Abstract
CRISPR gene editing holds great promise to modify DNA sequences in somatic cells to treat disease. However, standard computational and biochemical methods to predict off-target potential focus on reference genomes. We developed an efficient tool called CRISPRme that considers single-nucleotide polymorphism (SNP) and indel genetic variants to nominate and prioritize off-target sites. We tested the software with a BCL11A enhancer targeting guide RNA (gRNA) showing promise in clinical trials for sickle cell disease and β-thalassemia and found that the top candidate off-target is produced by an allele common in African-ancestry populations (MAF 4.5%) that introduces a protospacer adjacent motif (PAM) sequence. We validated that SpCas9 generates strictly allele-specific indels and pericentric inversions in CD34+ hematopoietic stem and progenitor cells (HSPCs), although high-fidelity Cas9 mitigates this off-target. This report illustrates how genetic variants should be considered as modifiers of gene editing outcomes. We expect that variant-aware off-target assessment will become integral to therapeutic genome editing evaluation and provide a powerful approach for comprehensive off-target nomination.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Sequencing data are deposited in the NCBI Sequence Read Archive database under accession number PRJNA733110. The data for 1000 G were downloaded from https://doi.org/10.12688/wellcomeopenres.15126.2. The data for the HGDP were downloaded from https://doi.org/10.1126/science.aay5012. The full CRISPRme results for NGG PAM gRNAs (including sg1617) are available at https://doi.org/10.5281/zenodo.7195706. Source data are provided with this paper.
Code availability
CRISPRme source code is available at https://github.com/pinellolab/crisprme and https://github.com/InfOmics/CRISPRme. The web app is available online at http://crisprme.di.univr.it. The versions of CRISPRme (1.8.8 and v1.7.7) used to generate the results presented in this manuscript have been deposited on Zenodo: https://doi.org/10.5281/zenodo.5047489. CRISPRitz v2.6.5 used to generate the data presented in this paper have been deposited to Zenodo: https://doi.org/10.5281/zenodo.7078220.
The scripts to generate the plots presented in the manuscript have been deposited to Zenodo: https://doi.org/10.5281/zenodo.7193131.
References
Anzalone, A. V., Koblan, L. W. & Liu, D. R. Genome editing with CRISPR-Cas nucleases, base editors, transposases and prime editors. Nat. Biotechnol. 38, 824–844 (2020).
Clement, K., Hsu, J. Y., Canver, M. C., Joung, J. K. & Pinello, L. Technologies and computational analysis strategies for CRISPR applications. Mol. Cell 79, 11–29 (2020).
Bao, X. R., Pan, Y., Lee, C. M., Davis, T. H. & Bao, G. Tools for experimental and computational analyses of off-target editing by programmable nucleases. Nat. Protoc. 16, 10–26 (2021).
Hsu, P. D. et al. DNA targeting specificity of RNA-guided Cas9 nucleases. Nat. Biotechnol. 31, 827–832 (2013).
Doench, J. G. et al. Optimized sgRNA design to maximize activity and minimize off-target effects of CRISPR-Cas9. Nat. Biotechnol. 34, 184–191 (2016).
Chaudhari, H. G. et al. Evaluation of homology-independent CRISPR-Cas9 off-target assessment methods. CRISPR J. 3, 440–453 (2020).
Lessard, S. et al. Human genetic variation alters CRISPR-Cas9 on- and off-targeting specificity at therapeutically implicated loci. Proc. Natl Acad. Sci. USA 114, E11257–E11266 (2017).
Scott, D. A. & Zhang, F. Implications of human genetic variation in CRISPR-based therapeutic genome editing. Nat. Med. 23, 1095–1101 (2017).
Concordet, J.-P. & Haeussler, M. CRISPOR: intuitive guide selection for CRISPR/Cas9 genome editing experiments and screens. Nucleic Acids Res. 46, W242–W245 (2018).
Listgarten, J. et al. Prediction of off-target activities for the end-to-end design of CRISPR guide RNAs. Nat. Biomed. Eng. 2, 38–47 (2018).
Labun, K. et al. CHOPCHOP v3: expanding the CRISPR web toolbox beyond genome editing. Nucleic Acids Res. 47, W171–W174 (2019).
Park, J., Bae, S. & Kim, J.-S. Cas-Designer: a web-based tool for choice of CRISPR-Cas9 target sites. Bioinformatics 31, 4014–4016 (2015).
Cancellieri, S., Canver, M. C., Bombieri, N., Giugno, R. & Pinello, L. CRISPRitz: rapid, high-throughput and variant-aware in silico off-target site identification for CRISPR genome editing. Bioinformatics 36, 2001–2008 (2020).
Lowy-Gallego, E. et al. Variant calling on the GRCh38 assembly with the data from phase three of the 1000 Genomes Project. Wellcome Open Res 4, 50 (2019).
Bergström, A. et al. Insights into human genetic variation and population history from 929 diverse genomes. Science 367, eaay5012 (2020).
Karczewski, K. J. et al. The mutational constraint spectrum quantified from variation in 141,456 humans. Nature 581, 434–443 (2020).
Frangoul, H. et al. CRISPR-Cas9 gene editing for sickle cell disease and β-thalassemia. N. Engl. J. Med. 384, 252–260 (2021).
Canver, M. C. et al. BCL11A enhancer dissection by Cas9-mediated in situ saturating mutagenesis. Nature 527, 192–197 (2015).
Wu, Y. et al. Highly efficient therapeutic gene editing of human hematopoietic stem cells. Nat. Med. 25, 776–783 (2019).
Walton, R. T., Christie, K. A., Whittaker, M. N. & Kleinstiver, B. P. Unconstrained genome targeting with near-PAMless engineered CRISPR-Cas9 variants. Science 368, 290–296 (2020).
Fennell, T. et al. CALITAS: A CRISPR-Cas-aware ALigner for In silico off-TArget Search. CRISPR J. 4, 264–274 (2021).
Frankish, A. et al. GENCODE reference annotation for the human and mouse genomes. Nucleic Acids Res. 47, D766–D773 (2019).
ENCODE Project Consortium. et al. Expanded encyclopaedias of DNA elements in the human and mouse genomes. Nature 583, 699–710 (2020).
Abadi, S., Yan, W. X., Amar, D. & Mayrose, I. A machine learning approach for predicting CRISPR-Cas9 cleavage efficiencies and patterns underlying its mechanism of action. PLoS Comput. Biol. 13, e1005807 (2017).
Demirci, S. et al. Durable and robust fetal globin induction without Anemia in rhesus monkeys following autologous hematopoietic stem cell transplant with BCL11A erythroid enhancer editing.Blood 134, 4632 (2019).
Schmid-Burgk, J. L. et al. Highly parallel profiling of Cas9 variant specificity. Mol. Cell 78, 794–800.e8 (2020).
Vakulskas, C. A. et al. A high-fidelity Cas9 mutant delivered as a ribonucleoprotein complex enables efficient gene editing in human hematopoietic stem and progenitor cells. Nat. Med. 24, 1216–1224 (2018).
Xu, L. et al. CRISPR/Cas9-mediated CCR5 ablation in human hematopoietic stem/progenitor cells confers HIV-1 resistance in vivo. Mol. Ther. 25, 1782–1789 (2017).
Xu, L. et al. CRISPR-edited stem cells in a patient with HIV and acute lymphocytic leukemia. N. Engl. J. Med. 381, 1240–1247 (2019).
Stadtmauer, E. A. et al. CRISPR-engineered T cells in patients with refractory cancer. Science 367, eaba7365 (2020).
Gillmore, J. D. et al. CRISPR-Cas9 in vivo gene editing for transthyretin amyloidosis. N. Engl. J. Med. 385, 493–502 (2021).
DeWitt, M. A. et al. Selection-free genome editing of the sickle mutation in human adult hematopoietic stem/progenitor cells. Sci. Transl. Med. 8, 360ra134 (2016).
Xu, S. et al. Editing aberrant splice sites efficiently restores β-globin expression in β-thalassemia. Blood 133, 2255–2262 (2019).
Métais, J.-Y. et al. Genome editing of HBG1 and HBG2 to induce fetal hemoglobin. Blood Adv. 3, 3379–3392 (2019).
Tsai, S. Q. et al. GUIDE-seq enables genome-wide profiling of off-target cleavage by CRISPR-Cas nucleases. Nat. Biotechnol. 33, 187–197 (2015).
Zeng, J. et al. Therapeutic base editing of human hematopoietic stem cells. Nat. Med. 26, 535–541 (2020).
Musunuru, K. et al. In vivo CRISPR base editing of PCSK9 durably lowers cholesterol in primates. Nature 593, 429–434 (2021).
Chu, S. H. et al. Rationally designed base editors for precise editing of the sickle cell disease mutation. CRISPR J. 4, 169–177 (2021).
Newby, G. A. et al. Base editing of haematopoietic stem cells rescues sickle cell disease in mice. Nature 595, 295–302 (2021).
Maeder, M. L. et al. Development of a gene-editing approach to restore vision loss in Leber congenital amaurosis type 10. Nat. Med. 25, 229–233 (2019).
De Dreuzy, E. et al. EDIT-301: An experimental autologous cell therapy comprising Cas12a-RNP modified mPB-CD34+ cells for the potential treatment of SCD. Blood 134, 4636–4636 (2019).
Zhao, M., Kim, P., Mitra, R., Zhao, J. & Zhao, Z. TSGene 2.0: an updated literature-based knowledgebase for tumor suppressor genes. Nucleic Acids Res. 44, D1023–D1031 (2016).
Finkel, R. S. et al. Nusinersen versus sham control in infantile-onset spinal muscular atrophy. N. Engl. J. Med. 377, 1723–1732 (2017).
Mercuri, E. et al. Nusinersen versus sham control in later-onset spinal muscular atrophy. N. Engl. J. Med. 378, 625–635 (2018).
Raal, F. J. et al. Inclisiran for the treatment of heterozygous familial hypercholesterolemia. N. Engl. J. Med. 382, 1520–1530 (2020).
Hickey, G. et al. Genotyping structural variants in pangenome graphs using the vg toolkit. Genome Biol. 21, 35 (2020).
Ameur, A. Goodbye reference, hello genome graphs. Nat. Biotechnol. 37, 866–868 (2019).
Center for Biologics Evaluation & Research. Human gene therapy products incorporating human genome editing. U.S. Food and Drug Administration https://www.fda.gov/regulatory-information/search-fda-guidance-documents/human-gene-therapy-products-incorporating-human-genome-editing (2022).
Corces, M. R. et al. Lineage-specific and single-cell chromatin accessibility charts human hematopoiesis and leukemia evolution. Nat. Gen. 48, 1193–1203 (2016).
Acknowledgements
L.P. received support from the National Institutes of Health (NIH) (R35 HG010717 and RM1 HG009490). D.E.B. was supported by the National Heart, Lung, and Blood Institute (OT2HL154984 and P01HL053749), Burroughs Wellcome Fund, Doris Duke Charitable Foundation and the St. Jude Children’s Research Hospital Collaborative Research Consortium. R.G received support from European Union’s ERA-NET JPCOFUND2 (JPND2019-466-037). We thank S. H. Orkin, G. Lettre, J. K. Joung, V. Pattanayak, K. Petri, A. H. Shen, E. Dirupo and F. Masillo for helpful input.
Author information
Authors and Affiliations
Contributions
S.C., L.Y.L., M.T., N.B., R.G. and L.P. created the software; J.Z., M.A.N., S.A.M., M.F.C., V.K., S.Q.T., M.A., S.A.W. and D.E.B. designed and conducted experiments; S.C., J.Z., L.Y.L., J.L., R.G., D.E.B. and L.P. performed data analysis; S.C., R.G., D.E.B. and L.P. conceived the work; S.C., J.Z., L.Y.L, R.G., D.E.B. and L.P. wrote the paper with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
L.P. has financial interests in Edilytics, Excelsior Genomics and SeQure Dx. L.P.’s interests were reviewed and are managed by Massachusetts General Hospital and Partners HealthCare in accordance with their conflict of interest policies. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Genetics thanks Krishanu Saha and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
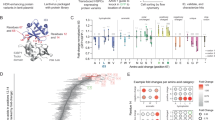
Extended Data Fig. 2 Plots with rank ordered correlation between CFD and CRISTA reported targets.
Scatter plots show from left to right, the correlation of ranked targets, extracted by selecting top 10000 targets ordered by CFD and CRISTA score, respectively. The left plot shows the rank correlation of targets with 0 bulges (Pearson’s correlation: 0.57, p < 1e-10, Spearman’s correlation: 0.55, p < 1e-10), the center plot shows rank correlation of targets with 1 bulge (Pearson’s correlation: −0.16, p < 1 e-10, Spearman’s correlation: −0.33, p < 1 e-10) and the right plot shows the rank correlation of targets with 2 bulges (Pearson’s correlation: −0.55, p < 1e-10, Spearman’s correlation: −0.80, p < 1e-10). The correlation values and p-values(two-sided) were calculated using standard functions from the Python scipy library. The colors represent the lowest count of bulges for each target, because the two scoring methods may prioritize different alignments and thus different number of mismatches and bulges of the same genomic target.
Extended Data Fig. 3 HGDP superpopulation distribution plots.
HGDP variant off-targets with CFD ≥ 0.2 and increase in CFD of ≥0.1. Individual samples from each of the seven superpopulations were shuffled 100 times to calculate the mean and 95% confidence interval. First panel shows distribution within all 54 discrete populations, colored by superpopulation. Additional seven panels show distribution of discrete populations within each listed superpopulation.
Extended Data Fig. 4 Candidate transcript off-targets introduced by common genetic variants for non-CRISPR sequence-based RNA-targeting therapeutic strategies.
a) A common SNP (in blue) introduces a candidate CDS off-target site with 2 mismatches for the FDA-approved antisense oligo Nusinersen. b) Top 1000 candidate transcript off-targets ranked by mismatches and bulges for Nusinersen from a search performed with the 1000 G and HGDP genetic variant datasets. c) A common insertion variant (in red) introduces a candidate 3’UTR off-target site with 4 mismatches + bulges for the FDA-approved RNAi therapy Inclisiran. d) Top 1000 candidate transcript off-targets ranked by mismatches and bulges for Inclisiran from a search performed with the 1000 G and HGDP genetic variant datasets.
Supplementary information
Supplementary Information
Supplementary Figures 1–13, Tables 1–5, Notes 1–9 and Data 1–4. Supplementary figures, tables and notesare included in the flat file. Supplementary data files are separate (see below).
Supplementary Data 1
Top candidate off-targets in CRISPRme search results for sg1617 using hg38, 1000 G and HGDP data with up to six mismatches and two bulges (including the integrated_results, all_results_with_alternative_alignments, and private_targets files).
Supplementary Data 2
Top candidate off-targets in CRISPRme search results for other example gRNAs with NGG PAMs.
Supplementary Data 3
Top candidate off-targets in CRISPRme search results for example gRNAs with non-NGG PAMs.
Supplementary Data 4
Top candidate off-targets in CRISPRme search results for example non-CRISPR-based, RNA-targeting strategies (antisense oligo and RNA interference).
Source data
Source Data Fig. 1
Gel image.
Source Data Fig. 2
ddPCR image files.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Cancellieri, S., Zeng, J., Lin, L.Y. et al. Human genetic diversity alters off-target outcomes of therapeutic gene editing. Nat Genet 55, 34–43 (2023). https://doi.org/10.1038/s41588-022-01257-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-022-01257-y
This article is cited by
-
Selection of extended CRISPR RNAs with enhanced targeting and specificity
Communications Biology (2024)
-
Reassessing human MHC-I genetic diversity in T cell studies
Scientific Reports (2024)
-
Detection of CRISPR/Cas9-Mediated Fetal Hemoglobin Reactivation in Erythroblasts Derived from Cord Blood-Hematopoietic Stem Cells
Molecular Biotechnology (2024)
-
Splice modulators target PMS1 to reduce somatic expansion of the Huntington’s disease-associated CAG repeat
Nature Communications (2024)
-
Interpretable neural architecture search and transfer learning for understanding CRISPR–Cas9 off-target enzymatic reactions
Nature Computational Science (2023)