Abstract
The gut microbiome can influence the development of tumours and the efficacy of cancer therapeutics1,2,3,4,5; however, the multi-omics characteristics of antitumour bacterial strains have not been fully elucidated. In this study, we integrated metagenomics, genomics and transcriptomics of bacteria, and analyses of mouse intestinal transcriptome and serum metabolome data to reveal an additional mechanism by which bacteria determine the efficacy of cancer therapeutics. In gut microbiome analyses of 96 samples from patients with non-small-cell lung cancer, Bifidobacterium bifidum was abundant in patients responsive to therapy. However, when we treated syngeneic mouse tumours with commercial strains of B. bifidum to establish relevance for potential therapeutic uses, only specific B. bifidum strains reduced tumour burden synergistically with PD-1 blockade or oxaliplatin treatment by eliciting an antitumour host immune response. In mice, these strains induced tuning of the immunological background by potentiating the production of interferon-γ, probably through the enhanced biosynthesis of immune-stimulating molecules and metabolites.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
16s rRNA and WGS data from human stool samples, the mouse intestinal RNA sequencing data, the bacterial WGS data and the bacterial RNA sequencing data have been deposited in the European Nucleotide Archive (accession number PRJEB26531). Source data are provided with this paper.
References
Viaud, S. et al. The intestinal microbiota modulates the anticancer immune effects of cyclophosphamide. Science 342, 971–976 (2013).
Daillère, R. et al. Enterococcus hirae and Barnesiella intestinihominis facilitate cyclophosphamide-induced therapeutic immunomodulatory effects. Immunity 45, 931–943 (2016).
Iida, N. et al. Commensal bacteria control cancer response to therapy by modulating the tumor microenvironment. Science 342, 967–970 (2013).
Sivan, A. et al. Commensal Bifidobacterium promotes antitumor immunity and facilitates anti-PD-L1 efficacy. Science 350, 1084–1089 (2015).
Gopalakrishnan, V. et al. Gut microbiome modulates response to anti-PD-1 immunotherapy in melanoma patients. Science 359, 97–103 (2018).
Thaiss, C. A., Zmora, N., Levy, M. & Elinav, E. The microbiome and innate immunity. Nature 535, 65–74 (2016).
Honda, K. & Littman, D. R. The microbiota in adaptive immune homeostasis and disease. Nature 535, 75–84 (2016).
Rosshart, S. P. et al. Wild mouse gut microbiota promotes host fitness and improves disease resistance. Cell 171, 1015–1028 (2017).
Vétizou, M. et al. Anticancer immunotherapy by CTLA-4 blockade relies on the gut microbiota. Science 350, 1079–1084 (2015).
Routy, B. et al. Gut microbiome influences efficacy of PD-1-based immunotherapy against epithelial tumors. Science 359, 91–97 (2018).
Matson, V. et al. The commensal microbiome is associated with anti-PD-1 efficacy in metastatic melanoma patients. Science 359, 104–108 (2018).
Eisenhauer, E. A. et al. New response evaluation criteria in solid tumours: revised RECIST guideline (version 1.1). Eur. J. Cancer 45, 228–247 (2009).
Guerrero-Ros, I. et al. The negative effect of lipid challenge on autophagy inhibits T cell responses. Autophagy 16, 223–238 (2020).
Gao, J. et al. Loss of IFN-γ pathway genes in tumor cells as a mechanism of resistance to anti-CTLA-4 therapy. Cell 167, 397–404.e9 (2016).
Patel, S. J. et al. Identification of essential genes for cancer immunotherapy. Nature 548, 537–542 (2017).
Medzhitov, R. Toll-like receptors and innate immunity. Nat. Rev. Immunol. 1, 135–145 (2001).
Kohwi, Y., Imai, K., Tamura, Z. & Hashimoto, Y. Antitumor effect of Bifidobacterium infantis in mice. GANN 69, 613–618 (1978).
Rafter, J. Probiotics and colon cancer. Best. Pract. Res. Clin. Gastroenterol. 17, 849–859 (2003).
Zhang, L. S. & Davies, S. S. Microbial metabolism of dietary components to bioactive metabolites: opportunities for new therapeutic interventions. Genome Med. 8, 46 (2016).
Desbonnet, L., Garrett, L., Clarke, G., Bienenstock, J. & Dinan, T. G. The probiotic Bifidobacteria infantis: an assessment of potential antidepressant properties in the rat. J. Psychiatr. Res. 43, 164–174 (2008).
Xiao, J. et al. Effects of milk products fermented by Bifidobacterium longum on blood lipids in rats and healthy adult male volunteers. J. Dairy Sci. 86, 2452–2461 (2003).
An, H. M. et al. Antiobesity and lipid-lowering effects of Bifidobacterium spp. in high fat diet-induced obese rats. Lipids Health Dis. 10, 116 (2011).
Wang, K. et al. Bifidobacterium bifidum TMC3115 can characteristically influence glucose and lipid profile and intestinal microbiota in the middle-aged and elderly. Probiotics Antimicrob. Proteins 11, 1182–1194 (2019).
Howie, D., Ten Bokum, A., Necula, A. S., Cobbold, S. P. & Waldmann, H. The role of lipid metabolism in T lymphocyte differentiation and survival. Front. Immunol. 8, 1949 (2018).
Long, J. et al. Lipid metabolism and carcinogenesis, cancer development. Am. J. Cancer Res. 8, 778–791 (2018).
Cruceriu, D., Baldasici, O., Balacescu. & Berindan-Neagoe, I. The dual role of tumor necrosis factor-alpha (TNF-α) in breast cancer: molecular insights and therapeutic approaches. Cell. Oncol. 43, 1–18 (2020).
Chen, X., Bäumel, M., Männel, D. N., Howard, O. Z. & Oppenheim, J. J. Interaction of TNF with TNF receptor type 2 promotes expansion and function of mouse CD4+ CD25+ T regulatory cells. J. Immunol. 179, 154–161 (2007).
Schioppa, T. et al. B regulatory cells and the tumor-promoting actions of TNF-α during squamous carcinogenesis. Proc. Natl Acad. Sci. USA 108, 10662–10667 (2011).
Zhao, X. et al. TNF signaling drives myeloid-derived suppressor cell accumulation. J. Clin. Invest. 122, 4094–4104 (2012).
Zheng, L. et al. Induction of apoptosis in mature T cells by tumour necrosis factor. Nature 377, 348–351 (1995).
Bertrand, F. et al. Blocking tumor necrosis factor α enhances CD8 T-cell-dependent immunity in experimental melanoma. Cancer Res. 75, 2619–2628 (2015).
Klindworth, A. et al. Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids Res. 41, e1 (2013).
Davis, M. P., van Dongen, S., Abreu-Goodger, C., Bartonicek, N. & Enright, A. J. Kraken: a set of tools for quality control and analysis of high-throughput sequence data. Methods 63, 41–49 (2013).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet.journal 17, 10–12 (2011).
Bolyen, E. et al. Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat. Biotechnol. 37, 852–857 (2019).
Callahan, B. J. et al. DADA2: high-resolution sample inference from Illumina amplicon data. Nat. Methods 13, 581–583 (2016).
DeSantis, T. Z. et al. Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl. Environ. Microbiol. 72, 5069–5072 (2006).
Segata, N. et al. Metagenomic biomarker discovery and explanation. Genome Biol. 12, R60 (2011).
Hill, D. A. et al. Metagenomic analyses reveal antibiotic-induced temporal and spatial changes in intestinal microbiota with associated alterations in immune cell homeostasis. Mucosal Immunol. 3, 148–158 (2010).
Segata, N. et al. Metagenomic microbial community profiling using unique clade-specific marker genes. Nat. Methods 9, 811–814 (2012).
Abubucker, S. et al. Metabolic reconstruction for metagenomic data and its application to the human microbiome. PLoS Comput. Biol. 8, e1002358 (2012).
Parks, D. H., Tyson, G. W., Hugenholtz, P. & Beiko, R. G. STAMP: statistical analysis of taxonomic and functional profiles. Bioinformatics 30, 3123–3124 (2014).
Ferrer-Font, L. et al. High-dimensional analysis of intestinal immune cells during helminth infection. eLife 9, e51678 (2020).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-seq data with or without a reference genome. BMC Bioinformatics 12, 323 (2011).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Bindea, G. et al. ClueGO: a Cytoscape plug-in to decipher functionally grouped gene ontology and pathway annotation networks. Bioinformatics 25, 1091–1093 (2009).
Bang, D. Y., Byeon, S. K. & Moon, M. H. Rapid and simple extraction of lipids from blood plasma and urine for liquid chromatography–tandem mass spectrometry. J. Chromatogr. A 1331, 19–26 (2014).
Lim, S., Byeon, S. K., Lee, J. Y. & Moon, M. H. Computational approach to structural identification of phospholipids using raw mass spectra from nanoflow liquid chromatography–electrospray ionization–tandem mass spectrometry. J. Mass Spectrom. 47, 1004–1014 (2012).
Horai, H. et al. MassBank: a public repository for sharing mass spectral data for life sciences. J. Mass Spectrom. 45, 703–714 (2010).
Ivanova, P. T., Milne, S. B., Myers, D. S. & Brown, H. A. Lipidomics: a mass spectrometry based systems level analysis of cellular lipids. Curr. Opin. Chem. Biol. 13, 526–531 (2009).
Bankevich, A. et al. SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 19, 455–477 (2012).
Gurevich, A., Saveliev, V., Vyahhi, N. & Tesler, G. QUAST: quality assessment tool for genome assemblies. Bioinformatics 29, 1072–1075 (2013).
Seemann, T. Prokka: rapid prokaryotic genome annotation. Bioinformatics 30, 2068–2069 (2014).
Page, A. J. et al. Roary: rapid large-scale prokaryote pan genome analysis. Bioinformatics 31, 3691–3693 (2015).
Pritchard, L., Glover, R. H., Humphris, S., Elphinstone, J. G. & Toth, I. K. Genomics and taxonomy in diagnostics for food security: soft-rotting enterobacterial plant pathogens. Anal. Methods 8, 12–24 (2016).
Bardou, P., Mariette, J., Escudié, F., Djemiel, C. & Klopp, C. jvenn: an interactive Venn diagram viewer. BMC Bioinformatics 15, 293 (2014).
Förstner, K. U., Vogel, J. & Sharma, C. M. READemption—a tool for the computational analysis of deep-sequencing-based transcriptome data. Bioinformatics 30, 3421–3423 (2014).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Acknowledgements
This research was supported by the Bio & Medical Technology Development Program of the National Research Foundation (NRF), funded by the Ministry of Science & ICT (NRF-2017M3A9F3046536 to H.P.); a GIST Research Institute (GRI) grant, funded by the GIST in 2020; Genome and Company (grant number GNC GR 17-01 to H.P.). This work was also supported by grants from National Cancer Centre, Korea (NCC-1911267 to H.P); the Post-Genome Technology Development Program (10067758, Business model development driven by clinico-genomic database for precision immuno-oncology to S.-H.L.) funded by the Ministry of Trade, Industry and Energy (MOTIE, Korea). S.-Y.C. is supported by the Bio & Medical Technology Development Program of the NRF, funded by the Ministry of Science & ICT (NRF-2018M3A9F3056902). This research was also supported by a grant from the Korea Health Technology R&D Project through the Korea Health Industry Development Institute (KHIDI), funded by the Ministry of Health & Welfare, Republic of Korea (grant number HI17C1076 to K.W.Y). This work was supported by the NRF grant funded by the Korea government (MSIT) (number 2017R1C1B2011196 to K.W.Y). M.H.M. is supported by grant NRF-2018R1A2A1A05019794 from the NRF of Korea. B.-N.J. is supported by the Basic Science Research Program through the NRF of Korea, funded by the Ministry of Education, Science and Technology (MEST) (2018R1C1B6005768). The International Research & Development Program of the NRF of Korea, funded by the Ministry of Science, ICT & Future Planning (grant no. 2015K1A4A3047851) also funded C.L. The Ewha Womans University Professorship is supported in part by the Ewha Womans University research grant of 2017-2019. This study is also supported in part by operational funds from The First Affiliated Hospital of Xi’an Jiaotong University.
Author information
Authors and Affiliations
Contributions
S.-H.L. and H.S.K. collected and analysed the clinical data. S.-H.L., S.-Y.C., Y.Y., C.P. and K.W.Y. wrote the manuscript. J.Y.K., S.W., J.-S.P., G.M.W., C.L. and H.P. revised the manuscript. S.-H.L., S.-Y.C., C.P. and G.K. analysed the 16S rRNA sequences and transcriptomics data. J.-J.J., B.-N.J., H.-S.Y. and Sarang Kim performed the cytometry analysis. Y.Y., J.S., S.L., Y.Y.K., Sujeong Kim, Yunjae Kim and S.G.K. performed the syngeneic mouse model experiments. Seonggon Kim. and J.-S.P. performed the orthotopic mouse model experiments. C.A., E.J.L., Yeongmin Kim and H.K. cultured the bacteria. H.-S.Y. and Sarang Kim performed the in vitro T-cell assays. M.J., H.C. and M.H.N. analysed metabolomics data. G.B.L. and M.H.M. analysed the lipidomics data. K.W.Y. and H.P. designed and supervised all experiments and analyses.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Microbiology thanks the anonymous reviewers for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Comparison of gut microbiome composition between non–small cell lung cancer (NSCLC) patients and healthy controls.
a, Scheme for human, syngeneic tumour model, and bacterial study. b, c Box-and-whisker plot illustrating alpha diversity (b) and principal coordinate analysis plot illustrating beta diversity (c) of gut microbiomes in NSCLC patients (red, n = 96) and healthy controls (blue, n = 139). p value of alpha diversity calculated using Kruskal-Wallis test. Box represents first, third quartiles, and median, and whiskers are range up to 1.5×Interquartile Ranges (IQR). d, A plot of linear discriminant analysis (LDA) scores from the linear discriminant analysis effect size (LEfSe) method illustrating the differential abundance of the indicated taxa in the gut microbiomes of NSCLC patients (red) and healthy controls (blue) (LDA score > 4). e, The abundance of B. bifidum in NSCLC patients except epidermal growth factor receptor (EGFR) tyrosine kinase inhibitor (TKI)-treated patients was determined by qPCR (n = 80). Data show means ± SEM. p value calculated using two-sided unpaired Mann-Whitney U test, p = 0.0128. f, Effect of B. bifidum and Leuconostoc strains on in vitro stimulation of CD8+ T cells by monocytes. Secreted IFN-γ was measured by ELISA in co-cultures of human autologous T cells and monocytes treated with B. bifidum and Leuconostoc strains (Supplementary Table 5). 3 technical replicates. Data show means ± SEM. p values calculated using one-way ANOVA with Tukey for multiple comparison. B. bif_M31 versus L. mes_G16, p = 0.0040; B. bif_M31 versus L. mes_K20, p = 0.0245; B. bif_M31 versus L. gar_K21, p = 0.0301. For all graphs, *p < 0.05, and **p < 0.01.
Extended Data Fig. 2 Synergistic anti-tumor effect of B. bifidum strains in combination with oxaliplatin or anti-PD-1 in syngeneic mouse tumor models.
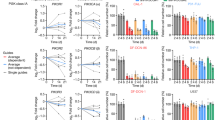
a, Abundance of B. bifidum in the mouse intestine after administration. Mice were administered a onetime dose of 1 × 109 CFU of B. bif_K57 and stool samples were collected at 4, 8, 12, and 24 hr after administration. B. bifidum abundance was determined using a qPCR assay with B. bifidum specific primers and normalized to 16S rRNA. n = 5 independent biological replicates. Data show means ± SEM. b, Flow cytometry analyses were performed on mouse spleen and tumor tissue. Upper: Heatmap represents the p values of differences in immune cell profile between oxaliplatin treatment and combined treatment with oxaliplatin and B. bifidum strains. p values calculated using two-sided unpaired t test. The exact p values are provided in Supplementary Table 14. Lower: Ratio of CD8+ T/Treg and effector CD8+ T/Treg in spleen and tumor. Ctrl, n = 9; Oxp or Oxp + B. bif_K57, n = 7; Oxp + B. bif_B06, n = 8 mice per group. Data show means ± SEM. p values calculated using one-way ANOVA with Tukey for multiple comparison. Oxp versus Oxp + B. bif_K57, p = 0.0002 (Ratio of CD8+ T/Treg and effector CD8+ T/Treg, tumor); Oxp versus Oxp + B. bif_B06, p = 0.0124 (Ratio of effector CD8+ T/Treg, tumor); Oxp versus Oxp + B. bif_K57, p = 0.0011 (Ratio of CD8+ T/Treg, spleen); Oxp versus Oxp + B. bif_K57, p = 0.0039 (Ratio of effector CD8+ T/Treg, spleen). c, Cytokine expression profiles in tumours from mice treated with oxaliplatin or both oxaliplatin and B. bifidum were measured by qPCR. Ctrl or Oxp groups, n = 6 independent biological replicates/group; oxaliplatin + B. bif_K57 or B. bif_ B06 groups, n = 8 independent biological replicates/group. Data show means ± SEM. p values calculated using one-way ANOVA with Tukey for multiple comparison within oxaliplatin treated groups. IFN-γ of Oxp versus Oxp + B. bif_K57, p = 0.0478; IL-2 of Oxp versus oxp + B. bif_K57, p < 0.0001; IL-2 of Oxp versus Oxp + B. bif_B06, p = 0.0006; TNF-α of Oxp versus Oxp + B. bif_K57, p = 0.0004; TNF-α of Oxp versus Oxp + B. bif_B06, p = 0.0040; IL-10 of Oxp versus Oxp + B. bif_K57, p < 0.0001; IL-10 of Oxp versus Oxp + B. bif_B06, p = 0.0063. d, Representative tumor growth curves after administration of B. bif_M31, with or without anti-PD-1 treatment. IgG, n = 8; anti-PD-1, n = 5; B. bif_M31, n = 10; anti-PD-1 + B. bif_M31, n = 9 mice per group. Data show means ± SEM. p value calculated using two-way ANOVA with Tukey for multiple comparison. anti-PD-1 versus anti-PD-1 + B. bif_M31, p < 0.0001. e, Representative tumor growth curves after administration of B. bif_C01, with or without anti-PD-1 treatment. IgG, n = 6; anti-PD-1 or B. bif_C01 or and anti-PD-1 + B. bif_C01, n = 7 mice per group. Data show means ± SEM. p value calculated using two-way ANOVA with Tukey for multiple comparison. f, Flow cytometry analysis of CD4+ T, NK and Treg cells in lamina propria of the intestine. n = 7 mice per group. Data show means ± SEM. p values calculated using one-way ANOVA with Tukey for multiple comparison. CD4+ T cells of anti-PD-1 versus anti-PD-1 + B. bif_K57, p = 0.0015; NK cells of anti-PD-1 versus anti-PD-1 + B. bif_K57, p = 0.0037. For all graphs, *p < 0.05, **p < 0.01, ***p < 0.001, and ns = not significant.
Extended Data Fig. 3 Transcriptomic and Lipidomic analysis of syngeneic tumor model treated with B. bifidum strains.
a, Network representation of enriched Gene Ontology (GO) biological pathways among differentially up-regulated genes in mice treated with anti-PD-1 + B. bif_K57 mice, determined using ClueGO. b, Principal component analysis of syngeneic mouse serum lipid profiles, based on lipids differential abundance among mice treated with anti-PD-1, anti-PD-1 + B. bif_B06, and anti-PD-1 + B. bif_K57 (>1.5-fold & p < 0.01). c, Heatmap of lipids showing significant differences (>1.5-fold & p < 0.01) in serum of syngeneic mice, as revealed by the lipidomic analysis. PE, phosphatidylethanolamine; PI, phosphoinositol; PC, phosphatidylcholine; LPG, lysophosphatidylglycerol; DG, diacylglycerol; TG, triglycerol.
Extended Data Fig. 4 Synergistic anti-tumor effect of B. bifidum strains in combination with anti-PD-1 in various syngeneic mouse tumor models.
a, Effect of B. bifidum strains on in vitro stimulation of CD8 T+ cells by monocytes. Secreted IFN-γ was measured by ELISA in co-cultures of human autologous T cells and monocytes treated with B. bif_K57 or B. bif_B06. 2 technical replicates. Data show means ± SEM. b, Upper: Experimental timeline: Animals were treated with the antibiotic cocktail (ABX) for 14 days. B. bif_K57 was then orally administered for 14 days before inoculation with MC38 colon cancer cells, followed by administration of anti-PD-1 via intraperitoneal injection twice a week for 21 days. Lower: MC38 tumor growth curves are shown for animals subjected to the indicated experimental treatments. IgG after PBS or anti-PD-1 after PBS, n = 5; anti-PD-1 + B. bif_K57 after PBS, n = 6; anti-PD-1 after ABX, n = 5; anti-PD-1 + B. bif_K57 after ABX, n = 7 mice per group. Data show means ± SEM. p values calculated using two-way ANOVA with Tukey for multiple comparison. anti-PD-1 after PBS versus anti-PD-1 + B. bif_K57 after PBS, p = 0.0053; anti-PD-1 after PBS versus anti-PD-1 + B. bif_K57 after ABX, p = 0.0004. c, Effect of ABX treatment on the abundance of gut microbiome. Stool 16S rRNA genes were analyzed by qPCR. n = 5 mice per group. Data show means ± SEM. p value calculated using two-sided unpaired t test, p < 0.0001. d, Upper: Experimental timeline: 14 days after initial oral administration of B. bif_K57, mice were inoculated with LLC1. anti-PD-1 was administered via intraperitoneal injection eight times, twice a week for 28 days. Lower: LLC1 tumor growth curves were generated for animals subjected to the indicated experimental treatments. n = 6 mice per group. Data show means ± SEM. p values calculated using two-way ANOVA with Tukey for multiple comparison. IgG versus anti-PD-1 + B. bif_K57, p < 0.0001. e, Macroscopic findings of tumor-inoculated lungs at day 23. The implanted tumors are indicated by yellow arrows. Scale bar, 50 mm. f, Upper: Experimental timeline: 14 days after initial oral administration of B. bif_K57, mice were inoculated with 4T1 breast cancer cells. anti-PD-1 was administered via intraperitoneal injection six times, twice a week for 21 days. Lower: 4T1 tumor growth curves were generated for animals subjected to the indicated experimental treatments. n = 10 mice per group. Data show means ± SEM. IgG versus anti-PD-1 + B. bif_K57, p < 0.0001. For all graphs, **p < 0.01, ***p < 0.001, and ns = not significant.
Extended Data Fig. 5 The abundance of peptidoglycan in B. bifidum strains.
The abundance of peptidoglycan of B. bif_K57 and B. bif_B06 was estimated using ELISA. Data show means ± SEM. 3 biological replicates. p value calculated using two-sided unpaired t test, p < 0.0006. For all graphs, ***p < 0.001.
Supplementary information
Supplementary Information
Supplementary Figs. 1–5.
Supplementary Tables
Supplementary Tables 1–14.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data and metabolomics source data.
Source Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Lipidomics source data.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 5
Statistical source data.
Rights and permissions
About this article
Cite this article
Lee, SH., Cho, SY., Yoon, Y. et al. Bifidobacterium bifidum strains synergize with immune checkpoint inhibitors to reduce tumour burden in mice. Nat Microbiol 6, 277–288 (2021). https://doi.org/10.1038/s41564-020-00831-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-020-00831-6
This article is cited by
-
Gut microbiome for predicting immune checkpoint blockade-associated adverse events
Genome Medicine (2024)
-
Unveiling the gastric microbiota: implications for gastric carcinogenesis, immune responses, and clinical prospects
Journal of Experimental & Clinical Cancer Research (2024)
-
Critical role of the gut microbiota in immune responses and cancer immunotherapy
Journal of Hematology & Oncology (2024)
-
Correlation of distribution characteristics and dynamic changes of gut microbiota with the efficacy of immunotherapy in EGFR-mutated non-small cell lung cancer
Journal of Translational Medicine (2024)
-
The impact of the gut microbiome on tumor immunotherapy: from mechanism to application strategies
Cell & Bioscience (2023)