Abstract
Inflammasome sensors detect pathogen- and danger-associated molecular patterns and promote inflammation and pyroptosis1. NLRP1 was the first inflammasome sensor to be described, and its hyperactivation is linked to autoinflammatory disease and cancer2,3,4,5,6. However, the mechanism underlying the activation and regulation of NLRP1 has not been clearly elucidated4,7,8. Here we identify ubiquitously expressed endogenous thioredoxin (TRX) as a binder of NLRP1 and a suppressor of the NLRP1 inflammasome. The cryo-electron microscopy structure of human NLRP1 shows NLRP1 bound to Spodoptera frugiperda TRX. Mutagenesis studies of NLRP1 and human TRX show that TRX in the oxidized form binds to the nucleotide-binding domain subdomain of NLRP1. This observation highlights the crucial role of redox-active cysteines of TRX in NLRP1 binding. Cellular assays reveal that TRX suppresses NLRP1 inflammasome activation and thus negatively regulates NLRP1. Our data identify the TRX system as an intrinsic checkpoint for innate immunity and provide opportunities for future therapeutic intervention in NLRP1 inflammasome activation targeting this system.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All data needed to evaluate the conclusions in the paper are presented in the paper and/or extended data figures and table. Additional data and resources related to this paper may be requested from the authors. The cryo-EM maps and related structure coordinates of the NLRP1–sfTRX complex have been deposited in the Electron Microscopy Data Bank (EMDB) under accession codes EMD-32484 (consensus map) and EMD-35591 (focused map) and in the Protein Data Bank (PDB) under accession 7WGE. Source data are provided with this paper.
References
Pandey, A., Shen, C., Feng, S. & Man, S. M. Cell biology of inflammasome activation. Trends Cell Biol. 31, 924–939 (2021).
Jin, Y. et al. NALP1 in vitiligo-associated multiple autoimmune disease. N. Engl. J. Med. 356, 1216–1225 (2007).
Martinon, F., Burns, K. & Tschopp, J. The inflammasome: a molecular platform triggering activation of inflammatory caspases and processing of proIL-β. Mol. Cell. 10, 417–426 (2002).
Taabazuing, C. Y., Griswold, A. R. & Bachovchin, D. A. The NLRP1 and CARD8 inflammasomes. Immunol. Rev. 297, 13–25 (2020).
Williams, T. M. et al. The NLRP1 inflammasome attenuates colitis and colitis-associated tumorigenesis. J. Immunol. 194, 3369–3380 (2015).
Zhong, F. L. et al. Germline NLRP1 mutations cause skin inflammatory and cancer susceptibility syndromes via inflammasome activation. Cell 167, 187–202. e117 (2016).
Bauernfried, S., Scherr, M. J., Pichlmair, A., Duderstadt, K. E. & Hornung, V. Human NLRP1 is a sensor for double-stranded RNA. Science 371, eabd0811 (2021).
Robinson, K. S. et al. ZAKα-driven ribotoxic stress response activates the human NLRP1 inflammasome. Science 377, 328–335 (2022).
Li, X., Kapos, P. & Zhang, Y. NLRs in plants. Curr. Opin. Immunol. 32, 114–121 (2015).
Platnich, J. M. & Muruve, D. A. NOD-like receptors and inflammasomes: A review of their canonical and non-canonical signaling pathways. Arch. Biochem. Biophys. 670, 4–14 (2019).
Kummer, J. A. et al. Inflammasome components NALP 1 and 3 show distinct but separate expression profiles in human tissues suggesting a site-specific role in the inflammatory response. J. Histochem. Cytochem. 55, 443–452 (2007).
Sanz, C. et al. NALP1 is a transcriptional target for cAMP-response-element-binding protein (CREB) in myeloid leukaemia cells. Biochem. J 384, 281–286 (2004).
Mitchell, P. S., Sandstrom, A. & Vance, R. E. The NLRP1 inflammasome: new mechanistic insights and unresolved mysteries. Curr. Opin. Immunol. 60, 37–45 (2019).
Finger, J. N. et al. Autolytic proteolysis within the function to find domain (FIIND) is required for NLRP1 inflammasome activity. J. Biol. Chem. 287, 25030–25037 (2012).
Tsu, B. V. et al. Diverse viral proteases activate the NLRP1 inflammasome. eLife 10, e60609 (2021).
Robinson, K. S. et al. Enteroviral 3C protease activates the human NLRP1 inflammasome in airway epithelia. Science 370, eaay2002 (2020).
Hollingsworth, L. R. et al. Mechanism of filament formation in UPA-promoted CARD8 and NLRP1 inflammasomes. Nat. Commun. 12, 189 (2021).
Gong, Q. et al. Structural basis for distinct inflammasome complex assembly by human NLRP1 and CARD8. Nat. Commun. 12, 188 (2021).
Zhong, F. L. et al. Human DPP9 represses NLRP1 inflammasome and protects against autoinflammatory diseases via both peptidase activity and FIIND domain binding. J. Biol. Chem. 293, 18864–18878 (2018).
Huang, M. et al. Structural and biochemical mechanisms of NLRP1 inhibition by DPP9. Nature 592, 773–777 (2021).
Hollingsworth, L. R. et al. DPP9 sequesters the C terminus of NLRP1 to repress inflammasome activation. Nature 592, 778–783 (2021).
D’Osualdo, A. et al. CARD8 and NLRP1 undergo autoproteolytic processing through a ZU5-like domain. PLoS ONE 6, e27396 (2011).
Gregory, S. M. et al. Discovery of a viral NLR homolog that inhibits the inflammasome. Science 331, 330–334 (2011).
Hsu, L.-C. et al. A NOD2–NALP1 complex mediates caspase-1-dependent IL-1β secretion in response to Bacillus anthracis infection and muramyl dipeptide. Proc. Natl Acad. Sci. USA 105, 7803–7808 (2008).
Bruey, J.-M. et al. Bcl-2 and Bcl-XL regulate proinflammatory caspase-1 activation by interaction with NALP1. Cell 129, 45–56 (2007).
Martin, J. L. Thioredoxin—a fold for all reasons. Structure 3, 245–250 (1995).
Sharif, H. et al. Structural mechanism for NEK7-licensed activation of NLRP3 inflammasome. Nature 570, 338–343 (2019).
Maekawa, S., Ohto, U., Shibata, T., Miyake, K. & Shimizu, T. Crystal structure of NOD2 and its implications in human disease. Nat. Commun. 7, 11813 (2016).
Hu, Z. et al. Crystal structure of NLRC4 reveals its autoinhibition mechanism. Science 341, 172–175 (2013).
Weichsel, A., Gasdaska, J. R., Powis, G. & Montfort, W. R. Crystal structures of reduced, oxidized, and mutated human thioredoxins: evidence for a regulatory homodimer. Structure 4, 735–751 (1996).
Dekker, C. et al. Crystal structure of NLRP3 NACHT domain with an inhibitor defines mechanism of inflammasome inhibition. J. Mol. Biol. 433, 167309 (2021).
Sandstrom, A. et al. Functional degradation: a mechanism of NLRP1 inflammasome activation by diverse pathogen enzymes. Science 364, eaau1330 (2019).
Zhang, L. et al. Cryo-EM structure of the activated NAIP2–NLRC4 inflammasome reveals nucleated polymerization. Science 350, 404–409 (2015).
Hu, Z. et al. Structural and biochemical basis for induced self-propagation of NLRC4. Science 350, 399–404 (2015).
Andreeva, L. et al. NLRP3 cages revealed by full-length mouse NLRP3 structure control pathway activation. Cell 184, 6299–6312.e22 (2021).
Xiao, L., Magupalli, V. G. & Wu, H. Cryo-EM structures of the active NLRP3 inflammasome disc. Nature 613, 595–600 (2023).
Kim, H.-E., Du, F., Fang, M. & Wang, X. Formation of apoptosome is initiated by cytochrome c-induced dATP hydrolysis and subsequent nucleotide exchange on Apaf-1. Proc. Natl Acad. Sci. USA 102, 17545–17550 (2005).
Wang, J. et al. Ligand-triggered allosteric ADP release primes a plant NLR complex. Science 364, eaav5868 (2019).
Wang, J. et al. Reconstitution and structure of a plant NLR resistosome conferring immunity. Science 364, eaav5870 (2019).
Hwang, J. et al. The structural basis for the negative regulation of thioredoxin by thioredoxin-interacting protein. Nat. Commun. 5, 2958 (2014).
Fritz-Wolf, K., Kehr, S., Stumpf, M., Rahlfs, S. & Becker, K. Crystal structure of the human thioredoxin reductase–thioredoxin complex. Nat. Commun. 2, 383 (2011).
Wang, Y. et al. Thioredoxin-1 attenuates atherosclerosis development through inhibiting NLRP3 inflammasome. Endocrine 70, 65–70 (2020).
Jia, J., Zhang, X., Xu, G., Zeng, X. & Li, L. Thioredoxin-1 inhibits amyloid-β25–35-induced activation of NLRP1/caspase-1/GSDMD pyroptotic pathway in PC12 cells. Mol. Biol. Rep. 49, 3445–3452 (2022).
Ball, D. P. et al. Oxidized thioredoxin-1 restrains the NLRP1 inflammasome. Sci. Immunol. 7, eabm7200 (2022).
Perkins, D. N., Pappin, D. J., Creasy, D. M. & Cottrell, J. S. Probability‐based protein identification by searching sequence databases using mass spectrometry data. Electrophoresis 20, 3551–3567 (1999).
Hunter, J. D. Matplotlib: A 2D graphics environment. Comput. Sci. Eng. 9, 90–95 (2007).
Liu, L. et al. Structural basis of toll-like receptor 3 signaling with double-stranded RNA. Science 320, 379–381 (2008).
Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. eLife 7, e42166 (2018).
Scheres, S. H. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Rohou, A. & Grigorieff, N. CTFFIND4: Fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Sanchez-Garcia, R. et al. DeepEMhancer: a deep learning solution for cryo-EM volume post-processing. Commun. Biol. 4, 874 (2021).
Reubold, T. F., Hahne, G., Wohlgemuth, S. & Eschenburg, S. Crystal structure of the leucine-rich repeat domain of the NOD-like receptor NLRP1: Implications for binding of muramyl dipeptide. FEBS Lett. 588, 3327–3332 (2014).
Tunyasuvunakool, K. et al. Highly accurate protein structure prediction for the human proteome. Nature 596, 590–596 (2021).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Goddard, T. D. et al. UCSF ChimeraX: meeting modern challenges in visualization and analysis. Protein Sci. 27, 14–25 (2018).
Lee, P. H., Huang, X. X., Teh, B. T. & Ng, L. M. TSA-CRAFT: a free software for automatic and robust thermal shift assay data analysis. SLAS Discov. 24, 606–612 (2019).
Yasudo, H. et al. A possible association between a nucleotide‐binding domain LRR‐containing protein family PYD‐containing protein 1 mutation and an autoinflammatory disease involving liver cirrhosis. Hepatology 74, 2296–2299 (2021).
Rueden, C. T. et al. PyImageJ: a library for integrating ImageJ and Python. Nat. Methods 19, 1326–1327 (2022).
Dickson, M. A. et al. Human keratinocytes that express hTERT and also bypass a p16INK4a-enforced mechanism that limits life span become immortal yet retain normal growth and differentiation characteristics. Mol. Cell. Biol. 20, 1436–1447 (2000).
Acknowledgements
The authors thank M. Kikkawa, H. Yanagisawa, A. Tsutsumi, Y. Sakamaki and Y. Kise for the management and support of the Graduate School of Medicine cryo-EM facility at the University of Tokyo; Y. Kamitsukasa for preparation of NLRP9 protein; K. Murakami, S. Sakurai, M. Masutani and X.-W. Zhu for initial plasmid construction; M. Kanai and S. A. Kawashima for providing HEK293T cells; and J. G. Rheinwald for providing N/TERT-1 keratinocytes. This work was supported by a Grant-in-Aid from the Japanese Ministry of Education, Culture, Sports, Science, and Technology Grant nos 22K15046 and 20K15730 (Z.Z.), 22H02556 and 23K18211 (U.O.) and 23H00366 (T. Shimizu.); CREST, JST (grant no. JPMJCR21E4) (T. Shimizu. and K.M.); JP22H05182 (T. Shimizu. and K.M.); the Research Foundation for Pharmaceutical Sciences (Z.Z.); the Mochida Memorial Foundation for Medical and Pharmaceutical Research (Z.Z.); and the Basis for Supporting Innovative Drug Discovery and Life Science Research (BINDS) from the Japan Agency of Medical Research and Development (AMED) (grant no. JP21am0101115; support no. 1570, 1846, 1848).
Author information
Authors and Affiliations
Contributions
Conceptualization: Z.Z. Investigation (recombinant protein generation and biochemical analysis): Z.Z. Investigation (cryo-EM analysis and image processing): Z.Z. and U.O. Investigation (cellular assay): T. Shibata, Z.Z., A.F., J.K. and K.M. Visualization: Z.Z. Validation, data curation and project administration: Z.Z. and U.O. Writing (original draft): Z.Z. and U.O. Writing (review and editing): Z.Z., U.O. and T. Shimizu. Supervision: U.O. and T. Shimizu. Funding acquisition: Z.Z., U.O. and T. Shimizu.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks the anonymous reviewer(s) for their contribution to the peer review of this work. Peer review reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Sequence alignment of NLRP1 from various species.
Sequence alignment of human, monkey, bovine, horse, mouse (NLRP1a and NLRP1b), rat (NLRP1a), and zebrafish NLRP1 was performed using the Clustal Omega web server. Secondary structures of human NLRP1 in this study are indicated by cylinders (α helices) and arrows (β-strands) above the sequence. Residues involved in thioredoxin binding are highlighted in yellow color. Walker A, Walker B, Sensor I motifs and a conversed histidine are boxed in blue lines.
Extended Data Fig. 2 Binding assay of hNLRP1 and nucleic acids, and putative dsRNA density.
a, Binding assay of hNLRP1230–994 and dsRNAs or dsDNAs using electrophoretic mobility shift assay (EMSA). The assay was independently performed twice and representative results are shown. b, The structure of the hNLRP195–994/sfTRX complex and the cryo-EM density map (consensus map). Cryo-EM density maps are displayed in semi-transparent surface representation with hNLRP195–994/sfTRX complex at two contour levels, 0.025 (left) and 0.012 (right). Densities around HD1-HD2-LRR interface that might be attributable to the dsRNA are indicated (right).
Extended Data Fig. 3 Cryo-EM image processing and density map.
a, Cryo-EM data processing workflow of the hNLRP195–994/sfTRX complex. Representative motion-corrected micrograph, 2D class averages, 3D class-averages, gold-standard FSC curves of the final 3D reconstruction (consensus map, resolution cut-off at FSC = 0.143), the consensus map, the focused map with its focus mask of hNLRP1(NOD)/sfTRX (purple meshes), and the consensus map (colored according to the local resolution) are shown. 2D class averages were calculated using the refined particles that were used for the final reconstruction. 3D classes selected for the following analyses are indicated with red boxes. b, Representative cryo-EM density map (B-factor sharpened maps of the consensus map or the focused map if indicated) of the hNLRP195–994/sfTRX complex. The density map is segmented and displayed by domain (sfTRX, NBD, HD1, WHD, HD2 and LRR). Densities around the C32-C35 disulfide bond and ATPγS are shown. Densities of the hNLRP1/sfTRX binding interface are shown. c, Model to map (B-factor sharpened consensus map) FSC curve.
Extended Data Fig. 4 Structural comparisons of hNLRP1 and other NLRs.
a, Structural comparison of the hNLRP195–994/sfTRX complex (this study), the NLRP3/NEK7 complex (PDB: 6NPY), NOD2 (PDB: 5IRN) and NLRC4 (PDB: 4KXF). Domains of each NLR are shown in the same colors. ATPγS and ADP are shown in sphere representations. b, Superposition of the NBD-HD1 and the bound sfTRX in the structure of the hNLRP195–994/sfTRX complex onto the NLRC4 inflammasome structure (PDB: 3JBL). NLRC4 protomers are colored gray or yellow.
Extended Data Fig. 5 Structural analysis of thioredoxin binding to NLRP1.
a, Superposition of the crystal structures of oxidized (PDB: 1ERU) and reduced (PDB: 1ERT) hTRX onto the structure of the hNLRP195–994/sfTRX complex (this study). C32 and C35 of TRX are shown in stick representations. b, Inter-molecular disulfide bond formation assay between hNLRP1230–994 (sfTRX removed) and hTRX (WT, C32S, C35S or C73Y). Reducing (left) and non-reducing (right) SDS-PAGE analyses (n = 1). c, Reducing and non-reducing SDS-PAGE of the cryo-EM sample of hNLRP195–994/sfTRX (n = 1). d, Superposition of hNLRP195–994/sfTRX complex (this study) onto hTXNIP/hTRX complex (PDB: 4LL1) (left), and hTRX reductase (hTRXR)/hTRX complex (PDB: 3QFB) (right). Structures are superimposed by TRX.
Extended Data Fig. 6 Purification of MBP-hNLRP1230–994 WT and L399E proteins.
a, Gel filtration purification of MBP-hNLRP1230–994 WT (upper) and L399E (lower) proteins. Absorbance at 280 nm is shown in blue lines. b, SDS-PAGE analysis (stained with CBB) of fractions shown in (a) (n = 1).
Extended Data Fig. 7 Comparison of the NLR NBD subdomains.
a, Sequence alignment of the NBD subdomains from various NLRs (NLRP1, NLRP3, NLRP9, NLRP7, NOD2, and NLRC4). The alignment was performed using the Clustal Omega web server. Residues involved in the thioredoxin binding in the structure of hNLRP195–994/sfTRX complex are highlighted in yellow color. b, Structure comparison of the NBD subdomains of NLRP1 (this study), NOD2 (PDB: 5IRN) and NLRP3 (PDB: 7ALV). Residues involved in the thioredoxin binding in the structure of hNLRP195–994/sfTRX complex and corresponding residues for NOD2 and NLRP3 are shown in stick representations. c, StrepII tag pull-down assay using StrepII-tagged hTRX protein (6xHis tag-cleaved) and hNLRP1230–994, rNOD2ΔC, hNOD2FL, hNLRP7ΔP, hNLRP9FL and mNLRP3FL proteins. The assay was independently performed three times and representative results are shown.
Extended Data Fig. 8 Thermal shift assay of hNLRP1230–994.
Thermal shift assay of hNLRP1230–994 (sfTRX removed) with or without hTRX (1.0 eq or 2.0 eq). Means of normalized fluorescence (n = 8) versus temperature were plotted as the melting curves (left). The Tm values (n = 8) were calculated using the Boltzmann sigmodal equation and plotted with dots (right). Tm values are presented as mean values ± SEM. hNLRP1230–994 with hTRX showed a significant high Tm values. P values were determined by one-way ANOVA with two-sided Sidak’s multiple comparison test. *P < 0.05, **P < 0.01, ***P < 0.001.
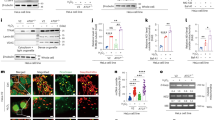
Extended Data Fig. 9 Cellular assay using N/TERT-1 keratinocyte.
a, Representative Western blot analysis of the cell lysates of N/TERT-1 keratinocytes (WT and three TRX-KO clones). b-c, The secreted level of mature IL-1β in N/TERT-1 keratinocytes (WT and three TRX-KO clones) treated with ANS (b) or poly(I:C) (c). Lipofectamine 2000 (LF) was used for intracellular delivery of poly(I:C). Bar plots are mean ± SEM. Dot plots indicate individual data points. Data are n = 3 independent biological replicates. P values were determined by two-way ANOVA with two-sided Dunnett’s multiple comparison test (control = WT). *P < 0.05, **P < 0.01, ***P < 0.001.
Extended Data Fig. 10 NLRP1 inflammasome activation and regulation models.
In steady state, NLRP1 forms complex with TRX. Release of TRX from NLRP1 induces hypersensitive state, which has higher inflammasome activation. Release of TRX is possibly regulated by antioxidants, TXNIP, and TRX reductase. TRX may also play a role in regulating other NLRs.
Supplementary information
Supplementary Information
This file contains Supplementary Figs. 1–3.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhang, Z., Shibata, T., Fujimura, A. et al. Structural basis for thioredoxin-mediated suppression of NLRP1 inflammasome. Nature 622, 188–194 (2023). https://doi.org/10.1038/s41586-023-06532-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-023-06532-4
This article is cited by
-
Stimulation of tumoricidal immunity via bacteriotherapy inhibits glioblastoma relapse
Nature Communications (2024)
-
NLRP1- A CINDERELLA STORY: a perspective of recent advances in NLRP1 and the questions they raise
Communications Biology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.