Abstract
Tissue regeneration and maintenance rely on coordinated stem cell behaviours. This orchestration can be impaired by oncogenic mutations leading to cancer. However, it is largely unclear how oncogenes perturb stem cells’ orchestration to disrupt tissue. Here we used intravital imaging to investigate the mechanisms by which oncogenic Kras mutation causes tissue disruption in the hair follicle. Through longitudinally tracking hair follicles in live mice, we found that KrasG12D, a mutation that can lead to squamous cell carcinoma, induces epithelial tissue deformation in a spatiotemporally specific manner, linked with abnormal cell division and migration. Using a reporter mouse capture real-time ERK signal dynamics at the single-cell level, we discovered that KrasG12D, but not a closely related mutation HrasG12V, converts ERK signal in stem cells from pulsatile to sustained. Finally, we demonstrated that interrupting sustained ERK signal reverts KrasG12D-induced tissue deformation through modulating specific features of cell migration and division.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Source data are provided with this study. All other data that support the findings of this study are available from the corresponding author upon reasonable request.
References
Bissell, M. J. & Hines, W. C. Why don’t we get more cancer? A proposed role of the microenvironment in restraining cancer progression. Nat. Med. 17, 320–329 (2011).
Schneider, G., Schmidt-Supprian, M., Rad, R. & Saur, D. Tissue-specific tumorigenesis: context matters. Nat. Rev. Cancer 17, 239–253 (2017).
Kakiuchi, N. & Ogawa, S. Clonal expansion in non-cancer tissues. Nat. Rev. Cancer 21, 239–256 (2021).
Amberg, N. et al. Mouse models of nonmelanoma skin cancer. Methods Mol. Biol. 1267, 217–250 (2015).
Lapouge, G. et al. Identifying the cellular origin of squamous skin tumors. Proc. Natl Acad. Sci. USA 108, 7431–7436 (2011).
Latil, M. et al. Cell-type-specific chromatin states differentially prime squamous cell carcinoma tumor-initiating cells for epithelial to mesenchymal transition. Cell Stem Cell 20, 191–204.e195 (2017).
White, A. C. et al. Stem cell quiescence acts as a tumour suppressor in squamous tumours. Nat. Cell Biol. 16, 99–107 (2014).
White, A. C. et al. Defining the origins of Ras/p53-mediated squamous cell carcinoma. Proc. Natl Acad. Sci. USA 108, 7425–7430 (2011).
Brown, S. et al. Correction of aberrant growth preserves tissue homeostasis. Nature 548, 334–337 (2017).
Pineda, C. M. et al. Hair follicle regeneration suppresses Ras-driven oncogenic growth. J. Cell Biol. 218, 3212–3222 (2019).
Joost, S. et al. The molecular anatomy of mouse skin during hair growth and rest. Cell Stem Cell 26, 441–457.e447 (2020).
Lee, J. & Tumbar, T. Hairy tale of signaling in hair follicle development and cycling. Semin. Cell Dev. Biol. 23, 906–916 (2012).
Rompolas, P. et al. Live imaging of stem cell and progeny behaviour in physiological hair-follicle regeneration. Nature 487, 496–499 (2012).
Rompolas, P., Mesa, K. R. & Greco, V. Spatial organization within a niche as a determinant of stem-cell fate. Nature 502, 513–518 (2013).
Xin, T., Gonzalez, D., Rompolas, P. & Greco, V. Flexible fate determination ensures robust differentiation in the hair follicle. Nat. Cell Biol. 20, 1361–1369 (2018).
Sequeira, I. & Nicolas, J. F. Redefining the structure of the hair follicle by 3D clonal analysis. Development 139, 3741–3751 (2012).
Morrow, A., Underwood, J., Seldin, L., Hinnant, T. & Lechler, T. Regulated spindle orientation buffers tissue growth in the epidermis. eLife 8, e48482 (2019).
Aikin, T. J., Peterson, A. F., Pokrass, M. J., Clark, H. R. & Regot, S. MAPK activity dynamics regulate non-cell autonomous effects of oncogene expression. eLife 9, e60541 (2020).
Aoki, K. et al. Propagating wave of ERK activation orients collective cell migration. Dev. Cell 43, 305–317.e305 (2017).
De Simone, A. et al. Control of osteoblast regeneration by a train of Erk activity waves. Nature 590, 129–133 (2021).
Hino, N. et al. ERK-mediated mechanochemical waves direct collective cell polarization. Dev. Cell 53, 646–660.e648 (2020).
Hiratsuka, T. et al. Intercellular propagation of extracellular signal-regulated kinase activation revealed by in vivo imaging of mouse skin. eLife 4, e05178 (2015).
Pokrass, M. J. et al. Cell-cycle-dependent ERK signaling dynamics direct fate specification in the mammalian preimplantation embryo. Dev. Cell 55, 328–340.e325 (2020).
Regot, S., Hughey, J. J., Bajar, B. T., Carrasco, S. & Covert, M. W. High-sensitivity measurements of multiple kinase activities in live single cells. Cell 157, 1724–1734 (2014).
Li, S., Balmain, A. & Counter, C. M. A model for RAS mutation patterns in cancers: finding the sweet spot. Nat. Rev. Cancer 18, 767–777 (2018).
Johnson, H. E. & Toettcher, J. E. Signaling dynamics control cell fate in the early Drosophila embryo. Dev. Cell 48, 361–370.e363 (2019).
Marshall, C. J. Specificity of receptor tyrosine kinase signaling: transient versus sustained extracellular signal-regulated kinase activation. Cell 80, 179–185 (1995).
Gagliardi, P. A. et al. Collective ERK/Akt activity waves orchestrate epithelial homeostasis by driving apoptosis-induced survival. Dev. Cell 56, 1712–1726.e1716 (2021).
Pond, K. W. et al. Live-cell imaging in human colonic monolayers reveals ERK waves limit the stem cell compartment to maintain epithelial homeostasis. eLife 11, e78837 (2022).
Valon, L. et al. Robustness of epithelial sealing is an emerging property of local ERK feedback driven by cell elimination. Dev. Cell 56, 1700–1711.e1708 (2021).
Bugaj, L. J. et al. Cancer mutations and targeted drugs can disrupt dynamic signal encoding by the Ras-Erk pathway. Science 361, eaao3048 (2018).
Dessauges, C. et al. Optogenetic actuator – ERK biosensor circuits identify MAPK network nodes that shape ERK dynamics. Mol. Syst. Biol. 18, e10670 (2022).
Hiratsuka, T., Bordeu, I., Pruessner, G. & Watt, F. M. Regulation of ERK basal and pulsatile activity control proliferation and exit from the stem cell compartment in mammalian epidermis. Proc. Natl Acad. Sci. USA 117, 17796–17807 (2020).
Aoki, K. et al. Stochastic ERK activation induced by noise and cell-to-cell propagation regulates cell density-dependent proliferation. Mol. Cell 52, 529–540 (2013).
Sugawara, K. et al. Spatial and temporal control of laminin-332 (5) and -511 (10) expression during induction of anagen hair growth. J. Histochem. Cytochem. 55, 43–55 (2007).
Rezza, A. et al. Signaling networks among stem cell precursors, transit-amplifying progenitors, and their niche in developing hair follicles. Cell Rep. 14, 3001–3018 (2016).
Tumbar, T. et al. Defining the epithelial stem cell niche in skin. Science 303, 359–363 (2004).
Mesa, K. R. et al. Niche-induced cell death and epithelial phagocytosis regulate hair follicle stem cell pool. Nature 522, 94–97 (2015).
Madisen, L. et al. A robust and high-throughput Cre reporting and characterization system for the whole mouse brain. Nat. Neurosci. 13, 133–140 (2010).
Jackson, E. L. et al. Analysis of lung tumor initiation and progression using conditional expression of oncogenic K-ras. Genes Dev. 15, 3243–3248 (2001).
Chen, X. et al. Endogenous expression of Hras(G12V) induces developmental defects and neoplasms with copy number imbalances of the oncogene. Proc. Natl Acad. Sci. USA 106, 7979–7984 (2009).
Barker, N. et al. Identification of stem cells in small intestine and colon by marker gene Lgr5. Nature 449, 1003–1007 (2007).
Muller-Rover, S. et al. A comprehensive guide for the accurate classification of murine hair follicles in distinct hair cycle stages. J. Invest. Dermatol. 117, 3–15 (2001).
Chi, W., Wu, E. & Morgan, B. A. Dermal papilla cell number specifies hair size, shape and cycling and its reduction causes follicular decline. Development 140, 1676–1683 (2013).
Greco, V. et al. A two-step mechanism for stem cell activation during hair regeneration. Cell Stem Cell 4, 155–169 (2009).
Acknowledgements
We thank the Greco Lab members for helpful discussion and thoughtful feedback. We thank H. Clevers for the Lgr5-IRES-CreER mice, E. Fuchs for the K14-H2BGFP mice, M. D. Muzumdar for the LoxP-STOP-LoxP-KrasG12D mice and S. Beronja for the LoxP-Hras-LoxP-HrasG12V mice. V.G. was a New York Stem Cell Foundation Robertson Investigator and HHMI Scholar. T.X. was supported by the New York Stem Cell Foundation Druckenmiller Fellowship and the Dermatology Foundation Research Grant. S.G. was supported by the Human Frontier Science Program. The Regot Lab is supported by a National Institutes of Health (NIH) NIGMS R35 (R35GM133499) and an NIH NCI R01 (R01CA279546). Research reported in this publication was supported by the National Institute of Arthritis and Musculoskeletal and Skin Diseases of the NIH under award number R01AR063663 (V.G. as principal investigator) and R01AR067755 (V.G. as principal investigator), and the National Institute on Aging of the NIH under award number DP1AG066590 (V.G. as principal investigator). The content is solely the responsibility of the authors and does not necessarily represent the official views of the NIH.
Author information
Authors and Affiliations
Contributions
T.X., S.R. and V.G. designed the experiments. T.X. performed the experiments and analysed the data. S.G. characterized and maintained the KrasG12D and HrasG12V mice. H.W. recorded some of the time lapses after MEKi injection. D.G.G. assisted with the two-photon imaging and performed cell migration analysis. C.M.-M. performed FACS work. H.M., H.A.P. and H.F. assisted with data analysis. T.X., K.C.S., L.E.G., S.R. and V.G. wrote the paper with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Cell Biology thanks Michiyuki Matsuda, Michael Rendl and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Information on hair cycle and hair type for data interpretation.
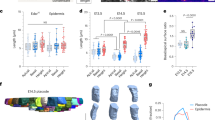
a, Schematic showing different stages of the hair follicle regeneration cycle. b, Representative images and percentages of each hair type in the region of the mouse ear skin where intravital imaging was conducted. c, Representative two-photon images of the KrasG12D hair follicles in the same mouse 13 days after induction. In the skin region that hair follicles entered late growth stages, bump-like deformation emerged in the ORS, while in the area of early and middle growth stages, hair follicles were normal. Scale bar, 20 µm.
Extended Data Fig. 2 Characteristics of ERK signal dynamics and expression levels of Kras and Hras.
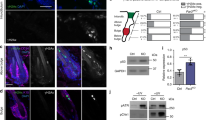
a, Representative two-photon time lapse frames of the wild type late growth hair follicle expressing the ERK biosensor showing wave-like ERK signal propagation in the ORS. In this example, the wave initiated from the middle ORS and propagated towards both the upper and lower ORS. Scale bar, 20 µm. b, Back skin cells at Anagen were processed for flow cytometry and gated for ORS cells using Lgr5-GFP and K14-H2BmCherry. Sorted ORS cells were then used to conduct qRT-PCR to compare the expression levels between Kras and Hras. n = 4 mice. ns, not significant, p = 0.0852. Two-sided paired t-test was used to calculate p value. Scale bar, 20 µm. c, Pulsing frequency of the ERK signal in the upper and lower ORS cells of the control and KrasG12D hair follicles. n = 236 upper and 162 lower ORS cells in 3 wild type mice, 1666 upper and 91 lower ORS cells in KrasG12D mice. ns, not significant, p = 0.6385 and 0.0571. Two-sided unpaired t-test was used to calculate p value. d, Cumulative ERK activity of the upper and lower ORS cells of the control and KrasG12D hair follicles. The same cells in c were analysed. ns, not significant, p = 0.5064, **, p = 0.0058. Two-sided unpaired t-test was used to calculate p value. Data are presented as mean with individual data points in c and violin plots with median and quartiles in d. Scale bar, 20 µm.
Extended Data Fig. 3 MEKi injection temporarily inhibits ERK without promoting apoptosis or differentiation.
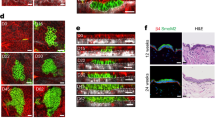
a, Representative two-photon time lapse frames of the wild type late growth hair follicles expressing the ERK biosensor 3 hours after intradermal injection of MEKi. Note that ERK activation began to emerge in the ORS (arrowheads) shortly after the time lapse started. b, Representative KrasG12D hair follicles treated with vehicle or MEKi in the whole mount skin stained for cleaved-Caspase3 (C-CASP3, red) and cell nuclei (DAPI, cyan). Apoptotic cell is indicated by arrowhead. c, Average numbers of apoptotic cells in the KrasG12D hair follicles treated with vehicle or MEKi. n = 3 mice (102 hair follicles from the skin treated with vehicle and 115 hair follicles from the skin treated with MEKi). ns, not significant, p = 0.1964. d, Representative images of the control and KrasG12D hair follicles stained for basal marker K14 or differentiation markers K75 and GATA3 (green). Cell nuclei were labelled by SiR-DNA or DAPI (magenta). Representative tissue deformations in the ORS are indicated by arrowheads. e, Representative images of the KrasG12D hair follicles after MEKi treatment stained for differentiation markers K75 and GATA3 (green). Cell nuclei were labelled by DAPI (magenta). Representative tissue deformations in the ORS are indicated by arrowheads. Border of the hair follicle is marked by white dashed lines in b, d and e. Two-sided unpaired t-test was used to calculate p values. Data are presented as mean ±S.D. with individual data points in c. Scale bars, 20 µm.
Supplementary information
Supplementary Information
Legends for Supplementary Videos 1–3.
Supplementary Video 1
Two-photon time lapse of a control and a KrasG12D late growth hair follicle expressing epithelial nuclei marker (K14-H2BmCherry) to compare cell migration.
Supplementary Video 2
Two-photon time lapse of the representative control, KrasG12D and HrasG12V late growth hair follicle expressing ERK sensor (ERK-KTRmClover).
Supplementary Video 3
Two-photon time lapse of the representative KrasG12D late growth hair follicle expressing epithelial nuclei marker (K14-H2BGFP) after MEKi treatment.
Source data
Source Data Fig. 1
Source Data for Fig. 1.
Source Data Fig. 2
Source Data for Fig. 2.
Source Data Fig. 3
Source Data for Fig. 3.
Source Data Fig. 4
Source Data for Fig. 4.
Source Data Fig. 5
Source Data for Fig. 5.
Source Data Extended Data Fig. 2
Source Data Extended Data for Fig. 2.
Source Data Extended Data Fig. 3
Source Data Extended Data for Fig. 3.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Xin, T., Gallini, S., Wei, H. et al. Oncogenic Kras induces spatiotemporally specific tissue deformation through converting pulsatile into sustained ERK activation. Nat Cell Biol (2024). https://doi.org/10.1038/s41556-024-01413-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41556-024-01413-y