding of the cell–cell circuits that control the fate of InfResMacs will provide therapeutic opportunities to reprogram the ResMac pool and accelerate the return to homeostasis.
References
Wynn, T. A., Chawla, A. & Pollard, J. W. Macrophage biology in development, homeostasis and disease. Nature 496, 445–455 (2013).
Gautier, E. L. et al. Gene-expression profiles and transcriptional regulatory pathways that underlie the identity and diversity of mouse tissue macrophages. Nat. Immunol. 13, 1118–1128 (2012).
Gosselin, D. et al. Environment drives selection and function of enhancers controlling tissue-specific macrophage identities. Cell 159, 1327–1340 (2014).
Lavin, Y. et al. Tissue-resident macrophage enhancer landscapes are shaped by the local microenvironment. Cell 159, 1312–1326 (2014).
Okabe, Y. & Medzhitov, R. Tissue biology perspective on macrophages. Nat. Immunol. 17, 9–17 (2016).
Scott, C. L. et al. Bone marrow-derived monocytes give rise to self-renewing and fully differentiated Kupffer cells. Nat. Commun. 7, 10321 (2016).
Shemer, A. et al. Engrafted parenchymal brain macrophages differ from microglia in transcriptome, chromatin landscape and response to challenge. Nat. Commun. 9, 5206 (2018).
Van de Laar, L. et al. Yolk sac macrophages, fetal liver, and adult monocytes can colonize an empty niche and develop into functional tissue-resident macrophages. Immunity 44, 755–768 (2016).
Ajami, B., Bennett, J. L., Krieger, C., Tetzlaff, W. & Rossi, F. M. V. Local self-renewal can sustain CNS microglia maintenance and function throughout adult life. Nat. Neurosci. 10, 1538–1543 (2007).
Hashimoto, D. et al. Tissue-resident macrophages self-maintain locally throughout adult life with minimal contribution from circulating monocytes. Immunity 38, 792–804 (2013).
Liu, Z. et al. Fate mapping via Ms4a3-expression history traces monocyte-derived cells. Cell 178, 1509–1525.e19 (2019).
Sawai, C. M. et al. Hematopoietic stem cells are the major source of multilineage hematopoiesis in adult animals. Immunity 45, 597–609 (2016).
Aegerter, H. et al. Influenza-induced monocyte-derived alveolar macrophages confer prolonged antibacterial protection. Nat. Immunol. https://doi.org/10.1038/s41590-019-0568-x (2020).
Chakarov, S. et al. Two distinct interstitial macrophage populations coexist across tissues in specific subtissular niches. Science 363, eaau0964 (2019).
Gibbings, S. L. et al. Three unique interstitial macrophages in the murine lung at steady state. Am. J. Respir. Cell Mol. Biol. 57, 66–76 (2017).
Schyns, J. et al. Non-classical tissue monocytes and two functionally distinct populations of interstitial macrophages populate the mouse lung. Nat. Commun. 10, 3964 (2019).
Guilliams, M. & Scott, C. L. Does niche competition determine the origin of tissue-resident macrophages? Nat. Rev. Immunol. 17, 451–460 (2017).
Guilliams, M. et al. Alveolar macrophages develop from fetal monocytes that differentiate into long-lived cells in the first week of life via GM-CSF. J. Exp. Med. 210, 1977–1992 (2013).
Schneider, C. et al. Alveolar macrophages are essential for protection from respiratory failure and associated morbidity following influenza virus infection. PLoS Pathog. 10, e1004053 (2014).
Trapnell, B. C. et al. Pulmonary alveolar proteinosis. Nat. Rev. Dis. Primers 5, 16 (2019).
Yu, X. et al. The cytokine TGF-β promotes the development and homeostasis of alveolar macrophages. Immunity 47, 903–912.e4 (2017).
Trapnell, B. C., Whitsett, J. A. & Nakata, K. Pulmonary alveolar proteinosis. N. Engl. J. Med. 349, 2527–2539 (2003).
Bonnardel, J. et al. Stellate cells, hepatocytes, and endothelial cells imprint the Kupffer cell identity on monocytes colonizing the liver macrophage niche. Immunity 51, 638–654.e9 (2019).
Ural, B. B. et al. Identification of a nerve-associated, lung-resident interstitial macrophage subset with distinct localization and immunoregulatory properties. Sci. Immunol. 5, eaax8756 (2020).
Bedoret, D. et al. Lung interstitial macrophages alter dendritic cell functions to prevent airway allergy in mice. J. Clin. Invest. 119, 3723–3738 (2009).
Sabatel, C. et al. Exposure to bacterial CpG DNA protects from airway allergic inflammation by expanding regulatory lung interstitial macrophages. Immunity 46, 457–473 (2017).
Muller, P. A. et al. Crosstalk between muscularis macrophages and enteric neurons regulates gastrointestinal motility. Cell 158, 300–313 (2014).
Schepper, S. D. et al. Self-maintaining gut macrophages are essential for intestinal homeostasis. Cell 175, 400–415.e13 (2018).
Kolter, J. et al. A subset of skin macrophages contributes to the surveillance and regeneration of local nerves. Immunity 50, 1482–1497.e7 (2019).
Ginhoux, F. & Guilliams, M. Tissue-resident macrophage ontogeny and homeostasis. Immunity 44, 439–449 (2016).
McGrath, K. E., Frame, J. M. & Palis, J. Early hematopoiesis and macrophage development. Semin. Immunol. 27, 379–387 (2015).
Perdiguero, E. G. & Geissmann, F. The development and maintenance of resident macrophages. Nat. Immunol. 17, 2–8 (2016).
Gibbings, S. L. et al. Transcriptome analysis highlights the conserved difference between embryonic and postnatal-derived alveolar macrophages. Blood 126, 1357–1366 (2015).
Yona, S. et al. Fate mapping reveals origins and dynamics of monocytes and tissue macrophages under homeostasis. Immunity 38, 79–91 (2013).
Hoeffel, G. et al. C-Myb+ erythro-myeloid progenitor-derived fetal monocytes give rise to adult tissue-resident macrophages. Immunity 42, 665–678 (2015).
Cohen, M. et al. Lung single-cell signaling interaction map reveals basophil role in macrophage imprinting. Cell 175, 1031–1044.e18 (2018).
Tan, S. Y. S. & Krasnow, M. A. Developmental origin of lung macrophage diversity. Dev. Camb. Engl. 143, 1318–1327 (2016).
Schulz, C. et al. A lineage of myeloid cells independent of Myb and hematopoietic stem cells. Science 336, 86–90 (2012).
Scott, C. L. et al. The transcription factor ZEB2 is required to maintain the tissue-specific identities of macrophages. Immunity 49, 312–325.e5 (2018).
Sajti, E. et al. Transcriptomic and epigenetic mechanisms underlying myeloid diversity in the lung. Nat. Immunol. https://doi.org/10.1038/s41590-019-0582-z (2020).
Sieweke, M. H. Waddington’s valleys and Captain Cook’s islands. Cell Stem Cell 16, 7–8 (2015).
Misharin, A. V. et al. Monocyte-derived alveolar macrophages drive lung fibrosis and persist in the lung over the life span. J. Exp. Med. 214, 2387–2404 (2017).
Machiels, B. et al. A gammaherpesvirus provides protection against allergic asthma by inducing the replacement of resident alveolar macrophages with regulatory monocytes. Nat. Immunol. 18, 1310–1320 (2017).
De Kleer, I. M. et al. Perinatal activation of the interleukin-33 pathway promotes type 2 immunity in the developing lung. Immunity 45, 1285–1298 (2016).
Camara, A. et al. Lymph node mesenchymal and endothelial stromal cells cooperate via the RANK–RANKL cytokine axis to shape the sinusoidal macrophage niche. Immunity 50, 1467–1481.e6 (2019).
Schneider, C. et al. Induction of the nuclear receptor PPAR-γ by the cytokine GM-CSF is critical for the differentiation of fetal monocytes into alveolar macrophages. Nat. Immunol. 15, 1026–1037 (2014).
Polosukhin, V. V. et al. Intratracheal bleomycin causes airway remodeling and airflow obstruction in mice. Exp. Lung Res. 38, 135–146 (2012).
Duffield, J. S., Lupher, M., Thannickal, V. J. & Wynn, T. A. Host responses in tissue repair and fibrosis. Annu. Rev. Pathol. Mech. Dis. 8, 241–276 (2013).
Aran, D. et al. Reference-based analysis of lung single-cell sequencing reveals a transitional profibrotic macrophage. Nat. Immunol. 20, 163–172 (2019).
Joshi, N. et al. A spatially restricted fibrotic niche in pulmonary fibrosis is sustained by M-CSF/M-CSFR signalling in monocyte-derived alveolar macrophages. Eur. Respir. J. https://doi.org/10.1183/13993003.00646-2019 (2019).
Aoshiba, K., Tsuji, T. & Nagai, A. Bleomycin induces cellular senescence in alveolar epithelial cells. Eur. Respir. J. 22, 436–443 (2003).
Tanjore, H. et al. Contribution of epithelial-derived fibroblasts to bleomycin-induced lung fibrosis. Am. J. Respir. Crit. Care 180, 657–665 (2009).
Wynn, T. A. & Ramalingam, T. R. Mechanisms of fibrosis: therapeutic translation for fibrotic disease. Nat. Med. 18, 1028–1040 (2012).
Satoh, T. et al. Identification of an atypical monocyte and committed progenitor involved in fibrosis. Nature 541, 96–101 (2016).
Netea, M. G. et al. Defining trained immunity and its role in health and disease. Nat. Rev. Immunol. 20, 375–388 (2020).
Guilliams, M., Mildner, A. & Yona, S. Developmental and functional heterogeneity of monocytes. Immunity 49, 595–613 (2018).
Svedberg, F. R. et al. The lung environment controls alveolar macrophage metabolism and responsiveness in type 2 inflammation. Nat. Immunol. 20, 571–580 (2019).
Ydens, E. et al. Profiling peripheral nerve macrophages reveals two macrophage subsets with distinct localization, transcriptome and response to injury. Nat. Neurosci. 23, 676–689 (2020).
Remmerie, A. et al. Osteopontin expression identifies a subset of recruited macrophages distinct from Kupffer cells in the fatty liver. Immunity 53, 641–657.e14 (2020).
McQuattie-Pimentel, A. C. et al. The aging microenvironment shapes alveolar macrophage identity in aging. Preprint at bioRxiv https://doi.org/10.1101/717033 (2019).
Meziani, L. et al. CSF1R inhibition prevents radiation pulmonary fibrosis by depletion of interstitial macrophages. Eur. Respir. J. 51, 1702120 (2018).
Misharin, A. V., Morales-Nebreda, L., Mutlu, G. M., Budinger, G. R. S. & Perlman, H. Flow cytometric analysis of macrophages and dendritic cell subsets in the mouse lung. Am. J. Respir. Cell Mol. Biol. 49, 503–510 (2013).
Knapp, S. et al. Alveolar macrophages have a protective antiinflammatory role during murine pneumococcal pneumonia. Am. J. Resp. Crit. Care 167, 171–179 (2003).
Naessens, T. et al. Innate imprinting of murine resident alveolar macrophages by allergic bronchial inflammation causes a switch from hypoinflammatory to hyperinflammatory reactivity. Am. J. Pathol. 181, 174–184 (2012).
Van Riel, D. et al. Highly pathogenic avian influenza virus H5N1 infects alveolar macrophages without virus production or excessive TNF-alpha induction. PLoS Pathog. 7, e1002099 (2011).
Serbina, N. V. & Pamer, E. G. Monocyte emigration from bone marrow during bacterial infection requires signals mediated by chemokine receptor CCR2. Nat. Immunol. 7, 311–317 (2006).
Lin, K. L., Suzuki, Y., Nakano, H., Ramsburg, E. & Gunn, M. D. CCR2+ monocyte-derived dendritic cells and exudate macrophages produce influenza-induced pulmonary immune pathology and mortality. J. Immunol. 180, 2562–2572 (2008).
Dawson, T. C., Beck, M. A., Kuziel, W. A., Henderson, F. & Maeda, N. Contrasting effects of CCR5 and CCR2 deficiency in the pulmonary inflammatory response to influenza A virus. Am. J. Pathol. 156, 1951–1959 (2000).
Archambaud, C. et al. Contrasting roles of macrophages and dendritic cells in controlling initial pulmonary Brucella infection. Eur. J. Immunol. 40, 3458–3471 (2010).
Kumagai, Y. et al. Alveolar macrophages are the primary interferon-α producer in pulmonary infection with RNA viruses. Immunity 27, 240–252 (2007).
Guarda, G. et al. Type I interferon inhibits interleukin-1 production and inflammasome activation. Immunity 34, 213–223 (2011).
Major, J. et al. Type I and III interferons disrupt lung epithelial repair during recovery from viral infection. Science https://doi.org/10.1126/science.abc2061 (2020).
Channappanavar, R. et al. Dysregulated type I interferon and inflammatory monocyte–macrophage responses cause lethal pneumonia in SARS-CoV-infected mice. Cell Host Microbe 19, 181–193 (2016).
Bourdonnay, E. et al. Transcellular delivery of vesicular SOCS proteins from macrophages to epithelial cells blunts inflammatory signaling. J. Exp. Med. 212, 729–742 (2015).
Westphalen, K. et al. Sessile alveolar macrophages communicate with alveolar epithelium to modulate immunity. Nature 506, 503–506 (2014).
Haczku, A. Protective role of the lung collectins surfactant protein A and surfactant protein D in airway inflammation. J. Allergy Clin. Immunol. 122, 861–879 (2008).
Sano, H. et al. Pulmonary surfactant protein A modulates the cellular response to smooth and rough lipopolysaccharides by interaction with CD14. J. Immunol. 163, 387–395 (1999).
Yamada, C. et al. Surfactant protein A directly interacts with TLR4 and MD-2 and regulates inflammatory cellular response. J. Biol. Chem. 281, 21771–21780 (2006).
Minutti, C. M. et al. Local amplifiers of IL-4Rα–mediated macrophage activation promote repair in lung and liver. Science 356, 1076–1080 (2017).
Bost, P. et al. Host–viral infection maps reveal signatures of severe COVID-19 patients. Cell 181, 1475–1488.e12 (2020).
Gao, T. et al. Highly pathogenic coronavirus N protein aggravates lung injury by MASP-2-mediated complement over-activation. Preprint at medRxiv https://doi.org/10.1101/2020.03.29.20041962 (2020).
Carvelli, J. et al. Association of COVID-19 inflammation with activation of the C5a–C5aR1 axis. Nature 588, 146–150 (2020).
Cohen, S. B. et al. Alveolar macrophages provide an early Mycobacterium tuberculosis niche and initiate dissemination. Cell Host Microbe 24, 439–446.e4 (2018).
Ghoneim, H. E., Thomas, P. G. & McCullers, J. A. Depletion of alveolar macrophages during influenza infection facilitates bacterial superinfections. J. Immunol. 191, 1250–1259 (2013).
Yao, Y. et al. Induction of autonomous memory alveolar macrophages requires T cell help and is critical to trained immunity. Cell 175, 1634–1650.e17 (2018).
Roquilly, A. et al. Alveolar macrophages are epigenetically altered after inflammation, leading to long-term lung immunoparalysis. Nat. Immunol. 21, 636–648 (2020).
Blériot, C., Chakarov, S. & Ginhoux, F. Determinants of resident tissue macrophage identity and function. Immunity 52, 957–970 (2020).
Kulikauskaite, J. & Wack, A. Teaching old dogs new tricks? The plasticity of lung alveolar macrophage subsets. Trends Immunol. 41, 864–877 (2020).
Goulding, J. et al. Respiratory infections: do we ever recover? Proc. Am. Thorac. Soc. 4, 618–625 (2007).
Ries, C. H. et al. Targeting tumor-associated macrophages with anti-CSF-1R antibody reveals a strategy for cancer therapy. Cancer Cell 25, 846–859 (2014).
Wainberg, Z. A. et al. Phase 1/2a study of double immune suppression blockade by combining a CSF1R inhibitor (pexidartinib/PLX3397) with an anti PD-1 antibody (pembrolizumab) to treat advanced melanoma and other solid tumors. J. Clin. Oncol. 34, TPS465 (2016).
Gosselin, D. et al. An environment-dependent transcriptional network specifies human microglia identity. Science 356, eaal3222 (2017).
Bohlen, C. J. et al. Diverse requirements for microglial survival, specification, and function revealed by defined-medium cultures. Neuron 94, 759–773.e8 (2017).
Bennett, F. C. et al. A combination of ontogeny and CNS environment establishes microglial identity. Neuron 98, 1170–1183.e8 (2018).
Ridker, P. M. et al. Antiinflammatory therapy with canakinumab for atherosclerotic disease. N. Engl. J. Med. 377, 1119–1131 (2017).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Ioana Visan was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Guilliams, M., Svedberg, F.R. Does tissue imprinting restrict macrophage plasticity?. Nat Immunol 22, 118–127 (2021). https://doi.org/10.1038/s41590-020-00849-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-020-00849-2
This article is cited by
-
The role of macrophage polarization and cellular crosstalk in the pulmonary fibrotic microenvironment: a review
Cell Communication and Signaling (2024)
-
Joint-specific memory, resident memory T cells and the rolling window of opportunity in arthritis
Nature Reviews Rheumatology (2024)
-
Mineralocorticoid receptor promotes cardiac macrophage inflammaging
Basic Research in Cardiology (2024)
-
Immune stimulation recruits a subset of pro-regenerative macrophages to the retina that promotes axonal regrowth of injured neurons
Acta Neuropathologica Communications (2023)
-
Kupffer cells prevent pancreatic ductal adenocarcinoma metastasis to the liver in mice
Nature Communications (2023)