Abstract
The myeloid transcription factor CEBPA is recurrently biallelically mutated (i.e., double mutated; CEBPADM) in acute myeloid leukemia (AML) with a combination of hypermorphic N-terminal mutations (CEBPANT), promoting expression of the leukemia-associated p30 isoform, and amorphic C-terminal mutations. The most frequently co-mutated genes in CEBPADM AML are GATA2 and TET2, however the molecular mechanisms underlying this co-mutational spectrum are incomplete. By combining transcriptomic and epigenomic analyses of CEBPA-TET2 co-mutated patients with models thereof, we identify GATA2 as a conserved target of the CEBPA-TET2 mutational axis, providing a rationale for the mutational spectra in CEBPADM AML. Elevated CEBPA levels, driven by CEBPANT, mediate recruitment of TET2 to the Gata2 distal hematopoietic enhancer thereby increasing Gata2 expression. Concurrent loss of TET2 in CEBPADM AML induces a competitive advantage by increasing Gata2 promoter methylation, thereby rebalancing GATA2 levels. Of clinical relevance, demethylating treatment of Cebpa-Tet2 co-mutated AML restores Gata2 levels and prolongs disease latency.
Similar content being viewed by others
Introduction
Acute myeloid leukemia (AML) is characterized by genetic alterations affecting the proliferation and/or differentiation of hematopoietic stem or progenitor cells (HSPCs). Thereby, the expansion of immature myeloid precursors, at the expense of normal hematopoiesis, ultimately leads to bone marrow (BM) failure if left untreated. Recent sequencing efforts have identified numerous recurrent mutations in AML and revealed patterns of mutational co-segregation, suggesting that synergism between certain lesions drives leukemogenesis1. While we now recognize these patterns, the mechanistic basis for context-specific positive or negative selection of certain lesions remains to be elucidated in most cases.
CCAAT enhancer binding protein alpha (CEBPA) is a hematopoietic lineage-specific transcription factor that binds and primes genes for myeloid development and is required for differentiation and maturation of granulocytes2. The gene encoding CEBPA is biallelically mutated (i.e., double mutated; CEBPADM) in 3–15% of de novo AML patients3,4,5,6,7,8,9. CEBPADM patients harbor either biallelic N-terminal mutations or a combination of a monoallelic N-terminal mutation together with a C-terminal mutation in the other allele. Whereas N-terminal CEBPA (CEBPANT) lesions promote the expression of the truncated p30 isoform, C-terminal mutations result in CEBPA variants that are unable to dimerize or bind DNA, thus rendering them inactive. Hence, CEBPA p30 homodimers are the sole entity with functional transcription factor activity in CEBPADM AML. This is in contrast to normal hematopoietic cells where the full-length p42 isoform is predominantly expressed2. CEBPA p30 lacks two of three transactivation elements present in p42, but retains one transcriptional activating element and the basic-region leucine-zipper, which enables dimerization and DNA-binding10. CEBPA p30 has functions distinct from CEBPA p42 and can bind an isoform-specific set of enhancers and regulate the expression of downstream effector genes, such as Nt5e and Msi211,12. Importantly, in the context of CEBPADM AML, the CEBPANT is hypermorphic, leading to higher levels of the transcription factor, and thus, increased binding to enhancers and subsequent deregulation of gene expression11. In line with these data, mice with CEBPA p30 expression driven from the endogenous Cebpa locus develop AML with full penetrance within a year13.
Most patients with CEBPADM AML also feature additional mutations in GATA2, TET2, WT1, NRAS, FLT3, or CSF3R9. Several of these mutations are found together with CEBPADM more frequently than expected by the individual frequency of each mutation, while other combinations are statistically underrepresented. Recent studies have shed light on the molecular mechanisms underlying mutational cooperativity for some of the co-mutated genes, i.e. GATA214 and CSF3R15, while mechanistic insight is still lacking for other subgroups of CEBPADM AML. Of particular importance are mutations in the gene encoding the methylcytosine dioxygenase TET2 which, by converting 5-methylcytosine to 5-hydroxymethylcytosine, promotes DNA demethylation. TET2 mutations (TET2MUT) are frequent in CEBPADM AML cases and are associated with inferior prognosis16,17. Moreover, loss of Tet2 has been implicated in accelerating and/or aggravating hematological malignancies in combination with several other recurrent gain-of-function and loss-of-function mutations18,19,20, reflecting the importance of appropriately regulated DNA demethylation in normal hematopoiesis. Importantly, while Tet2 loss alone only mildly affects hematopoiesis with myeloid skewing and increased competitiveness of HSCs18, as well as the increased propensity of leukemic blasts to switch to a more stem-like phenotype21, it does not induce overt leukemia per se22,23,24. Despite being extensively studied, mechanistic insights of how TET2 loss-of-function cooperates with other aberrations have been hampered by the fact that malignant cells have been compared to their normal, wild-type counterparts in many studies.
In the present work, we sought to overcome this limitation by comparing CEBPA-mutant AML in the presence and absence of additional mutations in TET2. By combining transcriptomic and epigenomic analyses of relevant in vitro and in vivo models as well as data from AML patients, we identified an intricate mechanism where TET2 loss-of-function rebalances Gata2 expression levels in CebpaDM AML, and hence drives an aggressive disease.
Results
TET2 mutations impair outcome for patients with CEBPA-mutant AML
To validate previous reports on the spectrum of co-occurring mutations in CEBPADM AML patients, we compiled data from 557 CEBPADM cases and evaluated the co-occurrence of other known leukemia driver mutations3,4,5,6,7,17. TET2 was the second most frequently co-mutated gene, with 1 in 5 CEBPADM cases harboring TET2 mutations (Fig. 1a; Supplemental Table 1). Importantly, the survival of TET2-mutant (TET2MUT) CEBPADM patients was significantly lower than TET2 wild-type (TET2WT) CEBPADM patients (Fig. 1b), consistent with previous reports16, while the presence of TET2 mutations did not cause a higher overall number of mutations in CEBPADM patients (Supplemental Fig. 1a).
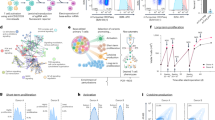
a Frequency of co-occurring mutations in CEBPADM AML cases, data aggregated from published cohorts3,4,5,6,7,8,17 (321–557 cases; detailed in Supplemental Table 1). b Overall survival of CEBPADM patients with wild-type (TET2WT; 84 patients) or mutated TET2 (TET2MUT; 35 patients). The data were analyzed by Mantel-Cox Log-rank test. c Volcano plot depicting differentially expressed genes dependent on TET2 mutational status in the cohort of CEBPA-mutant patients in the Beat AML dataset (TET2WT 11 and TET2MUT 5 patients). Differential analysis was performed with DESeq2 (P < 0.05). Source data are provided as a Source Data file.
To investigate the functional consequences of TET2 and CEBPA co-mutations, we analyzed RNA sequencing (RNA-seq) data from the Beat AML dataset1. We identified 1546 up- and 1201 downregulated genes in patients harboring a combination of CEBPA and TET2 mutations when compared to CEBPA-mutant patients with wild-type TET2 (Fig. 1c). Similarly, a slight overrepresentation of up-regulated genes was observed when comparing CEBPAWTTET2MUT patients to CEBPAWTTET2WT patients (601 up- and 527 downregulated). In line with the lower overall survival of TET2MUTCEBPADM patients, pathways related to inflammation, hypoxia, and aggressive cancer were upregulated in CEBPA-TET2 co-mutated patients (Supplemental Fig. 1b). The overrepresentation of up-regulated genes associated with TET2 deficiency in CEBPAMUT (and CEBPAWT) patients is somewhat surprising, as increased DNA methylation upon TET2 loss would be expected to cause global transcriptional repression. However, other co-occurring mutations and residual DNA demethylase activity from the TET2WT allele may cause a more complex pattern of gene expression.
These findings indicate that mutations in TET2 enhance the aggressiveness of CEBPA-mutant AML by deregulation of critical cellular pathways.
TET2 deficiency accelerates Cebpa-mutant AML
To study the effect of TET2 mutations in CEBPADM AML in pathophysiologically relevant in vitro and in vivo models, we utilized cell and murine models in which expression of the p30 isoform is retained (Cebpap30/p30 or CebpaΔ/p30), while the normal p42 isoform of CEBPA is completely lost13. Since TET2 is predominantly inactivated by loss-of-function mutations25, we modeled TET2 mutations either by the introduction of mutations with the CRISPR-Cas9 technology or by conditional knockout of the Tet2 alleles.
First, we introduced Tet2 mutations into a murine myeloid progenitor cell model (Cebpap30/p30) (Fig. 2a). Tet2-targeted cells displayed a selective advantage, as they outcompeted Cebpap30/p30 cells (Fig. 2b). Detailed analysis of the Tet2 mutation that was associated with the proliferative advantage showed that the Tet2 locus had acquired a + 1 insertion in exon 3, which resulted in a downstream premature termination codon (Supplemental Fig. 2a, b). In line with this, clones isolated from the targeted cell pool exhibited strongly reduced TET2 protein expression (Supplemental Fig. 2c). Gene expression analysis revealed that Tet2 loss in Cebpap30/p30 cells caused downregulated expression of 916 genes, while only 540 genes were upregulated (Fig. 2c). Gene set enrichment analysis (GSEA) showed higher expression of MYC and E2F targets in Cebpap30/p30 Tet2-mutated cells, consistent with their proliferative advantage (Supplemental Fig. 2d).
a Schematic representation of generation of Tet2-knockout clones with CRISPR/Cas9. The illustration was created with BioRender.com. b Proliferative outgrowth of Cebpap30/p30 cells with Tet2 indels. c Volcano plot depicting differentially expressed genes dependent on the Tet2 mutational status in Cebpap30/p30 cells (Tet2WT 7 and Tet2MUT 5 clones). Differential analysis was performed with DESeq2 (P < 0.05). d Experimental setup for evaluating the effect of Tet2-deficiency (Tet2Δ/Δ) in CebpaDM AML initiation in vivo. The illustration was created with BioRender.com. e Myeloid (Mac1+) contribution of donor-derived blood and bone marrow (BM) cells evaluated after BM transplantation and Cre-LoxP recombination. (Blood samples: Week 12; 6 mice per group. Week 24; CebpaΔ/p30Tet2+/+ and Cebpa+/ΔTet2Δ/Δ 6 mice per group and Cebpa+/ΔTet2Δ/Δ 3 mice. Week 36; 3 mice per group. BM samples: 3 mice per group.) Data are presented as mean±SEM and analyzed by one-way-ANOVA followed by Dunnett’s multiple comparisons correction. f Survival of lethally irradiated recipient mice after BM transplantation and Cre-LoxP recombination (CebpaΔ/p30Tet2+/+ 12 mice, CebpaΔ/p30Tet2Δ/Δ 14 mice, and Cebpa+/-Tet2Δ/Δ 14 mice). The data were analyzed by Mantel-Cox Log-rank test. g Volcano plot depicting differentially expressed genes dependent on Tet2 deficiency status in CebpaΔ/p30 leukemic blasts (samples from 3 mice per group). Differential analysis was performed with DESeq2 (P < 0.05). h Frequency of proliferating (Ki67+) cells in BM of moribund recipient mice (specimens from 3 mice per group). Data are presented as mean±SEM and analyzed by a two-tailed unpaired t-test. Source data are provided as a Source Data file.
In summary, these data show that CRISPR/Cas9-induced TET2 loss provides a competitive advantage to myeloid progenitors expressing the oncogenic CEBPA variant p30.
Next, we wanted to assess the impact of hematopoietic expression of CEBPA p30 (CebpaΔ/p30) with TET2-deficiency (Tet2−/−) on AML initiation in vivo. To do so, we transplanted lethally irradiated recipient mice with BM cells derived from mice with relevant allele combinations and, following hematopoietic reconstitution, induced hematopoietic-specific knockout of the Cebpa WT allele and/or the Tet2 alleles (Fig. 2d). The combination of CEBPA p30 expression with Tet2 loss led to an early expansion of myeloid (Mac1+) cells in the BM and blood compared to mice with hematopoietic cells featuring either alteration on its own (Fig. 2e; Supplemental Fig. 2e). Conforming to patient data and data obtained from Cebpap30/30 cells, CebpaΔ/p30Tet2Δ/Δ hematopoietic cells gave rise to AML with shorter latency than CebpaΔ/p30Tet2+/+ cells, with a median survival of 23 and 43 weeks, respectively (Fig. 2f). Mice transplanted with CebpaΔ/p30Tet2+/+ BM cells developed leukemia with similar latency as mice transplanted with Cebpap30/p30 fetal liver cells13. This is consistent with the matching expression of Cebpa in these two contexts (1.1 ± 0.24 vs. 1.0 ± 0.13 (relative expression) in Cebpap30/p30 and CebpaΔ/p30Tet2+/+ AML blasts n = 3/group, respectively). TET2 deficiency alone (CebpaΔ/+Tet2Δ/Δ) did not give rise to AML and cells which retained expression of the p42 isoform from one allele (Cebpa+/p30) only sporadically underwent leukemic transformation, in line with unaltered Cebpa expression levels in these cells (Fig. 2f; Supplemental Fig. 2f; 1.03 ± 0.14 vs. 1.0 ± 0.04 (relative expression) in Cebpafl/p30 and Cebpafl/+ cells n = 2–3/group, respectively). The transformed blasts expressed myeloid (Mac1+) and granulocytic (Gr1+) markers, confirming the myeloid origin of the leukemia (Supplemental Fig. 2g). The leukemias were transplantable into secondary recipients, and the shorter latency of the TET2-deficient CebpaDM AML was preserved in this setting (Supplemental Fig. 2h–i), indicating that TET2 not only has important tumor suppressive functions during malignant transformation but also during progression of AML.
We performed RNA-seq on CebpaΔ/p30 (Tet2 WT and knockout) AML blasts to assess changes in gene expression upon TET2 deficiency. Again, we found that the majority of differentially expressed genes was decreased in TET2-deficient AML blasts, with 176 down- vs. 58 upregulated genes (Fig. 2g). GSEA highlighted upregulation of genes involved in IL-6-JAK-STAT-signaling and hypoxia, in line with RNA-seq data from human TET2MUTCEBPAMUT cases (Supplemental Fig. 1b; Supplemental Fig. 2j). Furthermore, pathways related to cell cycle progression (G2M checkpoint and E2F targets) were enriched in TET2-deficient AML, indicating increased growth upon loss of TET2, consistent with the effects observed in the cell model (Supplemental Fig. 2d; Supplemental Fig. 2j). In line with this, we found that a higher frequency of CebpaΔ/p30Tet2Δ/Δ blasts expressed the proliferation marker Ki67 (Fig. 2h). In addition, we also observed increased proliferative capacity of CebpaΔ/p30Tet2Δ/Δ blasts compared to CebpaΔ/p30Tet2+/+ blasts ex vivo. This difference was dependent on Tet2 status, as the TET2 co-factor Vitamin C was able to mitigate proliferation of CebpaΔ/p30Tet2+/+ but not of CebpaΔ/p30TetΔ/Δ cells (Supplemental Fig. 2k).
Collectively, these data show that TET2 deficiency accelerates the establishment and progression of CEBPA p30-driven AML in vivo.
Loss of TET2 leads to reduced Gata2 levels in Cebpa-mutant AML
To find conserved gene targets of the CEBPA-TET2 axis, we integrated the transcriptomic data from our in vitro and in vivo models with gene expression analyses from AML patients harboring CEBPA and TET2 mutations. Three target genes exhibited downregulated expression in all three data sets; FUT8, GATA2, and SIRT5 (Fig. 3a; Supplemental Fig. 3a–c).
a Conserved targets of the CEBPA-TET2 axis visualized in a Venn-diagram of downregulated genes in CEBPA-TET2 co-mutated AML overlaid with corresponding data from Tet2-deficient in vivo and in vitro models of CebpaDM AML (P = 0.0264 vs. number of overlapping genes expected by random distribution assessed by Wilson/Brown binominal test). b Heatmap of differentially accessible regions assessed by assay for transposase-accessible chromatin sequencing (ATAC-seq; FDR < 0.05), and c genomic distribution of downregulated peaks (FDR < 0.05, Log2FC < 0) upon Tet2 mutation (4 clones per group). Differential analysis was performed with DiffBind and region enrichment analysis with GREAT. d Representative genome wide DNA-methylation status in leukemic blasts from the in vivo model assessed by whole genome bisulfite sequencing (WGBS) showing frequency of methyl-cytosine (mC) across the transcription start site (TSS) ±1000 base pairs, gene body scaled to 4000 base pairs, and transcription termination site (TES) ±1000 base pairs. Methylation analysis was performed with Bismark and visualized using deepTools. e Median and interquartile range of percent mC at promoters of down- (n = 172), not expressed (n = 6539), not differentially expressed (not DE; n = 14759) and up-regulated (n = 57) genes (averaged data generated from 2 CebpaΔ/p30Tet2+/+ and 3 CebpaΔ/p30Tet2Δ/Δ mice). Whiskers indicates max–min and data were analyzed by two-tailed unpaired t-test. f Promoter DNA methylation of conserved target genes in leukemic blast (samples from 2 CebpaΔ/p30Tet2+/+ and 3 CebpaΔ/p30Tet2Δ/Δ mice). Data are presented as mean±SEM. The data were log-transformed and analyzed by two-tailed unpaired t-test. g Gata2 variant mRNA expression in CebpaΔ/p30Tet2Δ/Δ and CebpaΔ/p30Tet2+/+ leukemic blasts (samples from 3 mice per group). Data are presented as mean ± SEM. The data were log-transformed and analyzed by a two-tailed unpaired t-test. h Schematic genomic view of the Gata2 locus, including representative examples of assay for transposase-accessible chromatin using sequencing (ATAC-seq) in Cebpap30/p30 cells. Source data are provided as a Source Data file.
Since the deregulation of these three genes was observed across species and differential experimental setups, we next aimed to investigate if their decreased gene expression was a direct result of TET2 deficiency. We therefore assessed chromatin accessibility and DNA methylation as a proxy for TET2 binding and activity26. Through assay for transposase-accessible chromatin sequencing (ATAC-seq), we identified 1809 differentially accessible regions in Cebpap30/p30Tet2MUT vs. Cebpap30/p30Tet2WT cells, and consistent with an activating effect of TET2, the majority of differential regions were less accessible in TET2-deficient cells (Fig. 3b). Half of the ATAC-seq peaks downregulated upon Tet2 mutation were located in promoters, and these regions were enriched for GATA and NFAT motifs (Fig. 3c; Supplemental Fig. 3d). Using whole genome bisulfite sequencing (WGBS), we observed a global increase in DNA methylation in CebpaΔ/p30Tet2Δ/Δ vs. CebpaΔ/p30Tet2+/+ AML blasts, consistent with a loss of demethylase activity in Tet2 knockout blasts (Fig. 3d). Increased DNA methylation was observed in promoter regions of genes whose expression were downregulated upon TET2 loss ( + 54%; Fig. 3e), while upregulated and not differently expressed genes did not show any marked changes. Strikingly, this pattern was not apparent when DNA methylation was evaluated across gene bodies (Supplemental Fig. 3e). Non-expressed genes exhibited equal increase in DNA methylation across promoters and gene bodies (Fig. 3e; Supplemental Fig. 3e). Since increased gene body methylation is not associated with gene repression27, we evaluated whether a gain in gene body methylation was coupled to a gain in promoter methylation for the down-regulated genes. In the presence of promoter hypermethylation, the bodies of down-regulated genes were more prevalently hypermethylated compared to neutral and up-regulated genes (34.8% [95%CI 18.8–55.11] vs. 18.1% [16.4–19.9], p = 0.0427). While, in the absence of promoter hypermethylation, the bodies of up-regulated genes tended to be hypermethylated compared to neutral and down-regulated genes (13.8% [95%CI 7.2–24.9] vs. 7.1% [6.8–7.4], p = 0.0518). Thus, loss of TET2 in CebpaDM cells caused decreased chromatin accessibility and increased methylation of DNA in promoters of TET2-responsive genes, consistent with previous reports showing that TET2 binding is enriched in promoters of TET2-regulated genes28.
To identify direct CEBPA-TET2 gene target(s), we evaluated the previously identified conserved candidates based on changes in DNA methylation of their promoters. Out of the three target genes, only the gene encoding the transcription factor GATA-binding factor 2 (GATA2) showed a gain of DNA methylation in the promoter of the gene variant 2 (Gata2 V2) upon TET2 deficiency ( + 46%; Fig. 3f). In line with this, specifically the Gata2 V2 mRNA isoform was downregulated in TET2-deficient CebpaDM AML blasts (−86%; Fig. 3g), while changes in mRNA expression and promoter methylation of Gata2 V1 did not reach statistical significance (Fig. 3f, g).
In summary, these analyses identify Gata2 (locus overview in Fig. 3h) as a conserved target of the CEBPA-TET2 axis across several settings. TET2 deficiency causes increased DNA methylation of the Gata2 promoter, resulting in reduced mRNA expression.
Moderate Gata2 reduction increases competitiveness of Cebpa-mutant AML
GATA2 is an essential transcription factor for hematopoietic cells and has profound effects on HSC maintenance. Moreover, it is recurrently mutated in AML29,30 and GATA2 lesions are overrepresented in CEBPADM AML8,16,31,32,33. Given these critical roles of GATA2, we next examined the consequences of reduced GATA2 levels in CEBPADM AML.
To test if reduced Gata2 expression would provide a competitive advantage in vivo, we set up an RNA-interference (RNAi) based competition assay (Fig. 4a) utilizing established Cebpap30/p30 leukemia cells, in which both Cebpa (+56–73%) and Gata2 (+45–56%) levels are increased modestly compared to primary CebpaΔ/p30Tet2+/+ blasts11,34. First, we identified four short hairpin RNAs (shRNA) which lowered Gata2 expression to a varying degree (Fig. 4b). Upon transplantation of shRNA-expressing cells, we observed a non-monotonic relationship between Gata2 expression levels and competitiveness, as measured by shGata2-to-shControl ratios. While efficient downregulation of Gata2 expression did not provide any competitive advantage to CebpaDM cells, moderate silencing imposed a three-fold increase in their ability to compete (Fig. 4c, d). Repetition of this experiment including only the most and least efficient shRNAs in a separate experiment yielded similar results (Supplemental Fig. 4a, b). These results were mirrored by increased expression of the proliferation marker Ki67 in cells expressing the least efficient Gata2-targeting shRNA but not the most efficient one (Supplemental Fig. 4c). To test if the same effects are observed in an in vitro setting, we targeted Gata2 in Cebpap30/p30 cells using the CRISPR/Cas9 approach. Gata2-targeted cells showed a proliferative advantage over Gata2WT cells, leading to their outgrowth (Fig. 4e, f). In accordance with previously published data that complete loss of Gata2 expression results in a loss of competitiveness35,36,37, we found that only clones with heterozygous Gata2 inactivation were viable, while clones with homozygous mutations in Gata2 could not be recovered (Supplemental Fig. 4d).
a Experimental setup for evaluating the effect of Gata2 knockdown, via short hairpin RNA (shRNA) mediated silencing, on Cebpap30/p30 leukemic cells in a competitive in vivo assay. The illustration was created with BioRender.com. b Gata2 mRNA in Cebpap30/p30 leukemic cells prior to transplantation. c Representative flow cytometry profiles of input and output of shControl (no knockdown), shGata2A (low knockdown), and shGata2D (high knockdown). d Competitive advantage of targeting shRNA (GFP+) vs. non-targeting shRNA (YFP+) cells in vivo assessed as by flow cytometry (Control 4, shGata2A 4, shGata2B 4, shGata2C 4, and shGata2D 3 mice). Data are presented as mean±SEM. Data were log-transformed and analyzed by one-way-ANOVA followed by Dunnett’s multiple comparisons correction. e Experimental setup for Gata2 CRISPR/Cas9 mutagenesis in Cebpap30/p30 cells, and outgrowth of heterozygous mutated clones. Percentages of Gata2 mutated clones are indicated. f Growth curve of Cebpap30/p30 clones with Gata2 mutation (Cebpap30/p30Gata2+/MUT, n = 10) or wild type Gata2 (Cebpap30/p30Gata2+/+, n = 3). Data are presented as mean±SEM and analyzed by two-tailed unpaired t-test. Red lines mark individual mutated clones. g Presence or absence of GATA2 mutations (GATA2MUT) in CEBPA double mutated (CEBPADM) AML cases with or without TET2 mutations (TET2MUT) in aggregated data from published cohorts3,4,5,7,8,17 (detailed in Supplemental Table 2a). Data were analyzed by Wilson/Brown binominal test. Source data are provided as a Source Data file.
If the pro-leukemogenic effect of TET2 mutations was, at least partly, caused by lowering GATA2 expression, we reasoned that concomitant mutations in both genes would be redundant and thus, the pattern of TET2 and GATA2 mutations would be mutually exclusive. Indeed, TET2MUTCEBPADM AML cases showed a lower frequency of GATA2 mutations than expected from the frequency of GATA2 mutations in TET2WTCEBPADM AML cases (Fig. 4g; Supplemental Table 2a), which was also true for all AML cases (3.5% in TET2MUT vs. 8.9% in TET2WT; Supplemental Fig. 4e; Supplemental Table 2b). Importantly, whereas mutations in WT1 followed the same pattern as TET2, CSF3R mutations appeared in equal frequency between TET2MUTCEBPADM and TET2WTCEBPADM AML cases, and ASXL1 mutations were increased in TET2MUTCEBPADM AML (Supplemental Fig. 4f; Supplemental Table 2c–e). While we favor a functional redundancy model, we cannot exclude that co-mutation of TET2 and GATA2 could induce synthetic lethality in AML cells, as Gata2-loss has been shown to induce terminal myeloid differentiation37.
Altogether, our data suggest that loss of TET2 in CebpaDM AML causes a moderate decrease in Gata2 expression, which in turn increases the competitive fitness of the leukemia. Hence, this indicates that TET2 and GATA2 mutations are partially redundant in CEBPADM AML, providing a mechanistic rationale for the mutational spectrum observed in this AML entity.
Increased CEBPA p30 binding to the Gata2 distal hematopoietic enhancer drives expression of Gata2 via TET2
We next asked if GATA2 expression is dependent on CEBPA mutational status. To this end, we exploited published transcriptomics data from human and mouse CEBPADM AML11. GATA2 expression was increased in human CEBPADM leukemic granulocyte/monocyte progenitors (GMPs) compared to GMPs from healthy donors ( + 77%; Supplemental Fig. 5a). Correspondingly, Gata2 was upregulated in murine Cebpap30/p30 leukemic GMPs as compared to normal GMPs ( + 43%; Fig. 5a). Likewise, analysis of AML patient data from the BEAT AML study1, revealed that both CEBPA and GATA2 expression were increased in CEBPANT AML compared to CEBPAWT AML ( + 91% and +37%, respectively), while GATA2 expression was reverted to CEBPAWT level in CEBPANTTET2MUT AML (Supplemental Fig. 5b, c). Since CEBPA is known to exert its transcription factor activity by binding to enhancers and thereby promote gene expression38, we assessed the binding of CEBPA to the crucial Gata2 distal hematopoietic enhancer (G2DHE; −77 kb in mouse) that governs Gata2 expression in hematopoietic stem and progenitor cells including GMPs11,39. Notably, we found substantially increased levels of CEBPA bound to the G2DHE in Cebpap30/p30 leukemic GMPs compared to their normal counterparts ( + 147%; Fig. 5b), while the binding levels associated with other known proximal and distal cis-regulatory elements of the Gata2 gene were unchanged (Supplemental Fig. 5d, e). Additionally, TET2 showed significant binding to the G2DHE in Cebpap30/p30 cells (Fig. 5c). However, DNA methylation at the G2DHE was low and unaltered upon Tet2 loss (Supplemental Fig. 5f). Importantly, CEBPA binding, as assessed by ChIP-qPCR, was not altered by introduction of Tet2 mutations in Cebpap30/p30 cells (Supplemental Fig. 5g).
a Gata2 mRNA expression in mouse Cebpap30/p30 leukemic granulocyte/monocyte progenitors (GMPs) vs normal GMPs (samples from 4 Cebpa+/+ and 2 Cebpap30/p30 mice) and, b CEBPA binding to the Gata2 distal hematopoietic enhancer (G2DHE; −77 kb) region (samples from 2 mice per group), data from Jakobsen et al.11. Data are presented as mean ± SEM. Differential analysis was performed with DESeq2 (P < 0.05). c Schematic genomic view of the Gata2 distal hematopoietic enhancer (G2DHE), including normalized chromatin immunoprecipitation sequencing (ChIP-seq) signal of CEBPA (data from Heyes et al.12), TET2 and H3K27Ac (data from Heyes et al.12), as well as assay for transposase-accessible chromatin using sequencing (ATAC-seq) in Cebpap30/p30 cells without (light blue) and with (green) mutation in Tet2. d Gata2 mRNA levels in response to targeting of the G2DHE by CRISPR-Cas9 in Cebpap30/p30 cells in vitro using indicated sgRNAs and e the averaged change in Gata2 mRNA levels of the 12 deletions (averaged data from 2 separate experiments). Data are presented as median ± range and analyzed by two-tailed Wilcoxon signed-rank test. f Experimental setup for evaluating the effects of Cebpa knockout on Gata2 V2 mRNA expression and DNA methylation of the CpG island at the promoter of Gata2 V2 in MLL-fusion driven AML (iMLL-AF9). The illustration was created with BioRender.com. g Gata2 V2 mRNA expression (leukemic cell lines generated from 2 separate mice were assayed on 2 separate days in 2–3 technical replicates each). Data are presented as mean ± SEM and the individual cell lines are indicated by circles or squares. Data were log-transformed and analyzed by two-tailed unpaired t-test. h DNA methylation of the Gata2 V2 promoter CpG-island (2 separate leukemic cell lines). Data are presented as median±range and the individual cell lines are indicated by circles or squares. i Experimental setup for evaluating the effects of Cebpa knockdown on TET2 binding to the G2DHE in Cebpap30/p30 cells with inducible expression of shRNA targeting Cebpa and control (Renilla), respectively. The illustration was created with BioRender.com. j TET2 binding to the G2DHE assessed by ChIP-qPCR (3 replicates per condition). Data are presented as mean±SEM and analyzed by two-tailed unpaired t-test. k Frequency of GATA2 and/or TET2 mutations (GATA2MUT and TET2MUT, respectively) in CEBPA high expressing (CEBPAHIGH; 45 cases) vs. CEBPA low expressing (CEBPALOW; 61 cases) AML cases, data from Beat AML cohort1. The distributions of GATA2WTTET2WT vs. GATA2MUT and/or TET2MUT cases were analyzed by Wilson/Brown binominal test. Source data are provided as a Source Data file.
These results prompted us to test if CEBPA binding to the G2DHE modulates Gata2 expression in CebpaDM AML. We deleted 250–500 bp fragments of the Gata2 enhancer encompassing the CEBPA binding site using the CRISPR/Cas9 approach in Cebpap30/p30 cells in vitro. Expression of total Gata2 mRNA, as well as both individual transcript variants, was decreased upon targeting the genomic region with strong CEBPA binding compared to non-targeting control (Fig. 5c–e, Supplemental Fig. 5h–k). In contrast, Gata2 expression was unchanged when G2DHE deletions were introduced in Cebpap30/p30Tet2MUT cells (Supplemental Fig. 5l, m). Combined, these data suggest that CEBPA binding to the G2DHE is important for promoting Gata2 expression in CebpaDM AML. Further, the G2DHE has been shown to primarily regulate expression of the hematopoietic specific Gata2 variant 2 (V2)40,41, conforming with our data that particularly the Gata2 V2 promoter displayed an increase in DNA methylation and that the Gata2 V2 mRNA was downregulated in TET2-deficient CebpaDM AML blasts (Fig. 3f–g).
Next, we tested if the reduction of CEBPA in AML cells influenced the expression and promoter DNA methylation of Gata2 V2. Given the dependence of CEBPADM AML on CEBPA for survival and maintenance, we utilized MLL-fusion-driven AML, in which CEBPA is dispensable for the maintenance of established leukemia42. Cre-mediated loss of Cebpa in leukemic cells expressing the inducible MLL-AF9 fusion-protein (iMLL-AF9+CebpaΔ/Δ; Fig. 5f) caused reduced Gata2 V2 mRNA levels compared to control cells (iMLL-AF9+Cebpafl/fl) ( − 72%; Fig. 5g). Importantly, the methylation frequency of the CpG island located at the Gata2 V2 promoter was increased in two separate leukemic lines ( + 186%; Fig. 5h; Supplemental Fig. 5n), suggesting that Gata2 V2 mRNA expression is regulated via the CEBPA-TET2 axis. Finally, we assessed TET2 binding to the G2DHE upon Cebpa knockdown in Cebpap30/p30 cells using ChIP-qPCR (Fig. 5i). Notably, we observed decreased TET2 binding to the G2DHE in cells expressing shCebpa compared to cells expressing control shRNA, verifying that CEBPA is important for recruitment of TET2 to the G2DHE (Fig. 5j).
In light of these findings, we asked whether elevated CEBPA level and not the CEBPA mutation(s) per se, drives the selective pressure for GATA2 and/or TET2 loss in AML to achieve moderate GATA2 levels that are optimal for leukemia growth. We therefore stratified AML cases in the Beat AML cohort1 based on CEBPA expression and assessed their GATA2 and TET2 mutational status. Indeed, the frequency of GATA2 and/or TET2 mutations was three-fold higher in CEBPAHIGH AML compared to the CEBPALOW samples (Fig. 5k). In line with previous data showing a hypermorphic effect of CEBPADM 11, the CEBPAHIGH group contained the majority of the CEBPA-mutant cases in the cohort (82 and 100% of CEBPASM and CEBPADM, respectively), while none of the cases in the CEBPALOW group were CEBPA-mutated.
In conclusion, our data show that elevated CEBPA binding to the G2DHE, driven by the hypermorphic effect of CebpaNT, increases TET2-mediated demethylation of the Gata2 promoter, which leads to elevated Gata2 levels in CebpaDM AML. In this context, CebpaDM AML cells gain a competitive advantage by loss of TET2, which in turn promotes an increase in DNA methylation at the Gata2 promoter resulting in the rebalancing of Gata2 levels.
Demethylating treatment restores Gata2 expression and prolongs survival in TET2-deficient Cebpa-mutant AML
Finally, we investigated if treatment with the demethylating agent 5-azacytidine (5-AZA) would be beneficial in TET2-deficient CEBPADM AML. Ex vivo treatment with 5-AZA restored Gata2 expression in CebpaΔ/p30Tet2Δ/Δ blasts to levels observed in CebpaΔ/p30Tet2+/+, while 5-AZA treatment did not affect Gata2 levels in CebpaΔ/p30Tet2+/+ cells (Supplemental Fig. 6a). Moreover, 5-AZA decreased the viability of blasts from both genotypes, although to a higher degree in the TET2-deficient setting (−82% and −40%, respectively, p < 0.01; Supplemental Fig. 6b).
To evaluate if the enhanced effect of 5-AZA treatment in TET2-deficient AML would also hold true in vivo, mice were transplanted with CebpaΔ/p30Tet2Δ/Δ or CebpaΔ/p30Tet2+/+ AML blasts and treated with 5-AZA for three consecutive days after disease establishment (Fig. 6a). While the blast frequency of TET2-deficient CebpaΔ/p30 AML decreased upon 5-AZA treatment ( − 62%; Fig. 6b), the treatment did not significantly decrease the frequency of TET2-proficient cells. Furthermore, 5-AZA treatment restored Gata2 levels in CebpaΔ/p30Tet2Δ/Δ blasts in vivo to the same level as in CebpaΔ/p30Tet2+/+ blasts (Fig. 6c). Intriguingly, two out of three individual CebpaΔ/p30Tet2Δ/Δ leukemic clones (A + B) responded to 5-AZA treatment with a pronounced increase of Gata2 levels and concomitant reduction of myeloid blasts, while one clone (C) appeared partially refractory to 5-AZA treatment, with limited increase of Gata2 and no reduction of leukemic burden (Fig. 6b, c). Importantly, a longer intermittent 5-AZA treatment prolonged the survival of mice transplanted with CebpaΔ/p30Tet2Δ/Δ blasts from one of the responding clones (A) (median survival +22%; Fig. 6d, e), while it did not affect disease latency of mice transplanted with CebpaΔ/p30Tet2+/+ blasts (A).
a Experimental setup for evaluating the effect of short-term 5-azacytidine (5-AZA) treatment in vivo. Recipient mice were sub-lethally irradiated and transplanted with leukemic BM from moribund secondary recipient mice. Three individual CebpaΔ/p30Tet2Δ/Δ clones (A–C) and two CebpaΔ/p30Tet2+/+ clones (A–B) were used, respectively. The illustration was created with BioRender.com. b Expansion of myeloid (Mac1+) donor-derived cells in bone marrow (BM) assessed by flow cytometry, and c Gata2 mRNA expression in sorted leukemic blasts by qPCR assessed 24 hours after the last of three injections of 5-AZA or vehicle (samples from 3 mice per clone and 6 and 9 mice per group, for CebpaΔ/p30Tet2+/+ and CebpaΔ/p30Tet2Δ/Δ, respectively). Dot plots showing individual mice for separate clones and bar graphs shows mean ± SEM for each group. Data were analyzed by Kruskal–Wallis test followed by Dunn’s correction for multiple comparisons. d Experimental setup for evaluating the effect of 5-AZA treatment on AML progression in vivo. The illustration was created with BioRender.com. e Survival of sub-lethally irradiated tertiary recipient mice after transplantation of leukemic BM from moribund secondary recipient mice (clone A from both genotypes) in response to intermittent 5-AZA treatment (5-AZA treated groups 8 mice and vehicle-treated groups 4 mice). The data were analyzed by Mantel–Cox Log-rank test. Source data are provided as a Source Data file.
In summary, we show that the demethylating agent 5-AZA can restore Gata2 expression levels in TET2-deficient CebpaDM AML to that of TET2-proficient CebpaDM AML, and concomitantly reduce leukemic burden and prolong survival of mice transplanted with TET2-deficient CebpaDM leukemic blasts.
Discussion
Mutational cooperativity is a fundamental driver of cancer development, progression, and aggressiveness. For CEBPADM AML, co-occurring lesions have been found in genes such as GATA2, TET2, WT1, FLT3, and CSFR3. While the mechanistic basis for the cooperation between CEBPA and GATA2/CSFR3 mutations has been investigated using mouse models14,15, we have very little insight into why other lesions, such as those in TET2, are overrepresented in CEBPADM AML. Here, we show that TET2 loss-of-function in CEBPADM AML leads to an aggressive disease phenotype by rebalancing the increased and suboptimal levels of GATA2 that are induced by hypermorphic CEBPANT mutations driving CEBPA-p30 isoform expression (see model in Fig. 7a). Specifically, loss of TET2 binding to the hematopoietic-specific G2DHE enhancer results in increased DNA methylation in the promoter region of the hematopoietic-specific Gata2 isoform (Gata2 V2). This proleukemic effect of TET2 loss can be reversed by the demethylating agent 5-AZA, suggesting that this could be a potential treatment option in CEBPADMTET2MUT patients. Altogether, our work proposes that CEBPA-mutant AMLs acquire additional lesions in genes such as GATA2 and TET2 to reestablish balanced GATA2 levels that permit leukemia development and progression.
a Model of Gata2 differential expression as a consequence of (I) elevated CEBPA p30 due to the hypermorphic effect of the CEBPANT, (II) TET2 deficiency and, (III) CEBPA deficiency. b Schematic illustration of two strategies for CEBPADM AML to rebalance GATA2 levels by (I) loss-of-function mutations in TET2 and (II) loss-of-function mutations in one GATA2 allele. The illustrations were created with BioRender.com.
Our work highlights the central importance of GATA2 regulation in CEBPA-mutant AML. Specifically, we show that GATA2 is a conserved target gene of CEBPA and TET2. Furthermore, the elevated levels of the CEBPA p30 variant likely mediate GATA2 upregulation in CEBPA-mutant AML. The increased expression of Gata2 is counteracted by loss of TET2 in vitro and in vivo models of CebpaDM AML as well as in CEBPA-TET2 co-mutated patients. This is accompanied by the gain of Gata2 promoter DNA methylation. These findings are consistent with previous data showing that Gata2 expression is TET2-dependent, as Gata2 was downregulated in various Tet2 knockout settings and that forced expression of Gata2 decreased the competitiveness of both normal and malignant TET2-deficient cells28,43,44,45. Further paralleling our data, TET2 deficiency in the context of Flt3ITD AML has been shown to accelerate leukemia by hypermethylation and consequent silencing of the Gata2 locus43.
Strikingly, we found that while moderate reduction of Gata2 expression increased competitiveness in CebpaDM AML both in vivo and in vitro, leukemia cells remain critically dependent on residual GATA2 function. Indeed, homozygous Gata2 lesions induced a strong inhibitory effect on CebpaDM AML in vitro37, which was also observed in other AML subtypes as well as in normal hematopoietic stem cells36,46,47,48. These findings are corroborated by a substantial body of genetic evidence supporting the importance of GATA2 regulation in CEBPA-mutant AML. First, heterozygous GATA2 lesions frequently co-occur with CEBPADM 4,8,16,17,31,32,33,49,50,51,52. Secondly, GATA2 allele-specific expression is strongly associated with CEBPADM AML and is neither found in AML with reduced CEBPA expression (i.e. t(8;21)) nor in CEBPA-silenced AML53. Thirdly, TET2MUT and GATA2MUT rarely co-occur in CEBPADM AML. Finally, we showed that mutations in GATA2 and TET2 are overrepresented in AML cases with high CEBPA expression. This supports the notion that unfavorable, high GATA2 levels in AML promoted by the CEBPA-TET2 axis are not limited to CEBPADM AML, but also include cases where CEBPA expression is high for other reasons. Further, this model also suggests that a major proleukemic effect of TET2 deficiency is to rebalance GATA2 levels in the context of CEBPADM AML (see Fig. 7b).
GATA2 expression is mainly driven by the conserved G2DHE in normal myeloid progenitors and leukemic blasts by promoting expression from the hematopoietic specific Gata2 V2 promoter39,40,44,54. Our data demonstrate that CEBPA plays a key role in regulating G2DHE activity. Specifically, we showed that the hypermorphic effects of CEBPADM 11, and experimental models thereof, resulted in increased GATA2 expression compared to CEBPAWT, and that CEBPA deficiency resulted in reduced Gata2 levels. Secondly, we observed increased CEBPA binding to the G2DHE in CebpaDM AML compared to normal progenitors and found that deletion or mutagenesis of the CEBPA-bound region of the enhancer resulted in lower expression of Gata2 in CebpaDM cells. In further support of the role of CEBPA, the G2DHE is highly active in CEBPADM AML, with both elevated eRNA expression and levels of H3K27ac53. An equally important role for CEBPA is observed in the context of inv(3) and t(3;3) AML in which inversions, translocations, and rearrangements involving the EVI1 gene at the MECOM locus, lead to hijacking of the G2DHE to promote EVII expression at the expense of GATA2 expression thus resulting in GATA2 haploinsufficiency55,56,57,58. Here, EVI1 expression was found to be downregulated following knockdown of CEBPA in inv(3) AML cells, and mutation of the CEBPA binding site in the hijacked enhancer reduced enhancer activity58. In this context, CEBPAMUT would not be favorable, and these lesions are indeed underrepresented in inv(3) and t(3;3) AML59,60,61.
We hypothesized that CEBPA recruits TET2 and thus mediates DNA demethylation of the Gata2 V2 promoter in a CEBPA- and TET2-dependent manner. Indeed, we observed reduced TET2-binding to the G2DHE upon knockdown of Cebpa in Cebpap30/p30 cells. Furthermore, Gata2 V2 levels were decreased, and Gata2 V2 promoter DNA methylation was increased upon Cebpa depletion in an MLL-AF9 leukemic setting where CEBPA is dispensable for maintenance of the leukemia. The concept of CEBPA as a recruiting factor for TET2 is also supported by previous findings showing that both the p30 and p42 isoforms of CEBPA interact with TET2 via the DNA binding domain of CEBPA62,63. Further, CEBPA binds preferentially to methylated DNA62,64, and has been classified as a binding site-directed DNA demethylation-inducing transcription factor62,65. Interestingly, TET2 binds genomic regions that are enriched for CEBP motifs in myeloid cells, particularly in myeloid enhancers such as the G2DHE26,62. Moreover, knockdown or knockout of Tet2 leads to impaired upregulation of myeloid-specific genes upon Cebpa induction, with corresponding increased promoter methylation66. Also, in TET2MUT or Tet2−/− leukemia an enrichment of CEBP motifs at or near hypermethylated CpGs was observed26,67. Importantly, AML with silenced CEBPA is associated with DNA hypermethylation, a feature that is not present in CEBPADM AML, which may suggest a broader function of CEBPA in the recruitment of TET268. In summary, we conclude that CEBPA plays an important role in the recruitment of TET2 to chromatin at the G2DHE, promoting DNA demethylation at the Gata2 V2 promoter and the induction of Gata2 expression. The extent to which this can be extended to other loci warrants further analysis but is supported by the data mentioned above.
While our findings suggest that GATA2MUT and TET2MUT both converge at rebalancing the increased expression of GATA2 in CEBPADM AML, patients with CEBPADM and GATA2MUT have a more favorable prognosis16,31,32,33,49 than patients harboring the CEBPADM and TET2MUT combination16,17. This suggests that while GATA2 deregulation plays an important role in leukemogenesis in the CEBPAMUT context, TET2 deficiency may likely contribute to malignancy through additional mechanisms that shall remain the subject of future work. Of clinical interest, we find that TET2 deficiency renders CebpaDM AML sensitive to 5-AZA and that TET2-deficient cells lose their proliferative advantage over TET2-proficient cells following 5-AZA treatment. In agreement with TET2-dependent Gata2 expression, ours and previous results show that 5-AZA treatment derepresses Gata2 expression in TET2-deficient cells44. Intriguingly, CEBPACT mutations have recently been reported to sensitize AML to treatment with hypomethylating agents by disrupting the inhibitory interaction with DNMT3A mediated by the wild-type CEBPA bZIP domain69. Taken together, this suggests that demethylating agents could be a particularly interesting treatment option in CEBPADMTET2MUT patients.
Finally, we note that although our mechanistic data have been acquired in experimental models of complete TET2 loss, data from AML patients indicates that TET2 haploinsufficiency is sufficient to rebalance GATA2 levels. We are also aware of the fact that our experimental models mimic AML in which the CEBPA p30 variant constitutes the sole CEBPA entity, which is different from the combination of N- and C-terminal mutations that constitutes the bulk of human CEBPADM AML cases. However, since our main findings from the murine Cebpap30/p30/CebpaΔ/p30 models are also observed in human CEBPADM AML (including upregulation of CEBPA and GATA2 in leukemic GMPs compared to normal GMPs, as well as rebalancing of GATA2 expression and worsened outcome by the acquisition of TET2 lesions), we believe that our observations indicate that a similar disease-relevant CEBPA-TET2 axis is active in human CEBPADM AML.
In conclusion, our results reveal that GATA2 is a conserved target of the CEBPA-TET2 mutational axis in CEBPADM AML and we propose an intricate mechanism by which elevated CEBPA p30 levels mediate recruitment of TET2 to regulatory regions of the Gata2 gene to promote its expression. We demonstrate that increased GATA2 levels are disadvantageous to CEBPADM leukemic cells and that this can be counteracted by TET2 loss thus providing an explanation for the co-occurrence of CEBPA and TET2 lesions in AML. Finally, increased Gata2 promoter methylation, inflicted by TET2 deficiency, can be restored by demethylating 5-AZA treatment, thereby providing entry points for the development of rational targeted therapies in AML patients with these mutations.
Methods
Patient data
Assessment of mutational status
To evaluate co-occurring mutations in CEBPADM AML cases, data from published studies3,4,5,6,7,8,17 including >40 CEBPADM cases were extracted, and co-occurring mutations were evaluated (Supplemental Table 1). To determine frequencies of target gene mutations between CEBPADM AML cases with TET2MUT compared to TET2 wild-type (TET2WT) AML cases, data from published studies3,4,5,7,8,17,51 with specified mutational status including >40 CEBPADM cases or corresponding cohorts were extracted and co-occurring mutations in TET2, GATA2, WT1, CSF3R, and ASXL1 were evaluated (Supplemental Table 2a–e). To examine how the mutational status of TET2 and GATA2 were affected by CEBPA expression levels in AML, we utilized the publicly available data from the Beat AML cohort (Oregon Health & Science University; OHSU)1, including 382 cases for which mutation and mRNA expression data were available. The cases were stratified based on CEBPA mRNA expression levels (z-score ±1.0 relative to all samples; CEBPAHIGH n = 45 and CEBPALOW n = 61) and frequencies of CEBPA, TET2, and GATA2 mutations were determined.
Survival analysis
The clinical data set comprises 298 patients with CEBPA mutations (MLL Münchner Leukämielabor GmbH), of which 152 harbored biallelic CEBPA mutations. Out of these 119 had specified TET2 mutational status and were included in the analyses (CEBPADMTET2WT n = 84, CEBPADMTET2MUT n = 35; Supplemental Table 3). All patients gave written informed consent for the use of data for scientific evaluations. The study was approved by the Internal Review Board and by the Bavarian Ethics Committee, the Bavarian State Medical Association (Bayerische Landesärztekammer) with the number 05117. The study adhered to the tenets of the Declaration of Helsinki.
Gene expression
The Beat AML dataset used in this study is available at http://vizome.org/aml and comprises 25 patients with CEBPA mutations (CEBPANT and/or CEBPACT) for which mutation and mRNA expression data is available. For the gene expression analysis, we excluded patients, which had co-occurring mutation(s) in WT1 or IDH1/2 since these have been shown to interfere with TET2 function70,71,72,73 as well as two patients with low CEBPA variant allele frequency (VAF). Gene expression analysis was conducted on data from 16 CEBPA-mutant patients of which 5 have a co-occurring mutation in TET2 (TET2MUT) (Supplemental Table 4). Differential expression analysis was performed with DESeq274 (v. 1.26.0, RRID:SCR_015687) and default parameters. To assess gene expression changes in CEBPAWT patients with TET2MUT vs TET2WT, we included patients with normal karyotype AML from the Beat AML dataset and excluded patients with mutation(s) in WT1 or IDH1/2 (CEBPAWTTET2MUT n = 34 and CEBPAWTTET2WT n = 167). To analyze CEBPA and GATA2 expression levels in CEBPAWT vs. CEBPANT AML mRNA expression of the two genes together with mutational and karyotype status data was retrieved from the Beat AML study1 via cBioPortal75,76 (RRID:SCR_014555). We evaluated patients for whom data was available for genomic profiling including mRNA expression, mutations, and karyotype. We included patients with normal karyotype AML and excluded patients with mutation(s) in WT1 or IDH1/2 (CEBPAWT n = 52 and CEBPANT n = 15).
In vitro experiments
Competitive CRISPR-targeting
For generation of Tet2 or Gata2 mutated clones, Cebpap30/p30 (♂) cells37 were electroporated with ribonucleoparticles containing recombinant Cas9 nuclease from Streptococcus pyogenes (Sp) (#1081058, IDT), tracrRNA (#1075927, IDT) and crRNAs (Alt-R® CRISPR-Cas9 crRNA, IDT) targeting Tet2 and Gata2, respectively. crRNAs were designed using the CHOPCHOP77 web tool (chopchop.cbu.uib.no, RRID:SCR_015723) (Supplemental Table 5). crRNA and tracrRNA molecules were complexed at room temperature and assembled with recombinant SpCas9 according to the manufacturer’s protocols (IDT). Pools of Tet2- or Gata2-targeted cells were screened at regular intervals to monitor the outgrowth of subpopulations. The genomic regions that were targeted with CRISPR/Cas9 technology were PCR-amplified, Sanger sequenced, and analyzed with the online tool Tracking of Indels by DEcomposition (TIDE)78 for insertions or deletions (indels) in the targeted region. Primers for PCR are provided in Supplemental Table 6.
Gata2 enhancer CRISPR-targeting
sgRNA sequences targeting the Gata2 distal hematopoietic enhancer (G2DHE) were obtained from the UCSC Genome Browser79 (genome.ucsc.edu, RRID:SCR_005780) and targets with a high predicted cleavage (Doench/Fusi 2016 Efficiency > 55)80 selected (Supplemental table 5). SpCas9-expressing Cebpap30/p30Tet2MUT cells were isolated after lentiviral expression of lenti-Cas9-Blast (#52962 Addgene). Cebpap30/p30 and Cebpap30/p30Tet2MUT cells were co-transduced with pLenti-hU6-sgG2DHE_A/B-IT-PGK-iRFP and LentiGuide-sgG2DHE_1–6-Puro-IRES-GFP. GFP+iRFP670+ cells were sorted via fluorescence-activated cell sorting (FACS) and frozen for subsequent analysis.
Cebpa shRNA-knockdown
Cepbap30/p30 rtTA3 cells expressing Dox-inducible shRNAs against Renilla luciferase (shRen, control) or Cebpa (shCebpa) were used as previously described12. Cebpa knockdown was induced by the addition of 4-hydroxytamoxifen (4-OHT; 1 μM; #H7904 Sigma-Aldrich) to the cell culture medium to activate shRNA expression (mean Cebpa knockdown efficiency > 90% compared to shRen control) and cells were collected for further analysis after 48 h.
In vivo experiments
Experiments were carried out according to protocols approved by the Danish Animal Ethical Committee. Mice were bred and housed locally at the Department of Experimental Medicine at the University of Copenhagen. The mice were housed in a temperature- and humidity-controlled room with a 06:00–18:00 h light cycle and fed a standard chow diet and tap water ad libitum. We used Tet2fl/fl 81, Cebpap30/+ 13, Cebpa+/fl 82 and Mx1-Cre+ 83 lines to generate inducible Tet2-deficient and Cebpa-mutant compound lines. The following genotypes were used for experiments: Cebpafl/p30Tet2+/+, Cebpa+/flTet2fl/fl, Cebpafl/p30Tet2fl/fl, Cebpafl/p30Tet2+/+Mx1-Cre+, Cebpa+/flTet2fl/flMx1-Cre+, and Cebpafl/p30Tet2fl/flMx1-Cre+. Cebpap30/p30 embryos were generated as previously described13. We used iMLL-AF9+ 84, Cebpa+/fl 82 and R26-CreER+ 85 lines to generate an iMLL-AF9+Cebpafl/flR26-CreER+ compound line. Primers for genotyping in are provided in Supplemental Table 6.
During the leukemia initiation and propagation experiments described below, the animals were monitored daily and euthanized when they showed sign(s) of sickness e.g., inactivity, hunched posture, white paws, and/or matted or puffed-up fur as well as pain assessed based on the NC3R’s mouse grimace scale86 or reduced bodyweight (maximal allowed reduction = 15%). The experiments were terminated after 12 months.
Leukemia initiation (CebpaΔ/p30 model)
C57BL/6 J.SJL congenic recipients (female, 10–12 weeks old) were lethally irradiated (900 cGy) 12–24 h prior to intravenous injection with 1 × 106 bone marrow (BM) cells from individual donor mice. The mice were given Ciprofloxacin (100 mg/l in acidified water; #17850 Sigma-Aldrich) in the drinking water to prevent infections 3 weeks post-irradiation. Recipient mice were allowed to recover for 6 weeks post-transplantation before Cre-LoxP recombination was induced by two intraperitoneal injections of Poly(I)-Poly(C) (300 μg in 200 μl PBS; #27-4732-01 GE Healthcare) with 48 h rest in-between. The day of the first injection was set as time-point zero for the survival study and mice were monitored for leukemia development and euthanized when moribund. To follow leukemia initiation in the recipients, a subgroup of mice was subjected to blood and BM sampling at 12, 24, and 36-week time-points. BM from moribund mice was collected and frozen (10% DMSO in FBS; #D8418 Sigma-Aldrich, #HYCLSV30160.03 Hyclone) for subsequent FACS and analysis.
Leukemia initiation (Cebpap30/p30 model)
C57BL/6 J.SJL congenic recipients (female, 10 weeks old) were lethally irradiated (900 cGy) 12–24 h prior to intravenous injection with 0.5-1 × 106 fetal liver cells from E15.5 Cebpap30/p30 embryos. The mice were given Ciprofloxacin (100 mg/l in acidified water) in the drinking water to prevent infections 3 weeks post-irradiation. Latency to leukemic initiation was 8-11 months.
Leukemia propagation
C57BL6/6 J.SJL recipients (female, 10–12 weeks old) were lethally irradiated (900 cGy) 12–24 h prior to being intravenously injected with 2 × 105 thawed live BM cells from moribund donor mice together with 4 × 105 freshly isolated BM cells from C57BL6/6 J.SJL mice. The day of the injection was set as time-point zero for the survival study and mice were monitored and euthanized when moribund. The mice were given Ciprofloxacin in the drinking water to prevent infections 3 weeks post-irradiation.
Competitive shRNA-knockdown
C57BL/6 J.SJL recipients (female, 10–12 weeks old) were sub-lethally irradiated (500 cGy) 12–24 h prior to being intravenously injected with a 1:1 mix of Cebpap30/p30 cells13 transduced with shRNA targeting Gata2 (detailed in ShRNA knockdown below) or with control-shRNA87. The ratio of Gata2- or control-shRNA-GFP+ to control-shRNA-YFP+ cells was analyzed by flow cytometry four weeks later.
5-azacytidine treatment
C57BL/6 J.SJL recipients (female, 10–12 weeks old) were sub-lethally irradiated (500 cGy) 12–24 h prior to being intravenously injected with 1 × 105 thawed live BM cells from moribund secondary recipient mice. The mice were given Ciprofloxacin in the drinking water to prevent infections 3 weeks post-irradiation. The mice received intraperitoneal injections with the demethylating agent 5-azacytidine (2.5 mg/kg/day in saline; #A2385 Sigma-Aldrich) at days 6–10 and 20–24 post-transplantation. The time of the BM cell injection was set as time-point zero for the survival study and mice were monitored and euthanized when moribund. To evaluate the effects of short-term 5-azacytidine treatment, recipient mice were treated at days 13–15 and euthanized 24 h after the last injection. BM was collected for FACS, and sorted cells were frozen for subsequent analysis.
Ex vivo cell culture
Establishment of ex vivo CebpaΔ/p30Tet2+/+ and CebpaΔ/p30Tet2Δ/Δ lines
Thawed cryo-preserved cells from primary AML were cultured in Lonza X-VivoTM 15 cell medium (#BE02-060Q Thermo Fisher Scientific) supplemented with Bovine Serum Albumin in Iscove’s MDM (10%; #09300 StemcellTM Technologies), Penicillin-Streptomycin (1%; #15140122 Gibco), β-mercaptoethanol (0.1 mM; #31350010 Gibco), L-glutamine (2 mM; #25030149 Gibco), and cytokines h-IL-6 (50 ng/ml; #130-093-032 Miltenyi Biotec), m-SCF (50 ng/µl; #250-03 Peprotech), m-IL-3 (10 ng/ml; #213-13 Peprotech), and m-GM-CSF (10 ng/ml; #315-03 Peprotech). Two clones of each genotype (CebpaΔ/p30Tet2+/+ and CebpaΔ/p30Tet2Δ/Δ) continued to expand beyond 40 days and withstood freeze-thawing, and these clones have been used for further experiments.
Vitamin C treatment
Cells were seeded at a density of 2 × 105 cells/ml and the cell culture medium was supplemented with vitamin C (100 μg/ml; L-ascorbic acid, #A8960 Sigma Aldrich). Live cells were counted using Solution 13 (AO-DAPI; #910-3013 Chemometec) on a NucleoCounter® NC-3000TM and reseeded at 2 × 105 cells/ml every third day. The experiments were run with a total of two biological replicates per genotype (performed on separate days) where each experiment assayed one leukemic line per genotype. Each condition (Vitamin C and vehicle) was performed in technical triplicates for each of the two biological replicates per genotype.
5-azacytidine treatment
Cells were seeded at a density of 2 × 105 cells/ml and medium supplemented with 5-azacytidine (5-AZA; 1 μg/ml; #A2385 Sigma-Aldrich). Live cells were counted using Solution 13 on a NucleoCounter® NC-3000TM and reseeded at 2 × 105 cells/ml days three and six. 24 hours later, up to 1 × 105 cells were isolated and resuspended in RA1 buffer (NucleoSpin RNA XS, #740902 Macherey-Nagel). The experiments were run with a total of two biological replicates per genotype (performed on separate days) where each experiment assayed one leukemic line per genotype. Each condition (5-AZA and vehicle) was performed in technical triplicates for each of the two biological replicates per genotype.
Establishment of ex vivo iMLL-AF9+Cebpafl/flR26-CreER+ lines
Sorted GMPs from iMLL-AF9+Cebpafl/flR26CreER+ mice, were cultured in MethoCult (M3434; #03434, Stemcell technologies) supplemented with doxycycline (1 μg/ml; #D9891 Sigma-Aldrich) for three replatings to induce expression of the MLL-fusion protein.
Cebpa knockout
Leukemic iMLL-AF9+Cebpafl/flR26CreER+ cells were cultured in RPMI 1640 medium (#21875034, Gibco) supplemented with FBS (10%), Penicillin-Streptomycin (1%), doxycycline (1 μg/ml), and cytokines m-IL-3 (6 ng/ml), m-SCF (50 ng/ml), and h-IL-6 (10 ng/ml). After two days, 4-hydroxytamoxifen (4-OHT; 1 μM; #H7904 Sigma-Aldrich) or vehicle was added to the cell culture medium to activate Cre-LoxP recombination, resulting in a reduction of Cebpa mRNA to 1.7 ± 0.3% vs. 100 ± 12.3% in vehicle control. Three days later cells were isolated and either frozen or resuspended in RA1 buffer (NucleoSpin RNA XS, # 740902 Macherey-Nagel). The experiments were run with a total of two biological replicates (performed on separate days). Each condition (4-OHT and vehicle) was performed in 2-3 technical triplicates for each of the two biological replicates.
ShRNA knockdown
Cloning of shRNA into pMLS vector
Murine shRNAs targeting Gata2 (shGata2) were cloned into MSCV-LTRmir30-SV40-GFP vector. Targeting sequences were identified from the Mission® shRNA library (Supplemental Table 7) and the sense and anti-sense sequences were incorporated with a miR-30-loop to generate a 97-mer target sequence. Oligonucleotides were amplified by PCR using miR30 common primers (Supplemental Table 6), which include restriction sites for XhoI and EcoRI. The resulting 138-mer PCR amplicons and the vector were digested with XhoI and EcoRI and products were ligated using T4 DNA Ligase (#15224025 Invitrogen). Bacterial transformation was performed to amplify individual ligation products, and correct inserts were verified by Sanger Sequencing. These, together with vectors containing a control non-targeting sequence (MSCV-LTRmir30-SV40-GFP and MSCV-LTRmir30-SV40-YFP), were used in subsequent transfection/transduction experiments, as previously described87,88.
Transduction of Cebpap30/p30 cells
Retroviral transduction was done as previously described87. Briefly, retroviral supernatants were generated by transfection of Phoenix-Eco cells (RRID:CVCL_H717). For transduction, retroviral supernatant was added onto retronectin-coated (1:25; #T100B TaKaRa) non-tissue culture treated plates and centrifuged at 2000 ×g for 60 min at 4 °C. After aspiration of the supernatant, Cebpap30/p30 cells were seeded at a density of 0.5–1 × 105 cells/cm2. The transduction was repeated the following day, and the cells were cultured for 24 h prior to FACS sorting of transduced (GFP+/YFP+) cells on a BD FACSAriaTM III (BD Bioscience). The efficiency of shRNA-mediated gene expression knockdown was assessed with qPCR and cells were used for transplantation and assessment of their competitiveness in vivo.
Immuno-staining
Flow cytometry
To analyze the composition of either freshly isolated or thawed cryopreserved BM and blood, cells were stained with fluorescently labelled antibodies. For blood analysis, 50 µl blood was collected from the facial vein and erythrocytes were lysed with lysing buffer (BD Pharm LyseTM, #555899 BD Bioscience). For BM analysis, cells were collected by crushing tibia, femur, and ilium and filtered through a 50 μm filcon cup (#340630 BD Bioscience). Blood or BM cells were washed in PBS with 3% FBS and stained with fluorescently labelled antibodies for 30 min at 4 °C (Supplemental Table 8). For cryopreserved cells, the cells were counterstained with DAPI (1:10000; #D3571 Invitrogen) to separate out dead cells. Fluorochrome-minus-one was used as controls. Flow cytometry data was obtained using a BD FACSAriaTM III or a BD LSR IITM (BD Bioscience) and analyzed using FlowJo software (v9, RRID:SCR_008520).
For downstream transcriptional and epigenetic analyses, live donor-derived non-lymphoid and non-erythroid cells (DAPI−CD45.2+CD3−B220−Ter119−) were sorted using a BD FACSAriaTM III, spun down and cell pellets were either snap-frozen or resuspended in RLT buffer (RNeasy Mini Kit, #74104 Qiagen).
For ex vivo cell culture of iMLL-AF9+Cebpafl/flR26-CreER+ cells, c-kit+ BM cells were enriched by magnetic sorting (mouse CD117 MicroBeads; #130-091-224, Miltenyi Biotec), and granulocyte/monocyte progenitors (GMPs; Lin−C-kit+Sca1−CD41−FcgRII+) were sorted using a BD FACSAriaTM III.
Immunohistochemistry
To evaluate the proliferative status of leukemia cells, cells from BM of moribund mice were spun on glass slides, air-dried, and fixed with methanol (#VWRC20846.292 VWR). After blocking of endogenous peroxidase activity with hydrogen peroxide (1%), slides were stained overnight at 4 °C with anti-Ki67 antibody (1:50; Clone SP6, RRID:AB_302459, #ab16667 Abcam) in antibody diluent (S3022 Dako). To visualize the primary antibody, EnVision HRP Rabbit (K4003 Dako) together with Vina GreenTM Chromogen Kit (BRR807 Biocare Medical) was utilized according to manufacturer’s instructions. The cells were counterstained with Mayer Hematoxylin (#51275 Sigma-Aldrich), dehydrated and coverslips mounted with Entellan (#107960 Sigma-Aldrich). Images were captured using a Leica microscope at 20X magnification and Ki67+ cells were quantified out of one hundred cells.
Western blotting
Western blotting for TET2 was performed according to standard laboratory protocols, using the following antibodies: anti-TET2 (1:100, Clone C-7, RRID:AB_2924805, #sc-398535 Santa Cruz) and anti-HSC70 (1:10000, Clone B-6, RRID:AB_627761, #sc-7298 Santa Cruz).
Quantitative PCR
RNA from sorted blasts or ex vivo-cultured cells was isolated using NucleoSpin RNA XS kit (#740902 Macherey-Nagel) or RNeasy Mini Kit (#74104 Qiagen) according to the manufacturers’ instructions and converted to cDNA using ProtoScript First Strand cDNA Synthesis Kit (#E6300 New England BioLabs). Quantitative PCR (qPCR) to assess knockdown efficiency was run using TaqMan Fast Advanced Master Mix (#4444556 Applied Biosystems) and TaqMan assay for Gata2 (Mm00492301_m1 FAM-MGB), in duplex with housekeeping gene 18 S (Hs99999901_s1 VIC-MGB-PL). TaqMan assay for Ki67 (Mm01278617_m1 FAM-MGB) was used to assess the expression of the proliferation marker. qPCR to evaluate mRNA levels of total Gata2, variant 1 (V1) and variant 2 (V2), respectively, was run in duplex using LightCycler 480 SYBR Green I Master (#04887352001 Roche) with primers for Gata2 and housekeeping gene Actb and Gapdh41 (Supplemental Table 6). Gene expression was calculated with the 2−ΔΔct method.
RNA from Cebpap30/p30 cell lines was isolated using RNeasy Plus Mini Kit (#74134 Qiagen) according to the manufacturer’s instructions and converted to cDNA with RevertAid First Strand cDNA Synthesis Kit (#K1622 Thermo Scientific). qPCR was run using SsoAdvanced Univ SYBR Grn Suprmix (#1725271, Bio-Rad Laboratories Ges.m.b.H.) and primers for Gata2 and Gapdh (Supplemental Table 6).
Bisulfite PCR
DNA was isolated using DNeasy Blood and tissue kit (#69504 Qiagen) and the DNA was bisulfite converted using EZ-DNA Methylation Gold Kit (#D5005 Zymo Research), both according to the manufacturer’s instructions. PCR was run using Pfu Turbo Cx Hotstart DNA polymerase (#600410 Agilent) with primers targeting a part of the CpG island in the Gata2 V2 promoter region (Supplemental table 6). After verification of their correct size, PCR products were cloned using Zero Blunt Topo PCR Cloning kit (#450245 Invitrogen), and single colonies were picked and amplified. Plasmid DNA was isolated using NucleoSpin Plasmid EasyPure (#740727.250 Macherey-Nagel), the correct insert size was verified after cleavage with restriction enzyme EcoRI (#R0101 New England Biolabs) and sent for Sanger sequencing using the M13 primer provided with the cloning kit.
Chromatin immunoprecipitation (ChIP)-qPCR
ChIP for CEBPA was performed as previously described37 using an anti-CEBPA antibody (1:60, C-18, RRID:AB_2078046, #sc-9314, Santa Cruz Biotechnology). ChIP for TET2 was performed using an anti-TET2 antibody (1:50; clone D6C7K, RRID:AB_2799102, #36449, Cell Signaling Technology), as previously described37, including a 30-minute incubation with 2 mM disuccunumidyl glutarate (DSG; #20593 Thermo Scientific) before the 1% formaldehyde crosslinking step. The sequences used for qPCR are listed in Supplemental table 6.
High-throughput sequencing and bioinformatic analyses
RNA-sequencing (RNA-seq) of cell line models
RNA was isolated from 1 × 106 cells using RNeasy Plus Mini Kit (#74134 Qiagen) according to the manufacturer’s instructions and quality was assessed on a Bioanalyzer 2100 G2939A (Agilent). 1 µg of RNA was used to generate sequencing libraries using QuantSeq 3′ mRNA-Seq Library Prep Kit (FWD) for Illumina, 96 preps (#015.96, Lexogen) and the PCR Add-on Kit for Illumina, 96 rxn (#020.96, Lexogen). The libraries were quantified on a Bioanalyzer 2100 G2939 (Agilent) and pooled in equimolar amounts. Multiplexed libraries were sequenced on a HiSeq4A (Illumina).
RNA-seq of leukemic cells from in vivo models
RNA was isolated from 5 × 105 sorted cells using RNeasy Mini Kit (#74104 Qiagen) according to the manufacturer’s instructions and quality was assessed by RNA 6000 Pico Kit (#5067-1513 Agilent) on a Bioanalyzer 2100 (Agilent). 200 ng RNA was used to generate sequencing libraries using TruSeq RNA Library Prep Kit v2 (#RS-122-2001 Illumina). The libraries were quantified using Qubit dsDNA BR Assay Kit (#32853 Thermo Fisher Scientific) and DNA 1000 Kit (#5067-1504 Agilent) and pooled in equimolar amounts. Multiplexed libraries were sequenced on a NextSeq 500 (Illumina) using NextSeq 500 High Output v2 Kit (75 cycles; #FC-404-2005 Illumina).
Bioinformatics analyses of RNA-seq data
RNA-seq analysis for in vitro Cebpap30/p30 cells was performed as previously described12,37. Quality check was done with FastQC89 (v. 0.11.4, RRID:SCR_014583) and preprocessing with PRINSEQ-lite90 (v. 0.20.4; RRID:SCR_005454), using parameters: -min_len 30 -min_qual_mean 30 -ns_max_n 5 -trim_tail_right 8 -trim_tail_left 8 -trim_qual_right 30 -trim_qual_left 30 -trim_qual_window 5. The remaining reads were aligned against the mouse reference genome (mm10) with BWA91 (v. 0.7.15; RRID:SCR_010910). RNA-seq analysis for in vivo CebpaΔ/p30 cells was performed as follows. RNA-seq reads were processed with the bcbio RNA-seq pipeline92 (https://github.com/bcbio/bcbio-nextgen, RRID:SCR_004316) and the bcbioRNASeq R package (https://github.com/hbc/bcbioRNASeq). In brief, transcript abundance estimates were obtained using Salmon93 (v. 0.12.0, RRID:SCR_017036) against reference transcriptome GRCm38/mm10 ENSEMBL release 94, summarized to gene level and imported into R using tximport94 (v. 1.10.1, RRID:SCR_016752) (using setting countsFromAbundance = “lengthScaledTPM”). Differential gene expression analysis between the CebpaΔ/p30Tet2+/+ and CebpaΔ/p30Tet2Δ/Δ genotype was performed using DESeq2 with standard parameters74 (v. 1.22.2, RRID:SCR_015687) excluding lowly expressed genes ( < 10 sum normalized counts across all samples) and running with alpha = 0.05.
Gene expression levels between primary CebpaΔ/p30Tet2+/+ and established Cebpap30/p30 leukemias were compared using edgeR (v. 3.32.1, RRID:SCR_012802).
Gene set enrichment analysis (GSEA)
GSEA was performed using the GSEA software95,96 (v. 4.1.0, RRID:SCR_003199) and the Molecular Signatures Database (RRID:SCR_016863).
Assay for transposase-accessible chromatin-sequencing (ATAC-seq)
ATAC-seq was performed as previously described12.
Bioinformatics analyses of ATAC-seq data
Analysis of ATAC-seq was performed as previously described12. HOMER97 (v. 4.11, RRID:SCR_010881) was used to identify motifs enriched in the ATAC peaks.
Bisulfite whole genome sequencing (WGBS)
DNA was isolated from 1 × 106 sorted cells using DNeasy Blood and tissue kit (#69504 Qiagen) according to the manufacturer’s instructions. Bisulfite conversion of DNA was done according to manufacturers’ instructions using EZ-DNA Methylation Gold Kit (#D5005 Zymo Research). Quality of bisulfite treated DNA was assessed by RNA 6000 Pico Kit (#5067-1513 Agilent) on a Bioanalyzer 2100. Libraries of bisulfite-converted DNA were prepared using Pico Methyl-Seq Library Prep Kit (#D5455 Zymo Research) according to manufacturer’s instructions and the final concentration and quality of the libraries was assessed using Qubit dsDNA HS Assay Kit (#Q32854 Thermo Fisher Scientific) and High Sensitivity DNA Analysis Kit (#5067-4626 Agilent) on a Bioanalyzer. Duplexed libraries were sequenced on a NextSeq 500 (Illumina) using NextSeq 500 High Output v2 Kit (75 cycles).
Bioinformatics analyses of WGBS data
Reads were trimmed and filtered using Trim Galore98 (v. 0.4.3, RRID:SCR_011847) with default parameters, and quality was assessed before and after using FastQC89 (v. 0.11.7). Trimmed reads were aligned to the mouse genome assembly GRCm38 (mm10) using Bismark99 (v. 0.19.1, RRID:SCR_005604) with option -non_directional (other parameter left at default values; this used Bowtie 2100 (v. 2.2.8 RRID:SCR_016368) with -q --score-min L,0,−0.2 --ignore-quals). After deduplication of alignments (using deduplicate_bismark), the methylation information for individual cytosines was extracted using bismark_methylation_extractor (--cytosine_report --comprehensive --gzip). To quantify DNA methylation of gene bodies and promoters (1000 bp up-and downstream of transcription start sites), we used the weighted methylation level (i.e., summarizing over all CpG positions in the given region, the number of reads supporting methylated cytosine divided by the number of all reads covering these positions). Plots of average methylation levels across extended gene bodies were generated using deepTools101 (v.3.1.3, RRID:SCR_016366) computeMatrix (scale-regions -m 4000 -a 1000 -b 1000 --unscaled5prime 1000 --unscaled3prime 1000) and plotProfile, for which Bismark-generated bedGraph files were converted to BigWig format (using UCSC’s bedGraphToBigWig102 (v. 4)).
Bioinformatic analyses of chromatin immunoprecipitation-sequencing (ChIP-seq)
ChIP-seq data from in vitro Cebpap30/p30 cells was processed as described12. ChIP-seq data from in vivo Cebpap30/p30 cells was processed as follows; raw reads derived from CEBPA (Cebpa+/+ and Cebpap30/p30) ChIP-seq experiments were mapped to mouse (mm10) genome assembly using Bowtie 2100 (v. 2.3.4.3). We used uniquely mapped and PCR duplicates (exact copies) collapsed as one read and extended to their fragment length by determining the read extension size using MACS2103 (v. 2.1.0.20151222; predicted parameter, RRID:SCR_013291). Raw read counts were normalized to TPM using deepTools101 (v. 3.3.1; bamCoverage). Raw read counts (CEBPA binding levels) mapping to Gata2 promoter and enhancer regions were computed using bedtools104 (v. 2.30.0; multicov, RRID:SCR_006646), and the differences in CEBPA binding between Cebpa+/+ and Cebpap30/p30 conditions were computed using DESeq274 (v. 1.30.1). Sequencing reads derived from TET2 ChIP-seq experiment were preprocessed with PRINSEQ-lite90 (v. 0.20.4; RRID:SCR_005454) and the remaining reads were mapped to the mouse reference genome sequence (mm10) using BWA91 (v. 0.7.17-r1188, RRID:SCR_010910). The resulting alignments were processed with samtools105 (v. 1.13; RRID:SCR_002105) and peak calling was done with MACS2103 (v. 2.1.0.20140616; RRID:SCR_013291). Aligned read counts were normalized to RPKM using the bamCoverage function from deeptools101 (v. 3.5.1; RRID:SCR_016366).
Statistics
Data were analyzed for significance using parametric tests, with prior log-transformation if necessary to achieve normal distribution. Normality was evaluated by Shapiro–Wilk test. Two-group analyses were done using an unpaired two-tailed t-test. Multiple-group analyses were done with one-way-ANOVA followed by multiple comparisons correction using Dunnett when comparing to a reference group, or two-way-ANOVA followed by multiple comparisons correction using Šídák test when comparing two independent factors across four groups. Data sets that did not pass normality tests were analyzed by Kruskal–Wallis test followed by multiple comparisons correction using Dunn’s test. Survival curves were analyzed using Mantel–Cox Log-rank test. To compare distributions Wilson/Brown binominal test was used. To compare a median against a hypothetical median Wilcoxon signed-rank test was used. p-values < 0.05 were considered statistically significant. Data was analyzed using GraphPad Prism (v. 9, RRID:SCR_002798). Data is shown as mean ± SEM unless otherwise stated.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The data generated in this study is publicly available in Gene Expression Omnibus (GEO) under accession numbers GSE214224 (RNA-seq, ATAC-seq, and TET2 ChIP-seq in vitro) and GSE213864 (RNA-seq and WGBS in vivo), and within the article and its supplementary files. The following other publicly available data was used in this study: CEBPA and H3K27Ac ChIP-seq from myeloid progenitor cell model for p30-driven AML12 is available under GSE158727. CEBPA ChIP from mouse Cebpa+/+ or Cebpap30/p30 GMPs11 is available under GSE118963. RNA-seq data from Cebpap30/p30 AML11,34 are available under GSE118963 and GSE141477. Patient data analyzed in this study were from the Beat AML study (accessed through cBioPortal75,76 [https://www.cbioportal.org/] or Vizome1 [http://www.vizome.org/]) or from published cohort studies (Supplemental tables 1, 2a–e). Source data are provided with this paper.
References
Tyner, J. W. et al. Functional genomic landscape of acute myeloid leukaemia. Nature 562, 526–531 (2018).
Ohlsson, E., Schuster, M. B., Hasemann, M. & Porse, B. T. The multifaceted functions of C/EBPalpha in normal and malignant haematopoiesis. Leukemia 30, 767–775 (2016).
Fasan, A. et al. The role of different genetic subtypes of CEBPA mutated AML. Leukemia 28, 794–803 (2014).
Papaemmanuil, E. et al. Genomic classification and prognosis in acute myeloid leukemia. N. Engl. J. Med. 374, 2209–2221 (2016).
Ahn, J. S. et al. Normal karyotype acute myeloid leukemia patients with CEBPA double mutation have a favorable prognosis but no survival benefit from allogeneic stem cell transplant. Ann. Hematol. 95, 301–310 (2016).
Su, L. et al. Mutational spectrum of acute myeloid leukemia patients with double CEBPA mutations based on next-generation sequencing and its prognostic significance. Oncotarget 9, 24970–24979 (2018).
Zhang, Y. et al. Companion gene mutations and their clinical significance in AML with double mutant CEBPA. Cancer Gene Ther. https://doi.org/10.1038/s41417-019-0133-7 (2019).
Taube, F. et al. CEBPA mutations in 4708 patients with acute myeloid leukemia: differential impact of bZIP and TAD mutations on outcome. Blood 139, 87–103 (2022).
Wilhelmson, A. S. & Porse, B. T. CCAAT enhancer binding protein alpha (CEBPA) biallelic acute myeloid leukaemia: cooperating lesions, molecular mechanisms and clinical relevance. Br. J. Haematol. 190, 495–507 (2020).
Leroy, H. et al. CEBPA point mutations in hematological malignancies. Leukemia 19, 329–334 (2005).
Jakobsen, J. S. et al. Mutant CEBPA directly drives the expression of the targetable tumor-promoting factor CD73 in AML. Sci. Adv. 5, eaaw4304 (2019).
Heyes, E. et al. Identification of gene targets of mutant C/EBPalpha reveals a critical role for MSI2 in CEBPA-mutated AML. Leukemia 35, 2526–2538 (2021).
Kirstetter, P. et al. Modeling of C/EBPalpha mutant acute myeloid leukemia reveals a common expression signature of committed myeloid leukemia-initiating cells. Cancer Cell 13, 299–310 (2008).
Di Genua, C. et al. C/EBPalpha and GATA-2 mutations induce bilineage acute erythroid leukemia through transformation of a neomorphic neutrophil-erythroid progenitor. Cancer Cell 37, 690–704.e698 (2020).
Braun, T. P. et al. Myeloid lineage enhancers drive oncogene synergy in CEBPA/CSF3R mutant acute myeloid leukemia. Nat. Commun. 10, 5455 (2019).
Grossmann, V. et al. CEBPA double-mutated acute myeloid leukaemia harbours concomitant molecular mutations in 76.8% of cases with TET2 and GATA2 alterations impacting prognosis. Br. J. Haematol. 161, 649–658 (2013).
Konstandin, N. P. et al. Genetic heterogeneity of cytogenetically normal AML with mutations of CEBPA. Blood Adv. 2, 2724–2731 (2018).
Kunimoto, H. & Nakajima, H. TET2: A cornerstone in normal and malignant hematopoiesis. Cancer Sci 112, 31–40 (2021).
Man, N. et al. p300 suppresses the transition of myelodysplastic syndromes to acute myeloid leukemia. JCI Insight 6, https://doi.org/10.1172/jci.insight.138478 (2021).
Reckzeh, K. et al. TET2 deficiency cooperates with CBFB-MYH11 to induce acute myeloid leukaemia and represents an early leukaemogenic event. Br. J. Haematol. https://doi.org/10.1111/bjh.18027 (2022).
Morinishi, L., Kochanowski, K., Levine, R. L., Wu, L. F. & Altschuler, S. J. Loss of TET2 affects proliferation and drug sensitivity through altered dynamics of cell-state transitions. Cell Syst 11, 86–94.e85 (2020).
An, J. et al. Acute loss of TET function results in aggressive myeloid cancer in mice. Nat. Commun. 6, 10071 (2015).
Li, R. et al. TET2 loss dysregulates the behavior of bone marrow mesenchymal stromal cells and accelerates tet2(-/-)-driven myeloid malignancy progression. Stem Cell Reports 10, 166–179 (2018).
Meisel, M. et al. Microbial signals drive pre-leukaemic myeloproliferation in a Tet2-deficient host. Nature 557, 580–584 (2018).
Weissmann, S. et al. Landscape of TET2 mutations in acute myeloid leukemia. Leukemia 26, 934–942 (2012).
Rasmussen, K. D. et al. TET2 binding to enhancers facilitates transcription factor recruitment in hematopoietic cells. Genome Res. 29, 564–575 (2019).
Jones, P. A. Functions of DNA methylation: islands, start sites, gene bodies and beyond. Nat. Rev. Genet. 13, 484–492 (2012).
Ito, K. et al. Non-catalytic Roles of Tet2 Are Essential to Regulate Hematopoietic Stem and Progenitor Cell Homeostasis. Cell Rep. 28, 2480–2490.e2484 (2019).
Lim, K. C. et al. Conditional Gata2 inactivation results in HSC loss and lymphatic mispatterning. J. Clin. Invest. 122, 3705–3717 (2012).
Koyunlar, C. & de Pater, E. From Basic Biology to Patient Mutational Spectra of GATA2 Haploinsufficiencies: What Are the Mechanisms, Hurdles, and Prospects of Genome Editing for Treatment. Front. Genome Ed. 2, 602182 (2020).
Greif, P. A. et al. GATA2 zinc finger 1 mutations associated with biallelic CEBPA mutations define a unique genetic entity of acute myeloid leukemia. Blood 120, 395–403 (2012).
Hou, H. A. et al. GATA2 mutations in patients with acute myeloid leukemia-paired samples analyses show that the mutation is unstable during disease evolution. Ann. Hematol. 94, 211–221 (2015).
Tien, F. M. et al. GATA2 zinc finger 1 mutations are associated with distinct clinico-biological features and outcomes different from GATA2 zinc finger 2 mutations in adult acute myeloid leukemia. Blood Cancer J. 8, 87 (2018).
Trempenau, M. L. et al. The histone demethylase KDM5C functions as a tumor suppressor in AML by repression of bivalently marked immature genes. Leukemia 37, 593–605 (2023).
Li, H. S. et al. Loss of c-Kit and bone marrow failure upon conditional removal of the GATA-2 C-terminal zinc finger domain in adult mice. Eur. J. Haematol. 97, 261–270 (2016).
Menendez-Gonzalez, J. B. et al. Gata2 as a crucial regulator of stem cells in adult hematopoiesis and acute myeloid leukemia. Stem Cell Reports 13, 291–306 (2019).
Schmidt, L. et al. CEBPA-mutated leukemia is sensitive to genetic and pharmacological targeting of the MLL1 complex. Leukemia 33, 1608–1619 (2019).
Pundhir, S. et al. Enhancer and transcription factor dynamics during myeloid differentiation reveal an early differentiation block in cebpa null progenitors. Cell Rep. 23, 2744–2757 (2018).
Johnson, K. D., Soukup, A. A. & Bresnick, E. H. GATA2 deficiency elevates interferon regulatory factor-8 to subvert a progenitor cell differentiation program. Blood Adv 6, 1464–1473 (2022).
Minegishi, N. et al. Alternative promoters regulate transcription of the mouse GATA-2 gene. J Biol Chem 273, 3625–3634 (1998).
Snow, J. W. et al. A single cis element maintains repression of the key developmental regulator Gata2. PLoS Genet 6, e1001103 (2010).
Ohlsson, E. et al. Initiation of MLL-rearranged AML is dependent on C/EBPalpha. J Exp Med. 211, 5–13 (2014).
Shih, A. H. et al. Mutational cooperativity linked to combinatorial epigenetic gain of function in acute myeloid leukemia. Cancer Cell 27, 502–515 (2015).
Duy, C. et al. Rational targeting of cooperating layers of the epigenome yields enhanced therapeutic efficacy against AML. Cancer Discov 9, 872–889 (2019).
Christen, F. et al. Modeling clonal hematopoiesis in umbilical cord blood cells by CRISPR/Cas9. Leukemia https://doi.org/10.1038/s41375-021-01469-x (2021).
Rodrigues, N. P. et al. Haploinsufficiency of GATA-2 perturbs adult hematopoietic stem-cell homeostasis. Blood 106, 477–484 (2005).
Miharada, N., Rydstrom, A., Rak, J. & Larsson, J. Uncoupling key determinants of hematopoietic stem cell engraftment through cell-specific and temporally controlled recipient conditioning. Stem Cell Reports 16, 1705–1717 (2021).
Thoms, J. A. I. et al. Disruption of a GATA2-TAL1-ERG regulatory circuit promotes erythroid transition in healthy and leukemic stem cells. Blood 138, 1441–1455 (2021).
Fasan, A. et al. GATA2 mutations are frequent in intermediate-risk karyotype AML with biallelic CEBPA mutations and are associated with favorable prognosis. Leukemia 27, 482–485 (2013).
Celton, M. et al. Epigenetic regulation of GATA2 and its impact on normal karyotype acute myeloid leukemia. Leukemia 28, 1617–1626 (2014).
Metzeler, K. H. et al. Spectrum and prognostic relevance of driver gene mutations in acute myeloid leukemia. Blood 128, 686–698 (2016).
Al Seraihi, A. F. et al. GATA2 monoallelic expression underlies reduced penetrance in inherited GATA2-mutated MDS/AML. Leukemia 32, 2502–2507 (2018).
Mulet-Lazaro, R. et al. Allele-specific expression of GATA2 due to epigenetic dysregulation in CEBPA double mutant AML. Blood https://doi.org/10.1182/blood.2020009244 (2021).
You, X. et al. Gata2 −77 enhancer regulates adult hematopoietic stem cell survival. Leukemia 35, 901–905 (2021).
Yamazaki, H. et al. A remote GATA2 hematopoietic enhancer drives leukemogenesis in inv(3)(q21;q26) by activating EVI1 expression. Cancer Cell 25, 415–427 (2014).
Groschel, S. et al. A single oncogenic enhancer rearrangement causes concomitant EVI1 and GATA2 deregulation in leukemia. Cell 157, 369–381 (2014).
Yamaoka, A. et al. EVI1 and GATA2 misexpression induced by inv(3)(q21q26) contribute to megakaryocyte-lineage skewing and leukemogenesis. Blood Adv .4, 1722–1736 (2020).
Kiehlmeier, S. et al. Identification of therapeutic targets of the hijacked super-enhancer complex in EVI1-rearranged leukemia. Leukemia 35, 3127–3138 (2021).
Lugthart, S. et al. Clinical, molecular, and prognostic significance of WHO type inv(3)(q21q26.2)/t(3;3)(q21;q26.2) and various other 3q abnormalities in acute myeloid leukemia. J. Clin. Oncol. 28, 3890–3898 (2010).
Groschel, S. et al. High EVI1 expression predicts outcome in younger adult patients with acute myeloid leukemia and is associated with distinct cytogenetic abnormalities. J. Clin. Oncol. 28, 2101–2107 (2010).
Groschel, S. et al. Mutational spectrum of myeloid malignancies with inv(3)/t(3;3) reveals a predominant involvement of RAS/RTK signaling pathways. Blood 125, 133–139 (2015).
Sardina, J. L. et al. Transcription Factors Drive Tet2-Mediated Enhancer Demethylation to Reprogram Cell Fate. Cell Stem Cell 23, 727–741.e729 (2018).
Ramberger, E. et al. PRISMA and BioID disclose a motifs-based interactome of the intrinsically disordered transcription factor C/EBPalpha. iScience 24, 102686 (2021).
Mann, I. K. et al. CG methylated microarrays identify a novel methylated sequence bound by the CEBPB|ATF4 heterodimer that is active in vivo. Genome Res 23, 988–997 (2013).
Suzuki, T. et al. A screening system to identify transcription factors that induce binding site-directed DNA demethylation. Epigenetics Chromatin 10, 60 (2017).
Kallin, E. M. et al. Tet2 facilitates the derepression of myeloid target genes during CEBPalpha-induced transdifferentiation of pre-B cells. Mol. Cell 48, 266–276 (2012).
Tulstrup, M. et al. TET2 mutations are associated with hypermethylation at key regulatory enhancers in normal and malignant hematopoiesis. Nat. Commun. 12, 6061 (2021).
Figueroa, M. E. et al. Genome-wide epigenetic analysis delineates a biologically distinct immature acute leukemia with myeloid/T-lymphoid features. Blood 113, 2795–2804 (2009).
Chen, X. et al. Tumor suppressor CEBPA interacts with and inhibits DNMT3A activity. Sci. Adv. 8, eabl5220 (2022).
Figueroa, M. E. et al. Leukemic IDH1 and IDH2 mutations result in a hypermethylation phenotype, disrupt TET2 function, and impair hematopoietic differentiation. Cancer Cell 18, 553–567 (2010).
Rampal, R. et al. DNA hydroxymethylation profiling reveals that WT1 mutations result in loss of TET2 function in acute myeloid leukemia. Cell Rep. 9, 1841–1855 (2014).
Wang, Y. et al. WT1 recruits TET2 to regulate its target gene expression and suppress leukemia cell proliferation. Mol. Cell 57, 662–673 (2015).
Wilson, E. R. et al. Focal disruption of DNA methylation dynamics at enhancers in IDH-mutant AML cells. Leukemia 36, 935–945 (2022).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Cerami, E. et al. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2, 401–404 (2012).
Gao, J. et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci. Signal 6, pl1 (2013).
Labun, K. et al. CHOPCHOP v3: expanding the CRISPR web toolbox beyond genome editing. Nucleic Acids Res. 47, W171–W174 (2019).
Brinkman, E. K., Chen, T., Amendola, M. & van Steensel, B. Easy quantitative assessment of genome editing by sequence trace decomposition. Nucleic Acids Res. 42, e168 (2014).
Lee, B. T. et al. The UCSC Genome Browser database: 2022 update. Nucleic Acids Res. 50, D1115–D1122 (2022).
Doench, J. G. et al. Optimized sgRNA design to maximize activity and minimize off-target effects of CRISPR-Cas9. Nat. Biotechnol. 34, 184–191 (2016).
Quivoron, C. et al. TET2 inactivation results in pleiotropic hematopoietic abnormalities in mouse and is a recurrent event during human lymphomagenesis. Cancer Cell 20, 25–38 (2011).
Lee, Y. H., Sauer, B., Johnson, P. F. & Gonzalez, F. J. Disruption of the c/ebp alpha gene in adult mouse liver. Mol Cell Biol 17, 6014–6022 (1997).
de Boer, J. et al. Transgenic mice with hematopoietic and lymphoid specific expression of Cre. Eur. J. Immunol. 33, 314–325 (2003).
Stavropoulou, V. et al. MLL-AF9 Expression in Hematopoietic Stem Cells Drives a Highly Invasive AML Expressing EMT-Related Genes Linked to Poor Outcome. Cancer Cell 30, 43–58 (2016).
Ventura, A. et al. Restoration of p53 function leads to tumour regression in vivo. Nature 445, 661–665 (2007).
Langford, D. J. et al. Coding of facial expressions of pain in the laboratory mouse. Nat. Methods 7, 447–449 (2010).
Ge, Y. et al. The splicing factor RBM25 controls MYC activity in acute myeloid leukemia. Nat. Commun. 10, 172 (2019).
Dickins, R. A. et al. Probing tumor phenotypes using stable and regulated synthetic microRNA precursors. Nat. Genet. 37, 1289–1295 (2005).
Andrews, S. FastQC: A Quality Control Tool for High Throughput Sequence Data [Online]. Available online at: http://www.bioinformatics.babraham.ac.uk/projects/fastqc/ (2010).
Schmieder, R. & Edwards, R. Quality control and preprocessing of metagenomic datasets. Bioinformatics 27, 863–864 (2011).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Steinbaugh, M. bcbioRNASeq: R package for bcbio RNA-seq analysis [version 2; peer review: 1 approved, 1 approved with reservations]. F1000Res 6, 1976 (2018).
Patro, R., Duggal, G., Love, M. I., Irizarry, R. A. & Kingsford, C. Salmon provides fast and bias-aware quantification of transcript expression. Nat. Methods 14, 417–419 (2017).
Soneson, C., Love, M. I. & Robinson, M. D. Differential analyses for RNA-seq: transcript-level estimates improve gene-level inferences. F1000Res 4, 1521 (2015).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Mootha, V. K. et al. PGC-1alpha-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nat. Genet. 34, 267–273 (2003).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Krueger, F. Trim galore. A wrapper tool around Cutadapt and FastQC to consistently apply quality and adapter trimming to FastQ files [Online]. Available online at: http://www.bioinformatics.babraham.ac.uk/projects/trim_galore/ (2015).
Krueger, F. & Andrews, S. R. Bismark: a flexible aligner and methylation caller for Bisulfite-Seq applications. Bioinformatics 27, 1571–1572 (2011).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Ramirez, F., Dundar, F., Diehl, S., Gruning, B. A. & Manke, T. deepTools: a flexible platform for exploring deep-sequencing data. Nucleic Acids Res. 42, W187–191, (2014).
Kent, W. J., Zweig, A. S., Barber, G., Hinrichs, A. S. & Karolchik, D. BigWig and BigBed: enabling browsing of large distributed datasets. Bioinformatics 26, 2204–2207 (2010).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Li, H. et al. The sequence alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Acknowledgements
Work in the Porse lab was supported by grants from the Copenhagen University Hospital, the Capital Region of Copenhagen, the Independent Research Fund Denmark (9039-00189B) and through a center grant from the Novo Nordisk Foundation (Novo Nordisk Foundation Center for Stem Cell Biology, DanStem; Grant Number NNF17CC0027852). This work has been performed in the context of the Danish Research Center for Precision Medicine in Blood Cancers funded by the Danish Cancer Society (R223-A13071) and Greater Copenhagen Health Science Partners. Work in the Grebien lab was supported by the European Union’s Horizon 2020 research and innovation program (European Research Council grant agreement No 636855 and Austrian Science Fund (FWF), grants no. TAI-490 and P35628. Work in the Schoof lab was supported by a grant from the Independent Research Fund Denmark (case no. 2067-00053B). ASW was supported by grants from The Lundbeck Foundation (R303-2018-2868) and The Swedish Research Council (2015-00517). EH was supported by a grant from Forschungsförderung from the Fellinger Krebsforschungsverein. We thank members of the Grebien and Porse laboratories for their discussions. We thank Anna Fossum for her excellent research assistance.
Author information
Authors and Affiliations
Contributions
E.H., A.S.W., G.M., M.B.S., E.R., T.D.A., A.K.F., C.G., J.W., and E.M.S. performed experiments. E.H., AS.W., A.W., T.E., M.B.S., S.P., and E.M.S. analyzed data. E.H., A.S.W., F.G., and B.T.P. designed experiments. J.F. contributed essential material. M.M. and T.H. provided clinical data. E.H., A.S.W., F.G., and B.T.P. drafted the manuscript. All authors have proofread and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Ross Levine, Cihangir Duy and Cristina Pina for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Heyes, E., Wilhelmson, A.S., Wenzel, A. et al. TET2 lesions enhance the aggressiveness of CEBPA-mutant acute myeloid leukemia by rebalancing GATA2 expression. Nat Commun 14, 6185 (2023). https://doi.org/10.1038/s41467-023-41927-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-023-41927-x
This article is cited by
-
Cell of origin epigenetic priming determines susceptibility to Tet2 mutation
Nature Communications (2024)
-
Prognostic impact of CEBPA mutational subgroups in adult AML
Leukemia (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.