Abstract
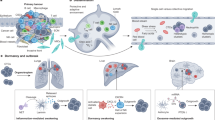
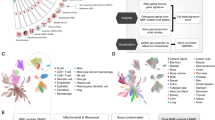
Intratumor heterogeneity underlies cancer evolution and treatment resistance, but targetable mechanisms driving intratumor heterogeneity are poorly understood. Meningiomas are the most common primary intracranial tumors and are resistant to all medical therapies, and high-grade meningiomas have significant intratumor heterogeneity. Here we use spatial approaches to identify genomic, biochemical and cellular mechanisms linking intratumor heterogeneity to the molecular, temporal and spatial evolution of high-grade meningiomas. We show that divergent intratumor gene and protein expression programs distinguish high-grade meningiomas that are otherwise grouped together by current classification systems. Analyses of matched pairs of primary and recurrent meningiomas reveal spatial expansion of subclonal copy number variants associated with treatment resistance. Multiplexed sequential immunofluorescence and deconvolution of meningioma spatial transcriptomes using cell types from single-cell RNA sequencing show decreased immune infiltration, decreased MAPK signaling, increased PI3K–AKT signaling and increased cell proliferation, which are associated with meningioma recurrence. To translate these findings to preclinical models, we use CRISPR interference and lineage tracing approaches to identify combination therapies that target intratumor heterogeneity in meningioma cell co-cultures.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
DNA sequencing and spatial transcriptomic data that support the findings of this study have been deposited in the Sequence Read Archive (https://www.ncbi.nlm.nih.gov/sra) under BioProject ID PRJNA950017. DNA methylation and RNA sequencing data that support the findings of this study have been deposited in the NCBI Gene Expression Omnibus (https://www.ncbi.nlm.nih.gov/geo) under accession numbers GSE228316 (DNA methylation) and GSE228433 (RNA sequencing). The publicly available GRCh38 (hg38) and CRCh37.p13 (hg19) were used in this study. Spatial proteomic data processed counts are provided in Supplementary Table 5. Source data are provided with this paper.
Code availability
No custom software, tools or packages were used. The open source software, tools and packages used for data analysis in this study are referenced in Methods where applicable and include Loupe Browser (10x Genomics, v6.3.0), Space Ranger (10x Genomics, v1.0), 2100 Bioanalyzer Expert Software (vB.02.10), Aggr (v2.0.0), AutoAnnotate (v1.3.5), Burrows–Wheeler aligner (v0.7.17), Clustree (v0.5.0), COMET Control Software (v1.0), Cutadapt (v3.7), Cytoscape (v3.7.2), DESeq2 (v1.36.0), EnrichmentMap (v3.3.4), FASTQC (v0.11.9), featureCounts (v2.0.0), Genome Analysis Toolkit (v4.3.0.0), GSEA (v4.3.2), Harmony (v0.1.1), HISAT2 (v2.2.0), Integrative Genome Viewer (v.2.16.0), inferCNV (1 v.14.0), MultiQC (v1.12), NxClinical (v6.0), Olympus cellSens Standard Imaging (v1.16), Picard (v2.27.5), R (v3.6.1 and v4.2.1), RStudio (v2022.07.02), SeSAMe (v3.10), Seurat (v4.3.0), spatialInferCNV (v0.1.0), SCDC (v 0.0.0.9000) and SpatialFeatureplot (Seurat v3). Code written for this project is available on GitHub (https://doi.org/10.5281/zenodo.10927737)66.
References
Ostrom, Q. T. et al. CBTRUS statistical report: primary brain and other central nervous system tumors diagnosed in the United States in 2015–2019. Neuro. Oncol. 24, v1–v95 (2022).
Goldbrunner, R. et al. EANO guideline on the diagnosis and management of meningiomas. Neuro. Oncol. 23, 1821–1834 (2021).
Lin, D. et al. Trends in intracranial meningioma incidence in the United States, 2004‐2015. Cancer Med. 8, 6458–6467 (2019).
Sahm, F. et al. DNA methylation-based classification and grading system for meningioma: a multicentre, retrospective analysis. Lancet Oncol. 18, 682–694 (2017).
Olar, A. et al. Global epigenetic profiling identifies methylation subgroups associated with recurrence-free survival in meningioma. Acta Neuropathol. 133, 431–444 (2017).
Choudhury, A. et al. Meningioma DNA methylation groups identify biological drivers and therapeutic vulnerabilities. Nat. Genet. 54, 649–659 (2022).
Patel, A. J. et al. Molecular profiling predicts meningioma recurrence and reveals loss of DREAM complex repression in aggressive tumors. Proc. Natl Acad. Sci. USA 116, 21715–21726 (2019).
Driver, J. et al. A molecularly integrated grade for meningioma. Neuro. Oncol. 24, 796–808 (2021).
Maas, S. L. N. et al. Integrated molecular-morphologic meningioma classification: a multicenter retrospective analysis, retrospectively and prospectively validated. J. Clin. Oncol. 39, 3839–3852 (2021).
Nassiri, F. et al. A clinically applicable integrative molecular classification of meningiomas. Nature 597, 119–125 (2021).
Choudhury, A. et al. Hypermitotic meningiomas harbor DNA methylation subgroups with distinct biological and clinical features. Neuro. Oncol. 25, 520–530 (2022).
Vasudevan, H. N. et al. Intratumor and informatic heterogeneity influence meningioma molecular classification. Acta Neuropathol. 144, 579–583 (2022).
Magill, S. T. et al. Multiplatform genomic profiling and magnetic resonance imaging identify mechanisms underlying intratumor heterogeneity in meningioma. Nat. Commun. 11, 1–15 (2020).
Louis, D. N. et al. The 2021 WHO classification of tumors of the central nervous system: a summary. Neuro. Oncol. 23, 1231–1251 (2021).
Chen, W. C. et al. Radiotherapy for meningiomas. J. Neurooncol. 160, 505–515 (2022).
Brastianos, P. K. et al. Advances in multidisciplinary therapy for meningiomas. Neuro. Oncol. 21, i18–i31 (2019).
Wen, P. Y., Quant, E., Drappatz, J., Beroukhim, R. & Norden, A. D. Medical therapies for meningiomas. J. Neurooncol. 99, 365–378 (2010).
Preusser, M. et al. Trabectedin for recurrent WHO grade 2 or 3 meningioma: a randomized phase II study of the EORTC Brain Tumor Group (EORTC-1320-BTG). Neuro. Oncol. 24, 755–767 (2021).
Bi, W. L. et al. Activity of PD-1 blockade with nivolumab among patients with recurrent atypical/anaplastic meningioma: phase II trial results. Neuro. Oncol. 24, 101–113 (2021).
Brastianos, P. K. et al. Phase 2 study of pembrolizumab in patients with recurrent and residual high-grade meningiomas. Nat. Commun. 13, 1325 (2022).
Chen, W. C. et al. Targeted gene expression profiling predicts meningioma outcomes and radiotherapy responses. Nat. Med. 29, 3067–3076 (2023).
Gilbert, L. A. et al. Genome-scale CRISPR-mediated control of gene repression and activation. Cell 159, 647–661 (2014).
Kline, C. N. et al. Targeted next-generation sequencing of pediatric neuro-oncology patients improves diagnosis, identifies pathogenic germline mutations, and directs targeted therapy. Neuro. Oncol. 9, 699–709 (2016).
Youngblood, M. W. et al. Correlations between genomic subgroup and clinical features in a cohort of more than 3000 meningiomas. J. Neurosurg. 133, 1345–1354 (2019).
Sahm, F. et al. TERT promoter mutations and risk of recurrence in meningioma. J. Natl Cancer Inst. 108, djv377 (2016).
Guyot, A. et al. Analysis of CDKN2A gene alterations in recurrent and non-recurrent meningioma. J. Neurooncol. 145, 449–459 (2019).
Youngblood, M. W. et al. Associations of meningioma molecular subgroup and tumor recurrence. Neuro. Oncol. 23, 783–794 (2021).
Chen, W. C. et al. Histopathological features predictive of local control of atypical meningioma after surgery and adjuvant radiotherapy. J. Neurosurg. 130, 443–450 (2018).
Tang, V. et al. Loss of p16 expression is a sensitive marker of CDKN2A homozygous deletion in malignant meningiomas. Acta Neuropathol. 145, 497–500 (2023).
Katz, L. M. et al. Loss of histone H3K27me3 identifies a subset of meningiomas with increased risk of recurrence. Acta Neuropathol. 135, 955–963 (2018).
Ravi, V. M. et al. Spatially resolved multi-omics deciphers bidirectional tumor-host interdependence in glioblastoma. Cancer Cell 40, 639–655.e13 (2022).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Couly, G. F. & Douarin, N. M. L. Mapping of the early neural primordium in quail-chick chimeras: II. The prosencephalic neural plate and neural folds: implications for the genesis of cephalic human congenital abnormalities. Dev. Biol. 120, 198–214 (1987).
Simões-Costa, M. & Bronner, M. E. Establishing neural crest identity: a gene regulatory recipe. Development 142, 242–257 (2015).
Merritt, C. R. et al. Multiplex digital spatial profiling of proteins and RNA in fixed tissue. Nat. Biotechnol. 38, 586–599 (2020).
Shankar, G. M. et al. Germline and somatic BAP1 mutations in high-grade rhabdoid meningiomas. Neuro. Oncol. 19, 535–545 (2017).
Vasudevan, H. N. et al. Comprehensive molecular profiling identifies FOXM1 as a key transcription factor for meningioma proliferation. Cell Rep. 22, 3672–3683 (2018).
Mangelinck, A. et al. MT2A is an early predictive biomarker of response to chemotherapy and a potential therapeutic target in osteosarcoma. Sci. Rep. 9, 12301 (2019).
Borchert, S. et al. Impact of metallothionein-knockdown on cisplatin resistance in malignant pleural mesothelioma. Sci. Rep. 10, 18677 (2020).
Patel, A. P. et al. Single-cell RNA-seq highlights intratumoral heterogeneity in primary glioblastoma. Science 344, 1396–1401 (2014).
Erickson, A. et al. Spatially resolved clonal copy number alterations in benign and malignant tissue. Nature 608, 360–367 (2022).
Nabors, L. B. et al. Central nervous system cancers, version 3.2020, NCCN clinical practice guidelines in oncology. J. Natl Compr. Canc. Netw. 18, 1537–1570 (2020).
Almeida et al. Clinical outcome, tumor recurrence, and causes of death: a long-term follow-up of surgically treated meningiomas. World Neurosurg. 102, 139–143 (2017).
Menke, J. R. et al. Somatostatin receptor 2a is a more sensitive diagnostic marker of meningioma than epithelial membrane antigen. Acta Neuropathol. 130, 441–443 (2015).
Doki, Y. et al. Nivolumab combination therapy in advanced esophageal squamous-cell carcinoma. N. Engl. J. Med. 386, 449–462 (2022).
Forde, P. M. et al. Neoadjuvant nivolumab plus chemotherapy in resectable lung cancer. N. Engl. J. Med. 386, 1973–1985 (2022).
Hellmann, M. D. et al. Nivolumab plus ipilimumab in lung cancer with a high tumor mutational burden. N. Engl. J. Med. 378, 2093–2104 (2018).
Motzer, R. J. et al. Nivolumab plus ipilimumab versus sunitinib in advanced renal-cell carcinoma. N. Engl. J. Med. 378, 1277–1290 (2018).
Hellmann, M. D. et al. Nivolumab plus ipilimumab in advanced non–small-cell lung cancer. N. Engl. J. Med. 381, 2020–2031 (2019).
Motzer, R. J. et al. Nivolumab versus everolimus in advanced renal-cell carcinoma. N. Engl. J. Med. 373, 1803–1813 (2015).
James, L. et al. Combined nivolumab and ipilimumab or monotherapy in untreated melanoma. N. Engl. J. Med. 373, 23–34 (2015).
Caroline, R. et al. Nivolumab in previously untreated melanoma without BRAF mutation. N. Engl. J. Med. 372, 320–330 (2015).
Webb, A. E., Kundaje, A. & Brunet, A. Characterization of the direct targets of FOXO transcription factors throughout evolution. Aging Cell 15, 673–685 (2016).
Corniola, M. V., Lemée, J.-M. & Meling, T. R. Histological transformation in recurrent WHO grade I meningiomas. Sci. Rep. 10, 11220 (2020).
Nakasu, S., Notsu, A., Na, K. & Nakasu, Y. Malignant transformation of WHO grade I meningiomas after surgery or radiosurgery: systematic review and meta-analysis of observational studies. Neurooncol. Adv. 2, vdaa129 (2020).
Chen, W. C. et al. Salvage therapy outcomes for atypical meningioma. J. Neurooncol. 138, 425–433 (2018).
Gauchotte, G. et al. Prognostic value of histopathological features and loss of H3K27me3 immunolabeling in anaplastic meningioma: a multicenter retrospective study. J. Neuropathol. Exp. Neurol. 79, 754–762 (2020).
Wang, J. et al. Clonal evolution of glioblastoma under therapy. Nat. Genet. 48, 768–776 (2016).
Andor, N. et al. Pan-cancer analysis of the extent and consequences of intratumor heterogeneity. Nat. Med. 22, 105–113 (2016).
Gerstung, M. et al. The evolutionary history of 2,658 cancers. Nature 578, 122–128 (2020).
Jamal-Hanjani, M. et al. Tracking the evolution of non–small-cell lung cancer. N. Engl. J. Med. 376, 2109–2121 (2017).
Wang, L. et al. A single-cell atlas of glioblastoma evolution under therapy reveals cell-intrinsic and cell-extrinsic therapeutic targets. Nat. Cancer 3, 1534–1552 (2022).
Lee, W. H. Characterization of a newly established malignant meningioma cell line of the human brain: IOMM-Lee. Neurosurgery 27, 389–395; discussion 396 (1990).
Mei, Y. et al. Genomic profile of human meningioma cell lines. PLoS ONE 12, e0178322 (2017).
Zhou, W., Triche, T. J., Laird, P. W. & Shen, H. SeSAMe: reducing artifactual detection of DNA methylation by Infinium BeadChips in genomic deletions. Nucleic Acids Res. 46, e123 (2018).
Mdm247. Mdm247/spatial_meningioma: v_1.0.0. Zenodo https://doi.org/10.5281/zenodo.10927737 (2024).
Acknowledgements
We thank A. Shai and the staff of the UCSF Brain Tumor Center Biorepository and Pathology Core, B. Ndjamen and the staff of the Gladstone Institutes Histology Core, M. Bernardi and H.-R. Lin and the staff of the Gladstone Institutes Genomics Core, E. Chow and the staff of the UCSF Center for Advanced Technology and E. Zacco and the staff of the UCSF Laboratory for Cell Analysis Genome Core. Lunaphore COMET multiplexed seqIF was made possible by a gift from the Stephen M. Coffman trust to the Northwestern University Malnati Brain Tumor Institute of the Lurie Cancer Center. We appreciate the thoughtful comments and critiques from J. Costello and the members of the Raleigh Laboratory during the inception, execution and dissemination of this study. This study was supported by National Institutes of Health (NIH) grants T32 CA15102 and P50 CA097257 to C.-H.G.L.; NIH grant P50 CA097257 to W.C.C.; NIH grants F30 CA246808 and T32 GM007618 and the UCSF Wolfe Meningioma Program Project to A.C.; NIH grant P50 CA097257 to S.J.L. and J.J.P.; NIH grant F32 CA213944, the UCSF Wolfe Meningioma Program Project and the Northwestern University Malnati Brain Tumor Institute to S.T.M.; NIH grants R01 CA120813, R01 NS120547 and P50 CA221747 to A.B.H.; NIH grants R01 NS117104, R01 NS118039 and P50 CA221747 to C.M.H.; and NIH grants R01 CA262311 and P50 CA097257, the UCSF Wolfe Meningioma Program Project and the Trenchard Family Charitable Fund to D.R.R.
Author information
Authors and Affiliations
Contributions
All authors made substantial contributions to the conception or design of the study; the acquisition, analysis or interpretation of data; or drafting or revising the paper. All authors approved the paper. All authors agree to be personally accountable for individual contributions and to ensure that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved and the resolution documented in the literature. C.-H.G.L. conceived and designed the study with supervision from D.R.R., analyzed bioinformatic data and performed and analyzed histological analyses with supervision from J.J.P., D.A.S. and A.P. K.S., K.F., C.D.E. and M.A.C. performed molecular biology, cell culture and co-culture experiments. K.M., W.C.C., N.Z., A.C. and S.J.L. performed bioinformatic analyses with supervision from H.N.V. H.N. performed meningioma immunofluorescence microscopy with supervision from C.M.H. and A.B.H. S.T.M. generated primary human meningioma cells and curated archival nucleic acid samples from the validation cohort. D.R.R. conceived, designed and supervised the study.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Genetics thanks the anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Meningioma spatial transcriptome analysis.
a, Uniform manifold approximation and projection (UMAP) analysis of spatial transcriptomes from the 16 meningioma samples in this study shaded by unsupervised hierarchical clusters in each sample. b, Distribution of unsupervised hierarchical meningioma spatial transcriptome clusters overlayed onto H&E-stained sections of each sample. Scale bar, 1mm. c, UMAP analysis of meningioma spatial transcriptomes shaded by sample of origin without Harmony batch correction suggesting significant batch effects across meningioma spatial transcriptomes.
Extended Data Fig. 2 Meningioma spatial transcriptome clusters and spatial protein analysis.
a, Spatial transcriptomes in reduced dimensionality clusters from each meningioma sample after Harmony batch correction. b, Stacked bar showing spatial transcriptomes from each meningioma sample after Harmony batch correction. Colors as in a. c, Top 92 differentially expressed genes (Supplementary Table 4) across 30 unsupervised hierarchical spatial transcriptome clusters from all meningiomas after Harmony batch correction. d, Principal component (PC) analysis of spatial protein profiling from all meningioma samples (72 proteins, 82 regions).
Extended Data Fig. 3 High-grade meningiomas are distinguished by divergent intratumor gene and protein expression programs.
a, Preoperative T1 post-contrast magnetic resonance imaging (MRI) of meningiomas with driver mutations associated with adverse clinical outcomes, such as BAP1 loss (M1), CDKN2A/B loss (M2), or TERT promoter mutation (M3). b, Representative H&E-stained sections and immunohistochemistry (IHC) for Ki-67, p16, or H3K27me3 from M1-3. Scale bar, 1mm. c, Uniform manifold approximation and projection (UMAP) of M1-3 spatial transcriptomes after Harmony batch correction shaded by sample of origin. d, UMAP of M1-3 spatial transcriptomes after Harmony batch correction shaded by unsupervised hierarchical clusters. e, Distribution of unsupervised hierarchical spatial transcriptome clusters from M1-3 after Harmony batch correction.
Extended Data Fig. 4 WHO grade 1 meningiomas demonstrate reduced diversity and number of spatial transcriptomes compared to high-grade meningiomas.
a, Uniform manifold approximation and projection (UMAP) of M1-3 and LGM1-2 spatial transcriptomes after Harmony batch correction shaded by sample of origin. b, UMAP of M1-3 and LGM1-2 spatial transcriptomes after Harmony batch correction shaded by unsupervised hierarchical clusters. c, Distribution of unsupervised hierarchical spatial transcriptome clusters from M1-3 and LGM1-2 after Harmony batch correction. d, Top 101 differentially expressed genes across unsupervised hierarchical spatial transcriptome clusters from M1-3 and LGM1-2 after Harmony batch correction. e, Spatial transcriptome expression of MKI67, FOXM1, or CCN3 across M1-3 and LGM1-2. f, InferCNV traces of copy number variants across unsupervised hierarchical transcriptome clusters from M1-3 and LGM1-2 after Harmony batch correction, revealing no significant CNVs in C2 comprising LGM1, and relative loss of chromosomes 1p and 22q in C1 and C7 comprising LGM2. Loses are shown in blue. Gains are shown in red. g, UMAP of LGM1-2 spatial transcriptomes after Harmony batch correction shaded by sample of origin. h, UMAP of LGM1-2 spatial transcriptomes after Harmony batch correction shaded by unsupervised hierarchical clusters. i, Spatial distribution of unsupervised hierarchical spatial transcriptome clusters from LGM1-2. Scale bar, 1mm. j, Distribution of unsupervised hierarchical spatial transcriptome clusters from LGM1-2 after Harmony batch correction. k, Top 116 differentially expressed genes across unsupervised hierarchical spatial transcriptome clusters from LGM1-2 after Harmony batch correction. l, Representative H&E morphology of spatial transcriptome clusters from LGM1 and LGM2. Colors correspond to spatial transcriptomes from h-j. Scale bars, 1mm and 10μm.
Extended Data Fig. 5 Primary and recurrent meningiomas are distinguished by divergent intratumor gene expression programs.
a, Representative H&E-stained sections and immunohistochemistry (IHC) for Ki-67, p16, or H3K27me3 from matched pairs of primary and recurrent meningiomas. Dots show regions of spatial protein profiling. Scale bar, 1mm. b, UMAP analysis of matched pairs of primary and recurrent meningioma spatial transcriptomes after Harmony batch correction shaded by unsupervised hierarchical clusters. Scale bar, 1mm. c, Spatial distribution of unsupervised hierarchical spatial transcriptome clusters from matched pairs of primary and recurrent meningiomas after Harmony batch correction. Scale bar, 1mm. d, Distribution of unsupervised hierarchical spatial transcriptome clusters from matched pairs of primary and recurrent meningiomas after Harmony batch correction.
Extended Data Fig. 6 Primary and recurrent meningiomas are distinguished by divergent copy number variants.
a, InferCNV traces of copy number variants across unsupervised hierarchical transcriptome clusters from matched pairs of primary and recurrent meningiomas after Harmony batch correction. Loses are shown in blue. Gains are shown in red.
Extended Data Fig. 7 Regionally distinct meningiomas are distinguished by divergent intratumor gene and protein expression programs and copy number variants.
a, Preoperative T1 post-contrast magnetic resonance imaging (MRI) of meningioma M8 from patient 7, which was regionally distinct compared to M7 and M7’ (Fig. 3a). b, Representative H&E-stained sections and immunohistochemistry (IHC) for Ki-67, p16, or H3K27me3 from M8. Dots show regions of spatial protein profiling. Scale bar, 1mm. c, Principal component (PC) analysis of spatial protein profiling from M7, M7’, and M8. d, Uniform manifold approximation and projection (UMAP) of M7, M7’, and M8 spatial transcriptomes after Harmony batch correction shaded by sample of origin. e, UMAP of M7, M7’, and M8 spatial transcriptomes after Harmony batch correction shaded by unsupervised hierarchical clusters. f, Spatial distribution of unsupervised hierarchical spatial transcriptome clusters from M7, M7’, and M8 after Harmony batch correction. Scale bar, 1mm. g, Distribution of unsupervised hierarchical spatial transcriptome clusters from M7, M7’, and M8 after Harmony batch correction. h, Spatial distribution of unsupervised hierarchical transcriptome clusters harboring chromosome 10q loss in M8 from InferCNV (Fig. 3c). Scale bar, 1mm. i, InferCNV traces of copy number variants across unsupervised hierarchical transcriptome clusters from M8 after Harmony batch correction. Loses are shown in blue. Gains are shown in red.
Extended Data Fig. 8 Primary and recurrent meningiomas are distinguished by divergent intratumor protein expression programs.
a and b, Principal component (PC) analysis of spatial protein profiling from matched pairs of primary and recurrent meningiomas. c, Spatial proteins from matched pairs of primary and recurrent meningiomas. d, Quantification of differentially expressed spatial proteins from at least 3 of 4 matched pairs of primary (M4 n = 7, M5 n = 6, M6 n = 5, M7 n = 3) and recurrent (M4’ n = 2, M5’ n = 4, M6’ n = 3, M7’ n = 4) meningiomas. Lines represent means and error bars represent standard error of the means. Student’s t tests, one-sided, * ≤ 0.05, **P ≤ 0.01, ***P ≤ 0.0001. e, Multiplexed seqIF microscopy showing sparse lymphocytes in the meningioma microenvironment. Scale bar, 20μm, similar across 3 high-power field replicates per tumor.
Extended Data Fig. 9 High-grade meningiomas are distinguished by regionally distinct intratumor protein expression programs.
a, Preoperative T1 post-contrast magnetic resonance imaging (MRI) of meningiomas M9 and M10. b, Representative H&E-stained sections and immunohistochemistry (IHC) for Ki-67, p16, or H3K27me3 from regionally distinct meningioma samples. Dots show regions of spatial protein profiling. Scale bar, 1mm. c, Principal component (PC) analysis of spatial protein profiling from M9 and M10.
Extended Data Fig. 10 Validation of pharmacological strategies to overcome intratumor heterogeneity in high-grade meningiomas.
a, M10G patient-derived meningioma cells stably expressing CRISPRi machinery (dCas9-KRAB) and sgRNAs suppressing CDKN2A (sgCDKN2A, n = 3), CDKN2B (sgCDKN2B, n = 4), ARID1A (sgARID1A, n = 6), or non-targeted control sgRNAs (sgNTC, n = 3, 4, 6, respectively). Cells were labeled with red or green fluorescence proteins and integrated into 3D co-cultures for pharmacologic and live cell imaging experiments. b, IOMM-Lee meningioma cells that lack endogenous CDKN2A/B stably expressing CRISPRi machinery (dCas9-Zim3) and sgARID1A (n = 4) or sgNTC (n = 4). Cells were labeled with red or green fluorescence proteins and integrated into 3D co-cultures for pharmacologic and live cell imaging experiments. c, Combination molecular therapy treatments of 3D co-cultures of IOMM-LeedCas9-Zim3 meningioma cells expressing sgARID1A or sgNTC. Scale bar, 100μm. d, Quantification of combination molecular therapy treatments of 3D co-cultures of IOMM-LeedCas9-Zim3 meningioma cells expressing sgARID1A or sgNTC. Representative of 8 biological replicates per condition. e, MTT cell viability results normalized to vehicle control after 5 days of molecular therapy of SF12964, a patient-derived meningioma cell line from a WHO grade 3 meningioma (Hypermitotic DNA methylation group, NF2 p.Q165* mutation, chromosome 1p and 22q deletion, CDKN2A/B homozygous deletion, chromosome 1q amplification) that underwent reoperation after 5 months of abemaciclib treatment following prior surgeries and prior radiotherapy treatments. Representative of 8 biological replicates per condition. Lines represent means and error bars represent standard error of the means. Student’s t tests, one-sided, **P ≤ 0.01, ***P ≤ 0.0001.
Supplementary information
Supplementary Tables
Supplementary Table 1. Meningioma discovery cohort clinical data. Supplementary Table 2. Meningioma targeted next-generation DNA sequencing. Supplementary Table 3. Meningioma targeted gene expression profiling. Supplementary Table 4. Meningioma spatial transcriptomic cluster marker genes. Supplementary Table 5. Meningioma spatial protein profiling. Supplementary Table 6. Low-grade meningioma clinical data and targeted next-generation DNA sequencing. Supplementary Table 7. RNA sequencing differential expression analyses across meningiomas with BAP1 inactivation, CDKN2A/B homozygous deletion, or TERT promoter mutation. Supplementary Table 8. Antibodies for multiplexed sequential immunofluorescence. Supplementary Table 9. MAPK and PI3K–AKT target gene expression from RNA sequencing of 502 primary or recurrent meningiomas. Supplementary Table 10. M10GdCas9-KRAB patient-derived meningioma cell RNA sequencing differential expression analyses of protein coding genes.
Source data
Source Data Fig. 7
Numerical values for dot-plot histograms and drug treatment heatmaps/graphs (statistical source data).
Source Data Extended Data Fig. 10
Numerical values for dot-plot histograms and drug treatment heatmaps/graphs (statistical source data).
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Lucas, CH.G., Mirchia, K., Seo, K. et al. Spatial genomic, biochemical and cellular mechanisms underlying meningioma heterogeneity and evolution. Nat Genet (2024). https://doi.org/10.1038/s41588-024-01747-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41588-024-01747-1