Abstract
Rational design of self-assembled DNA nanostructures has become one of the fastest-growing research areas in molecular science. Particular attention is focused on the development of dynamic DNA nanodevices whose configuration and function are regulated by specific chemical inputs. Herein, we demonstrate the concept of metal-mediated base-pair switching to induce inter- and intramolecular DNA strand displacement in a metal-responsive manner. The 5-hydroxyuracil (UOH) nucleobase is employed as a metal-responsive unit, forming both a hydrogen-bonded UOH–A base pair and a metal-mediated UOH–GdIII–UOH base pair. Metal-mediated strand displacement reactions are demonstrated under isothermal conditions based on the base-pair switching between UOH–A and UOH–GdIII–UOH. Furthermore, metal-responsive DNA tweezers and allosteric DNAzymes are developed as typical models for DNA nanodevices simply by incorporating UOH bases into the sequence. The metal-mediated base-pair switching will become a versatile strategy for constructing stimuli-responsive DNA nanostructures, expanding the scope of dynamic DNA nanotechnology.
Similar content being viewed by others
Introduction
Rational design of self-assembled DNA nanostructures, termed DNA nanotechnology1,2, is one of the fastest-growing research areas in molecular science. Hybridization based on strict base-paring rules has made DNA an excellent building block for constructing precisely defined molecular structures at the nanometer scale3,4. Of particular interest is the development of dynamic DNA nanodevices whose configuration and function are regulated by specific chemical inputs5,6,7. Such complex DNA systems are usually manipulated by toehold-mediated DNA strand displacement reactions (SDRs)8,9, which use DNA or RNA strands as input signals. There is also growing interest in the operation of DNA nanodevices by diverse signals such as pH10, light11,12,13, and small molecules14,15. Precise chemical modifications16 and conjugation with aptamers17 and antibodies18 have shown that DNA molecules can respond to specific chemical inputs or environmental stimuli.
Metal ions are often employed as chemical inputs in supramolecular chemistry19. Over the past few decades, a variety of metal-responsive molecular switches and machines have been synthesized20,21,22 by exploiting the specific affinity between ligands and metal species. However, the construction of metal-responsive DNA systems is still in its infancy, as it has relied on limited types of metal–DNA interactions, such as potassium-dependent induction of G-quadruplexes23 and metal-mediated formation of interstrand T–HgII–T and C–AgI–C complexes24 (termed as metal-mediated base pairs25,26,27).
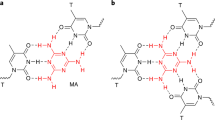
In this study, we demonstrate the concept of metal-mediated base-pair switching to induce inter- and intramolecular strand exchange in response to metal ions (Fig. 1). First, metal-mediated regulation of SDRs, one of the most fundamental processes for operating DNA nanodevices and computing circuits, is examined. Metal-mediated manipulation of DNA tweezers28 and regulation of the catalytic activity of a deoxyribozyme (DNAzyme)29 are also studied as typical models of DNA molecular devices. To enable rational sequence design of metal-responsive DNA structures, we utilize a noncanonical bifacial nucleobase, 5-hydroxyuracil (UOH), which can form both a Watson–Crick type hydrogen-bonded UOH–A base pair and a metal-mediated UOH–M–UOH base pair (M = GdIII, etc.) with its bidentate metal-binding site (i.e., an adjacent 4-carbonyl and a 5-hydroxy groups)30,31,32. Our previous studies have shown that the thermal stability of DNA duplexes containing UOH–UOH mismatches is significantly enhanced by the formation of UOH–GdIII–UOH base pairs, while duplexes with UOH–A base pairs are destabilized by the addition of GdIII. This raises the possibility that the base-pairing partner of the UOH bases can be switched in response to GdIII. We therefore conceived of the idea that GdIII-mediated base-pair switching between UOH–A and UOH–GdIII–UOH can be applied to the construction of dynamic DNA systems such as SDRs, DNA tweezers, and allosteric DNAzymes. Since UOH nucleobases can be easily synthesized and incorporated into DNA strands, sequence design strategies using metal-responsive UOH bases are expected to greatly expand the toolbox of dynamic DNA nanotechnology.
Base-pair switching between hydrogen-bonded UOH–A pairs and metal-mediated unnatural UOH–GdIII–UOH pairs can induce SDRs in response to GdIII ions. UOH nucleotides in the DNA strands are highlighted in navy blue. Other possible coordinating ligands on the GdIII ions, such as water molecules and neighboring nucleobases, are not shown for simplicity30.
Results
Metal-mediated DNA strand displacement reactions (SDRs)
Dynamic DNA nanotechnology relies heavily on SDRs8,9. SDRs are usually initiated by the binding of an invading strand to a single-stranded segment called a toehold. The development of SDRs triggered by stimuli other than oligonucleotides is increasingly attracting attention as an expansion of the DNA nanotechnology toolbox. In the past, binding of small molecules to aptamers15, pH changes33, photo-irradiation34,35, and metal–ligand interaction36,37 have been employed to trigger SDRs. In this study, we investigated toehold-free, metal-mediated SDRs based on base-pair switching between UOH–A and UOH–GdIII–UOH at the terminus of each strand.
Prior to the SDR experiments, metal-mediated regulation of DNA hybridization was first studied using oligonucleotides containing four UOH bases. Three 16-mer DNA strands were designed so that strand 1 can hybridize with strand 2 via UOH–GdIII–UOH base pairing and with strand 3 via UOH–A base pairing at their termini (Fig. 2a). The thermal stability of each duplex (i.e., 1·2 and 1·3) in the absence and presence of GdIII ions was evaluated by melting analysis (Supplementary Figs. 1a and 2a). Under conditions without GdIII ions, the melting temperature (Tm) of duplex 1·3 containing four UOH–A pairs was higher (Tm = 45.3 °C) than that of duplex 1·2 containing four UOH–UOH mismatch pairs (37.9 °C). Similar to previous studies30,31, the addition of GdIII ions significantly increased the stability of duplex 1·2 (ΔTm = +26.2 °C with 4 equiv of GdIII ions). The stabilization of the duplex by GdIII was ascribed to the formation of interstrand UOH–GdIII–UOH complexes, which was confirmed by UV titration experiments and ESI-TOF mass spectrometry (Supplementary Fig. 1). On the other hand, duplex 1·3 was destabilized by the addition of GdIII ions (ΔTm = –6.4 °C). This may be due to the weakening of the UOH–A base pairing by the binding of GdIII ions to the UOH bases30,31,32 (Supplementary Fig. 2). As a result, duplex 1·2 containing UOH–GdIII–UOH base pairs was found to be much more stable than duplex 1·3 containing UOH–A base pairs upon addition of GdIII (Fig. 2b). These results indicate that the addition of GdIII ions reversed the order of the stability of the duplexes. The same trend was observed for DNA strands containing three UOH bases (Supplementary Fig. 3). Addition of GdIII ions (3 equiv) stabilized duplex 1′·2′ with three UOH–UOH pairs (ΔTm = +25.2 °C) and destabilized duplex 1′·3′ with three UOH–A pairs (ΔTm = −4.4 °C). The degree of duplex (de)stabilization (ΔTm) was slightly greater when four UOH bases were incorporated (Supplementary Fig. 4). Incorporation of more than four UOH bases was thought to cause undesirable intramolecular metal complexation. Therefore, DNA strands 1 and 2 containing four consecutive UOH bases were employed in the SDR experiments.
a Schematic representation and base sequences. b Melting temperatures (Tm) of duplexes 1·2 (with UOH–UOH base pairs) and 1·3 (with UOH–A base pairs) in the absence and presence of GdIII ions (4 equiv). N = 3 independent experiments. Data are presented as mean values ± SEM. c Native PAGE analysis of an equimolar mixture of strands 1, 2, and 3 in the absence and presence of GdIII ions (4 equiv). The samples were annealed prior to the analysis. The strand 3 was labeled with FAM for detection. The authentic samples (strand 3 and pre-annealed duplex 1·3) were employed as the markers. Three independent experiments were performed. d UV absorption spectra of the mixture at 25 °C. l = 1 cm. The samples were annealed after the addition of GdIII ions (4 equiv). e The yields of duplex 1·3 and single strand 3 in the presence of GdIII ions at different concentrations. The yields were estimated by comparing the band intensities on the gel with those of the markers. N = 3 independent experiments. Data are presented as mean values ± SEM. f Reversible regulation of the hybridization behaviors of UOH-containing strand 1 by the alternate addition and removal of GdIII ions under isothermal conditions (25 °C). N = 4 independent experiments. Data are presented as mean values ± SEM. [DNA strand] = 2.0 μM each, [GdCl3] = 0 or 8.0 μM (1 equiv per UOH–UOH pair, otherwise noted) in 10 mM HEPES (pH 8.0), 100 mM NaCl. Source data are provided as a Source Data file.
Next, an equimolar mixture of strands 1, 2, and 3 (labeled with a fluorophore FAM) was annealed in the absence and presence of GdIII ions to see which duplex (1·2 or 1·3) is preferentially formed. Native polyacrylamide gel electrophoresis (PAGE) analysis confirmed products containing strand 3 by visualization with FAM fluorescence (Fig. 2c) and all products by further staining (Supplementary Fig. 5). The results revealed that duplex 1·3 was mainly formed (81%) under the GdIII-free condition. The addition of GdIII ions (4 equiv) resulted in the release of strand 3 (90%) and the formation of duplex 1·2. UV spectroscopy showed that hybridization of duplex 1·2 was accompanied by metal complexation of the UOH bases (Fig. 2d). The yield of the formed duplex increased as the amount of GdIII ions increased from 0 to 4.0 equiv and was little changed by the addition of excess GdIII ions (Fig. 2e). These results show that hybridization preference of UOH-containing strand 1 was changed in response to GdIII.
Under isothermal conditions, sequential addition and removal of GdIII ions resulted in reversible and alternating formation of duplexes 1·2 and 1·3 (Fig. 2f). Four equivalents of GdIII ions and a chelating agent EDTA were added every 2 h at 25 °C. Native PAGE analysis of the hybridization products showed that the removal of GdIII ions by EDTA induced the formation of duplex 1·3 containing UOH–A pairs. Subsequent addition of GdIII ions was found to regenerate duplex 1·2 via the interstrand UOH–GdIII–UOH complexation. Thus, the hybridization partner of strand 1 containing UOH bases was confirmed to be reversibly replaced by the addition and removal of GdIII ions.
Based on these results, metal-triggered SDRs were investigated by using longer DNA strands (Fig. 3a). Since the starting duplex (4·6 or 7·9) is long enough, the SDR should proceed via binding of the invading strand (5 or 8) and subsequent branch migration process to replace the incumbent strand (6 or 9). Like strands 1–3, strands 4 and 5 have four consecutive UOH bases, and strand 6 has four A bases at their termini. Strands 7, 8, and 9 have an additional natural nucleotide (C or G) at the terminus to stabilize the duplexes. Even with the terminal G–C base pairs, the addition of GdIII reversed the thermal stability of the duplex containing UOH–UOH pairs and the duplex containing UOH–A pairs (Supplementary Fig. 6). To examine metal-mediated SDRs, 4 equiv of GdIII ions were added to a mixture of a duplex containing UOH–A base pairs (4·6 or 7·9) and a UOH-modified invading strand (5 or 8, respectively). Two of the three strands (i.e., 4 and 5, or 7 and 8) were labeled with a fluorophore and a quencher, respectively. Thus, the progression of SDRs to form a duplex containing UOH–GdIII–UOH base pairs (4·5 or 7·8) can be monitored by fluorescence quenching.
a Design of metal-mediated SDRs. Strands 4 and 7 were labeled with a fluorophore (FAM) and strands 5 and 8 with a quencher (Dabcyl). b Time-course analysis of GdIII-triggered SDRs. The reaction was started by the addition of GdIII ions (4 equiv). In the control experiment, strands 7t and 8t, in which UOH bases are replaced with natural T bases, were used instead of strands 7 and 8. The toehold-mediated reaction was carried out with FAM-labeled 7t, 8 s (21-mer that lacks the terminal 5′-GUOHUOHUOHUOH-3′), and Dabcyl-labeled 9. c Time-course analysis of EDTA-triggered SDRs. The reaction was started by the addition of EDTA (4 equiv) to remove GdIII ions. [DNA strand] = 2.0 µM each in 10 mM HEPES buffer (pH 8.0), 100 mM NaCl, 25 °C. The SDRs were monitored by the changes in the fluorescence of the FAM. λex = 495 nm, λem = 519 nm. Source data are provided as a Source Data file.
The results of the GdIII-triggered SDRs are shown in Fig. 3b. The fluorescence decreased immediately after the addition of GdIII ions, suggesting that SDR was initiated in response to GdIII. The SDR proceeded at a similar rate regardless of the terminal G–C base pair. When the UOH bases of strands 7 and 8 were replaced with natural T bases (7t and 8t), little SDRs progressed with the addition of GdIII ions. This result confirms that the SDRs were triggered by metal complexation of the UOH bases, inducing base-pair switching from UOH–A to UOH–GdIII–UOH. Under these conditions, GdIII-mediated SDRs were slower than conventional toehold-mediated SDRs8,9. Complexation of UOH–GdIII–UOH is completed within 1 min, as suggested by UV absorption analysis (Supplementary Fig. 7a). The results suggest that the subsequent branch migration is slowed down due to the structural distortion caused by UOH–GdIII–UOH base pairing. In addition, the strand displacement may be delayed by the binding of the UOH-modified invading strands (5 or 8) in a parallel orientation or by homo-dimerization of the invading strands via the UOH–GdIII–UOH complexation (Supplementary Fig. 8). It is worth noting that the SDR using UOH-containing strands is specifically triggered by certain lanthanide ions (Supplementary Fig. 9). The lanthanide ion EuIII triggered the SDR as GdIII does, while other transition metal ions such as CuII and ZnII hardly induced SDR. The metal specificity is in good agreement with the fact that DNA duplexes containing UOH–UOH base pairs are stabilized only in the presence of lanthanide ions30.
The addition of EDTA can remove GdIII ions from the UOH–GdIII–UOH base pairs inside the duplexes, although the demetallation reaction is slower than complexation (Supplementary Fig. 7b). Under GdIII-free conditions, the SDR can be initiated by adding strand 9 containing AAAAG at the terminus to duplex 7·8 with UOH–UOH mismatches (Supplementary Fig. 10). Thus, it was expected that the addition of EDTA would induce SDRs from duplexes with UOH–GdIII–UOH (4·5 or 7·8) to duplexes with UOH–A pairs (4·6 or 7·9, respectively). When EDTA ([EDTA]/[GdIII] = 1.0) was added to a mixture of a GdIII-bridged duplex (4·5 or 7·8) and the other strand (6 or 9), the fluorescence was gradually increased. This result indicates that SDRs proceeded in the reverse direction (Fig. 3c). The slow rate of the EDTA-induced SDRs could be improved by increasing the number of UOH bases or by adding terminal bases only to the toehold and the incoming strand.
As discussed above, GdIII-triggered SDRs using the UOH base as a metal-binding site are now possible. GdIII destabilized the UOH–A base pairs and induced the formation of the UOH–GdIII–UOH base pairs. This base-pair switching promoted the displacement of the A-containing incumbent strands with the UOH-containing invading strand. The SDRs were found to proceed in the reverse direction upon removal of the GdIII ions, which induces base-pair switching from UOH–GdIII–UOH to UOH–A. Note that the SDRs developed in this study are reversible even though there are no exposed toehold regions in the strands. Since GdIII ions are unlikely to interact with natural nucleobases, the GdIII-triggered SDRs using unnatural UOH nucleobases would have significant advantages, especially when integrated into cascade SDRs such as DNA computing circuits. Thus, SDRs based on the base-pair switching between UOH–A and UOH–GdIII–UOH may provide a useful strategy for designing a wide variety of stimuli-responsive DNA molecular systems.
Development of metal-responsive DNA tweezers using UOH bases
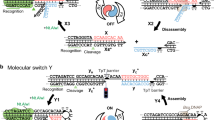
Based on the results of the strand displacement experiments, we further applied the UOH bases to the development of metal-responsive DNA molecular devices. As the simplest model, a pair of DNA tweezers28,38,39,40 was created that switch its structure upon metal complexation of the UOH bases (Fig. 4a). DNA tweezers are nanodevices consisting of two arms that take two forms, closed and open. Despite their simple structure, DNA tweezers have been applied to capture target proteins41 and regulate enzyme functions42, making them an important prototype of DNA molecular machines. The base sequence of the metal-responsive tweezers was designed based on Yurke’s original DNA tweezers28 operated by toehold-mediated SDR using fuel strands (Fig. 4b). The tweezers consist of two DNA duplex arms connected by a single-strand hinge. Hybridization of complementary oligonucleotides (strand d) at both dangling ends locks the DNA tweezers into a closed state. The UOH bases were introduced into strand d so that the tweezers are closed by UOH–A base pairing. The addition of GdIII ions was expected to destabilize the UOH–A base pairs and to induce the release of strand d associated with the intramolecular UOH–GdIII–UOH base pairing, which would open the tweezers.
a Design of GdIII-responsive DNA tweezers. U represents UOH nucleotides. b The base sequence of the UOH-modified DNA tweezers used in this study. c Native PAGE analysis of the structures of DNA tweezers in the absence and presence of GdIII ions. Strand b was labeled with FAM for detection. Two authentic samples, the DNA tweezers without strand d (open state) and with a closing strand containing T in place of UOH (closed state), were used as the markers. 15% gel at 20 °C. Three independent experiments were performed. (*) A dimeric structure28 in which two DNA tweezers are connected by binding of strand d. d Fluorescence analysis of the tweezers. Strand b was labeled with FAM and Dabcyl at both termini. λex = 495 nm, 25 °C. The samples were annealed prior to the measurement. e Opening of the tweezers triggered by the GdIII addition. λex = 495 nm, λem = 519 nm, 25 °C. f Closure of the tweezers triggered by the addition of EGTA (ethyleneglycol bis(2-aminoethyl ether)-N,N,N’,N’-tetraacetic acid), which selectively removes GdIII ions in the buffer containing MgII ions. Closure caused by the addition of strand d is also shown as a broken line. g, h Repeated opening and closing of the tweezers by alternating addition and removal of GdIII ions observed by native PAGE. Three independent experiments were performed. The bands were assigned by comparison with the authentic samples. h Repeated opening and closing of the tweezers observed by fluorescence analysis. [DNA] = 2.0 µM each in 10 mM HEPES buffer (pH 8.0), 100 mM NaCl, 3 mM MgCl2, [GdCl3] = 0 or 24 µM (otherwise noted), [EGTA] = 0 or 24 µM. Source data are provided as a Source Data file.
The structure of the UOH-modified DNA tweezers was first evaluated by native PAGE analysis (Fig. 4c). Each band was characterized by comparing with the mobility of reference samples in which UOH bases were replaced with T bases. When all strands were annealed in the absence of GdIII ions, a band corresponding to the closed state was mainly observed. In the presence of GdIII ions, a new band with higher mobility appeared, indicating the formation of an open state consisting of strands a, b, and c only. The open state was formed more efficiently by increasing the amount of GdIII ions. In the following experiments, the DNA tweezers were operated with 12 equiv of GdIII ions.
The metal-mediated switching between the closed and open states was further examined by FRET analysis (Fig. 4d). The hinge strand b was labeled with a fluorophore and a quencher. The fluorescence observed in the presence of GdIII ions (red solid line) was significantly higher than that observed in the absence of GdIII ions (black solid line). This suggests that the closed state was formed in the absence of GdIII ions, in which the FRET pairs were in close proximity to each other, while the open state was formed upon metal addition. The yields of both states were approximated by comparison with the control experiments (broken lines). The results showed that 73 ± 4% of the tweezers were closed in the absence of GdIII ions, whereas 94 ± 1% were open after annealing with 12 equiv of GdIII ions. Both values were in good agreement with the yields estimated from the band intensities in the PAGE analysis (Fig. 4c).
Next, the time course of the metal-mediated tweezer action was analyzed at a constant temperature of 25 °C. The fluorescence increased rapidly with the addition of GdIII ions, indicating that the metal addition triggered the tweezer opening under isothermal conditions (Fig. 4e). The opening reaction was faster than the SDR progression, reaching equilibrium within 5 min. This may be because the structure transformation of the DNA tweezers is based on the intramolecular UOH–GdIII–UOH base pairing and the strand d is sufficiently short to dissociate easily. Conversely, the addition of a chelating agent EGTA (ethyleneglycol bis(2-aminoethyl ether)-N,N,N’,N’-tetraacetic acid), which can selectively remove GdIII ions in the buffer containing MgII, caused the tweezers to close (Fig. 4f). The closing reaction was nearly complete in 1 h, but the rate of closure was slower than that induced by the addition of strand d (broken line). This may be due to the somewhat slower rate at which GdIII ions are removed from the UOH–GdIII–UOH base pairs inside the duplexes (Supplementary Fig. 7b). These results indicate that the UOH-modified DNA tweezers can be operated reversibly in response to GdIII ions.
Furthermore, the successive opening and closing of the tweezers were demonstrated by the sequential addition and removal of GdIII ions. GdIII ions (12 equiv) and EGTA (12 equiv) were added alternately to the pre-annealed DNA tweezers at a time interval of 2 h at 25 °C. Native PAGE and FRET analyses showed that the DNA tweezers were opened and closed repeatedly, undergoing structural changes four times (Fig. 4g, h). Although GdIII–EGTA complexes accumulated in this reaction, metal complexation proved to be useful in manipulating DNA tweezers under isothermal conditions. All results shown here suggest that the tweezers operate based on metal-mediated base-pair switching of the UOH bases without any toehold regions. Thus, the UOH nucleobase can be used as a versatile building block for constructing metal-responsive DNA nanodevices other than molecular tweezers.
Development of metal-dependent allosteric DNAzymes using UOH bases
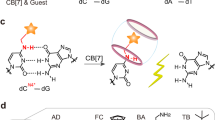
Deoxyribozymes (DNAzymes) are DNA oligomers with catalytic activity that were originally obtained by an in vitro selection method (SELEX) using nucleic acid libraries with a random sequence29. Among a variety of DNAzymes, RNA-cleaving DNAzymes, which cleave oligonucleotide substrates containing a ribonucleotide, are widely used to construct DNA molecular machines and computing circuits43,44. Allosteric regulation of DNAzyme activity is of growing interest as a promising strategy for constructing stimuli-responsive DNA systems45. We have previously synthesized CuII-responsive allosteric DNAzymes by introducing metal-mediated artificial base pairs (H–CuII–H or ImC–CuII–ImC) that stabilize DNA duplexes only in the presence of specific metal ions46,47,48,49,50. Here, we developed a metal-responsive allosteric DNAzyme by applying a base-pair switching system between hydrogen-bonded UOH–A base pairs and metal-mediated UOH–GdIII–UOH base pairs (Fig. 5a).
a Schematic representation of the GdIII-responsive allosteric DNAzyme. U represents UOH nucleotides. b Base sequence of the UOH-modified DNAzyme (UOH-DNAzyme). DNA strands containing an adenosine ribonucleotide (rA) at the cleavage site were used as the substrate. Both the catalytically inactive form (without GdIII) and the active form (with GdIII) are shown. The original NaA43 DNAzyme was modified by introducing UOH bases, and surrounding nucleobases (shown in orange) were further altered to stabilize the inactive structure in the absence of GdIII ions. c RNA-cleaving activity of UOH-DNAzyme, the original NaA43 DNAzyme, and T-DNAzyme in the absence and presence of GdIII ions (3 equiv). N = 3 (for UOH-DNAzyme) and N = 4 (for the others) independent experiments. Data are presented as mean values ± SEM. T-DNAzyme has T bases in place of UOH bases, thus lacking the metal binding affinity. The substrate cleavage was analyzed by denaturing PAGE using a substrate labeled with a fluorophore (FAM). d Activation of UOH-DNAzyme by the addition of GdIII ions (3 equiv). The activity of UOH-DNAzyme in the presence and absence of GdIII ions (3 equiv) is also shown by the red lines. N = 3 independent experiments. Data are presented as mean values ± SEM. e Deactivation of UOH-DNAzyme by removing GdIII ions. An equimolar amount of EDTA was added to remove the GdIII ions. N = 4 independent experiments. Data are presented as mean values ± SEM. [UOH-DNAzyme] = 1.0 μM, [substrate] = 10 μM, [GdCl3] = 3.0 μM (3 equiv, where applicable), [EDTA] = 3.0 μM (3 equiv, where applicable) in 10 mM HEPES (pH 7.0), 100 mM NaCl, 25 °C. Source data are provided as a Source Data file.
The metal-responsive DNAzyme was designed based on the secondary structure of the RNA-cleaving NaA43 DNAzyme reported previously51 (Fig. 5b). Three UOH–UOH base pairs were incorporated into the stem duplex as metal-binding sites. The sequence of the surrounding bases was modified to form a catalytically inactive structure with UOH–A base pairs in the absence of GdIII ions. The addition of GdIII ions was expected to switch the base pairing from UOH–A to UOH–GdIII–UOH, resulting in a structure transformation from the inactive to the catalytically active form.
The RNA-cleaving activity of the UOH-modified DNAzyme (named as UOH-DNAzyme) was evaluated in the absence and presence of GdIII ions. The reaction was initiated by the addition of a fluorophore-labeled substrate (10 equiv), and the time course of the reaction was monitored by denaturing PAGE (Fig. 5c). In the absence of GdIII ions, the activity of UOH-DNAzyme was efficiently suppressed (kobs = 4.8 × 10−3 h−1) compared to that of the original NaA43 DNAzyme (4.2 × 10−1 h−1). This result indicates that the UOH-DNAzyme adopted a catalytically inactive structure in the absence of GdIII ions. The addition of 3 equiv of GdIII ions enhanced the catalytic reaction of UOH-DNAzyme (kobs = 6.9 × 10−2 h−1). The activity of the unmodified DNAzyme was also increased by the addition of GdIII ions (kGd+/kGd– = 1.9), but the GdIII-induced activity enhancement of the UOH-DNAzyme was much more pronounced (kGd+/kGd– = 14.4). These results show that the interactions between UOH and GdIII play a pivotal role in DNAzyme activation. In control experiments using T-DNAzyme, in which all UOH bases were replaced with thymine (T) bases, no substrate cleavage was observed in the presence of GdIII ions, suggesting that UOH-DNAzyme was activated by the complexation of UOH bases with GdIII ions. UV spectroscopy also revealed the appearance of a new absorption band around 310 nm by the addition of GdIII ions (Supplementary Fig. 11), suggesting metal complexation of the UOH bases30,31. It can be concluded that the activity of UOH-DNAzyme is allosterically enhanced by the formation of UOH–GdIII–UOH base pairs, stabilizing the catalytically active structure. It should also be noted that the UOH-DNAzyme showed more efficient switching capability (kGd+/kGd− = 14.4) than the CuII-responsive DNAzyme (kCu+/kCu− = 5.9)47 developed by incorporating an H–CuII–H base pair into the same parent DNAzyme. This is thought to be due to a unique design strategy based on the bifacial nature of the UOH bases, whereby the inactive secondary structure (with UOH–A base pairs) and the active structure (with UOH–GdIII–UOH pairs) are formed stably in the absence and presence of GdIII ions, respectively.
We found that the DNAzyme activity during the reaction can be controlled by adding and removing GdIII ions. The substrate was treated with UOH-DNAzyme in the absence of GdIII ions for 2 h, and then GdIII ions were added. Time-course analysis showed that the addition of GdIII ions increased the DNAzyme activity to the level observed with GdIII ions from the beginning (Fig. 5d). When the chelating agent EDTA was added to remove GdIII ions, UOH-DNAzyme was immediately inactivated (Fig. 5e). It is interesting to note that the DNAzyme activity changed a short time after the addition or removal of GdIII ions. This is probably because the intramolecular structure transformation requires a little time.
All results presented here show that the activity of UOH-DNAzyme is allosterically regulated in response to GdIII ions under isothermal conditions. This is due to molecular design based on the base-pair switching between UOH–A and UOH–GdIII–UOH. Although the duplex structure around the UOH–GdIII–UOH base pairs is presumed to be distorted30, the UOH-modified DNAzyme showed catalytic activity in the presence of GdIII ions. Note that UOH bases served as metal-responsive switches to induce DNA structural conversion even when located at internal positions. Sequence design based on the base-pair switching of UOH may be applicable not only to allosteric DNAzymes but also to the development of various metal-responsive functional DNA devices.
Discussion
In this study, we have shown that GdIII-dependent DNA nanostructure transformation is possible by base-pair switching between hydrogen-bonded UOH–A base pairs and metal-mediated UOH–GdIII–UOH base pairs. In the absence of GdIII ions, UOH forms UOH–A base pairs, which are similar to Watson–Crick T–A pairs. When GdIII ions are added as external stimuli, the UOH–A base pair is destabilized and the UOH–GdIII–UOH base pairs can be preferentially formed instead. The addition and removal of GdIII ions result in the exchange of hybridization partners of the UOH-containing strands even under isothermal conditions. The introduction of multiple UOH bases at the termini of DNA strands enabled a GdIII-triggered strand displacement reaction (SDR), although it is not as fast as standard toehold-mediated SDRs. This metal-responsive SDR is reversible and does not require any toehold sequences, making it a versatile approach to creating dynamic DNA molecular devices.
The GdIII-mediated base-pair switching was further applied to the operation of DNA tweezers as a simple model for DNA molecular devices. Gel electrophoresis and FRET analysis confirmed that the DNA tweezers repeatedly open and close in response to GdIII ions. We also confirmed that this method can be applied to control the activity of a catalytic DNA (DNAzyme). Here, we designed a GdIII-dependent transformation between a catalytically inactive structure containing UOH–A base pairs and an active structure with UOH–GdIII–UOH base pairs. As a result, we succeeded in activating and inactivating the DNAzyme in response to the metal. Thus, UOH nucleobases are expected to be widely applied to manipulate DNA molecular devices and machines using metal ions as external stimuli.
The use of metal ions as external stimuli in dynamic DNA nanotechnology has the following characteristics. (1) Because coordination bonds are generally more stable than hydrogen bonds, metal-mediated base pairing can significantly alter the stability of DNA duplexes25. This is advantageous for inducing strand displacement and structure transformation of DNA constructs. (2) Metal coordination is fundamentally reversible, allowing repeated manipulation of metal-DNA systems, as demonstrated with the DNA tweezers in this study. Reversible SDRs could be useful in developing advanced DNA computing circuits. (3) A wide variety of metal ions are available for this purpose, and equimolar amounts or small excesses of metal ions are sufficient as stimuli when suitable ligands are used. Multiple metal species can also be used as orthogonal stimuli based on specific interactions between metal ions and ligands46,48. (4) SDRs triggered by biologically relevant metal ions have a wide range of biological applications.
Metal-mediated base pairs composed of natural nucleobases (i.e., T–HgII–T and C–AgI–C)52,53,54,55 have been used to construct metal-responsive DNA devices. However, these metal ions can bind to T and C bases at undesired positions that are difficult to predict, making sequence design not easy. We have worked to rationally design metal-responsive DNA supramolecules by introducing metal-mediated base pairs with unnatural ligand-type nucleobases46,47,48,49,50. Compared to these conventional approaches, metal-mediated base-pair switching dramatically alters duplex stability and hybridization behavior, making it suitable for dynamic control of DNA nanostructures. Moreover, because only modified nucleotides were used as additional building blocks, the strategies presented here are highly compatible with standard DNA nanotechnologies.
The basic behaviors seen in the metal-mediated SDRs and DNA nanodevice presented here are based on the unique concept of metal-mediated base-pair switching of modified pyrimidine bases. In conventional pH-responsive10 and light-responsive systems11,12,13, only (de)hybridization of DNA strands is controlled by external stimuli. In contrast, our approach can reversibly switch two different states: a structure with UOH–A base pairs (i.e., inactive state) and one with UOH–GdIII–UOH base pairs (active state), as clearly demonstrated using the UOH-DNAzyme. These features provide a useful basis for the development of DNA nanodevices, molecular machines, computing circuits, and more advanced systems56. In contrast to the selection methods by which various metal-dependent DNAzymes have been found57, our strategy is primarily focused on the rational design of metal-responsive DNA materials. Modification of existing functional DNAs is also a suitable approach, as the DNA sequences can be easily designed by replacing natural T bases with UOH bases. UOH nucleoside and UOH-modified oligonucleotides are readily available, and UOH bases function as metal-triggered switches not only at the termini of the DNA strands but also internally. Therefore, the UOH bases will be a useful building block for designing metal-responsive dynamic DNA architectures.
It should also be mentioned that metal-mediated SDR at lower concentrations is required for practical application in DNA nanotechnology. Because of the potential interaction of GdIII ions with phosphate groups, the use of other metal ions that rarely interact with natural DNA may be more appropriate for applications in complex systems consisting of many DNA strands. Metal-mediated base-pair switching can also be possible with other 5-substituted pyrimidine bases such as 5-carboxyuracil (caU)58 and N,N-dicarboxymethyl-5-aminouracil (dcaU)59, both of which were found to form more stabilizing metal-mediated base pairs (caU–CuII–caU and dcaU–GdIII–dcaU) as well as hydrogen-bonded base pairs (caU–A and dcaU–A). We believe that this method can be extended to other metal ions by designing bifacial nucleobases with different coordination functionality at the 5-position or by modifying cytosine bases. Thus, the principle of metal-mediated base-pair switching appears to be a versatile approach that extends the scope of dynamic DNA nanotechnology. The application of these bifacial nucleobases to the metal-dependent reconfiguration of DNA nanostructures is currently under investigation in our laboratory.
Methods
Materials and equipment
All natural DNA strands, including FAM-labeled strands, Dabcyl-labeled strands, and substrate strands containing a riboadenosine (rA) were purchased from Japan Bio Service Co., Ltd. (Saitama, Japan) at HPLC purification grade. The sequences of the DNA strands used in this study are summarized in Supplementary Table 1. Oligonucleotide concentrations were determined based on UV absorbance at 260 nm. GdCl3·6H2O (99.9% purity) purchased from Soekawa Chemical Co. was used as a metal source without further purification. The gels were analyzed using Gel Doc EZ Imager and Image Lab software ver. 6.1.0 (Bio-Rad). All the graphs were produced by KaleidaGrapgh ver. 5.0.3 (HULINKS).
DNA synthesis
DNA strands containing 5-hydroxyuracil (UOH) nucleobases were synthesized on an automated DNA synthesizer (NTS M-2-MX DNA/RNA synthesizer)30. The DNA synthesis was carried out on a 1-μmol scale in a DMTr-on mode with ultramild deprotection phosphoramidites and reagents (Glen Research). Note that the phosphoramidite monomer of UOH deoxynucleoside was synthesized following the literature60,61 or purchased from Glen Research (catalog No. 10-1053). The coupling time of UOH was extended to 15 min. The products were deprotected using 28% NH3 aqueous solution at room temperature for 2–3 h and then purified and detritylated using a PolyPak II cartridge (Glen Research). Some of the UOH-containing strands were prepared by the ligation of a shorter UOH-containing strand and a natural strand by using a T4 DNA ligase. The oligonucleotides were purified by reverse-phase HPLC (Waters XBridge C18 column) or by denaturing polyacrylamide gel electrophoresis. All DNA strands were identified by MALDI-TOF or ESI-TOF mass spectrometry (Supplementary Table 2).
Duplex melting analysis
The samples were prepared by mixing the DNA strands (2 µM) in 10 mM HEPES buffer (pH 8.0) containing 100 mM NaCl. After the addition of GdIII ions (0–6 equiv), the solutions were heated to 85 °C and cooled slowly to 4 °C at a rate of −1.0 °C/min. Absorbance at 260 nm was monitored by UV-1800 and UV-1900 spectrophotometers (Shimadzu) equipped with a TMSPC-8 temperature controller while the temperature was raised from 4 to 85 °C at a rate of 0.2 °C/min. Normalized absorbance shown in the figures was calculated as follows:
The melting temperature (Tm) was determined as the inflection point of the melting curve using LabSolutions Tm analysis software ver. 1.40 (Shimadzu) with a 17-point adaptive smoothing program. The average Tm values of at least three independent runs were calculated.
Mass spectrometry of the DNA duplexes
Electrospray ionization-time-of-flight (ESI-TOF) mass spectra were recorded on a Waters Micromass LCT premier. The samples (40 µM) were prepared in 20 mM NH4OAc buffer (pH 7.0) and annealed just before the measurements (from 85 to 4 °C, −1.0 °C/min).
Metal-dependent regulation of the hybridization behavior of UOH-containing DNA strands
Strand 3 was labeled with FAM. The samples were prepared by mixing the DNA strands (2 µM each) in 10 mM HEPES buffer (pH 8.0) containing 100 mM NaCl. The solutions were annealed (85 °C → 4 °C, −1.0 °C/min) in the absence or presence of GdIII ions. For the reversible regulation shown in Fig. 2f, after the solutions were annealed (85 °C → 25 °C, −1.0 °C/min), GdCl3 (4 equiv) and EDTA (4 equiv) were alternately added every 2 h at 25 °C. Native gel electrophoresis was carried out with 20% polyacrylamide gel in a cool incubator (4 °C). The bands were detected by FAM fluorescence, and the yield of each product was calculated by comparing the band intensities of the authentic samples on the same gel. The averages of at least three runs are summarized. The gel shown in Supplementary Fig. 5 was stained with SYBR Gold. The UV spectra of the samples were also recorded at 5 °C on a JASCO V-730 spectrometer.
Metal-mediated DNA strand displacement reactions (SDRs)
Strands 4 and 7 were labeled with a fluorophore (FAM) and strands 5 and 8 with a quencher (Dabcyl). The initial duplexes (4·6 or 7·9 for GdIII-triggered reaction, and 4·5 or 7·8 for EDTA-triggered reaction) were prepared by combining the two strands (2.0 µM each) in 10 mM HEPES buffer (pH 8.0) containing 100 mM NaCl in the absence or presence of GdIII ions (4 equiv). After annealing from 85 to 25 °C (−1.0 °C/min), the mixture was kept at 25 °C. After the addition of the other strand, the SDR was initiated by the addition of GdIII ions (4 equiv) or EDTA (4 equiv). The reaction progress was monitored by the changes in the fluorescence intensity of the FAM (λex = 495 nm, λem = 519 nm) using an FP-8350 fluorometer (JASCO). The fluorescent intensity was recorded in 10-min intervals for 8 h and subsequently in 30-min intervals. In the control experiments, the toehold-mediated SDR was started by adding the invading strands to the pre-annealed duplex. In the metal selectivity test (Supplementary Fig. 9), the fluorescence intensity (λex = 495 nm, λem = 520 nm) was recorded on a Varioskan LUX plate-reader (Thermo Fisher).
Metal-mediated operation of the DNA tweezers
Strand b was labeled with FAM at the 5′ end for native PAGE analysis or with FAM and Dabcyl at both termini for FRET analysis. The samples were prepared by combining DNA strands a, b, c, and d (2 µM each) in 10 mM HEPES buffer (pH 8.0) containing 100 mM NaCl and 3 mM MgCl2. The solutions were annealed (85 °C → 25 °C, −1.0 °C/min) in the absence or presence of GdIII ions (4, 8, 12, and 20 equiv). Native PAGE was carried out with 15% polyacrylamide gel in an incubator (20 °C). The bands were detected by FAM fluorescence, and the yields of the closed and the open states were estimated by comparing the band intensities. The averages of at least three runs are summarized. A mixture of strands a, b, and c was used as a marker corresponding to the open state. A mixture of strands a, b, c, and d′, in which UOH nucleotides were replaced with T, was used to indicate the closed state. For the FRET analysis, the fluorescence spectra were measured at 25 °C (λex = 495 nm).
Time-course study of the operation of the DNA tweezers
Strand b was labeled with FAM and Dabcyl at both termini. DNA strands a, b, c, and d (2 µM each) were mixed in 10 mM HEPES buffer (pH 8.0) containing 100 mM NaCl and 3 mM MgCl2. The mixtures were annealed (85 °C → 25 °C, −1.0 °C/min) in the absence or presence of GdIII ions (12 equiv). GdIII ions (12 equiv) or EGTA (12 equiv) were added while maintaining 25 °C. The change in the fluorescence intensity was monitored in 30-s intervals at 25 °C (λex = 495 nm, λem = 519 nm). In the control experiments, strand d was added to the pre-annealed mixture of strands a, b, and c.
Repetitive opening and closing of the tweezers
The sample was prepared in the absence of GdIII ions, as described above. With keeping the temperature at 25 °C, GdIII ions (12 equiv) or EGTA (12 equiv) were alternately added at the defined time points. The reaction progress was monitored by native PAGE and FRET analysis.
Metal-mediated regulation of DNAzyme activity
A DNAzyme strand (1.0 µM) was annealed (85 °C → 25 °C, −1.0 °C/min) in a reaction buffer (10 mM HEPES (pH 7.0), 100 mM NaCl) in the presence or absence of GdCl3 (3 equiv). The RNA-cleaving reaction was initiated by adding a FAM-labeled substrate (10 equiv). After incubating at 25 °C, the reaction was stopped by adding a 3:1 mixture of 7 M urea and the loading buffer. The progress of the DNAzyme reaction was analyzed by denaturing PAGE. The percentage of the cleaved product (F) was calculated as follows:
where Ic and Iu are the band intensities of the cleaved and the uncleaved substrate, respectively. The apparent first-order rate constants (kobs) were calculated from the initial velocity determined from the time points when F was less than 20%.
Regulation of DNAzyme activity during the reaction
For the GdIII-triggered DNAzyme activation, the RNA-cleaving reaction was initiated in the absence of GdIII ions. After 2 h, GdCl3 (3 equiv) was added. For the deactivation by the removal of GdIII ions, the reaction was started in the presence of GdIII ions (3 equiv). After 2 h, a chelating agent EDTA (3 equiv) was added to remove GdIII ions. The reaction progress was monitored by denaturing PAGE as described above.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All the data supporting the results of this study are available in this paper and the Supplementary Information. Source data are provided with this paper.
References
Seeman, N. C. & Sleiman, H. F. DNA nanotechnology. Nat. Rev. Mater. 3, 17068 (2017).
Seeman, N. C. Structural DNA Nanotechnology (Cambridge University Press, 2016).
Dey, S. et al. DNA origami. Nat. Rev. Methods Primers 1, 13 (2021).
Hong, F., Zhang, F., Liu, Y. & Yan, H. DNA origami: scaffolds for creating higher order structures. Chem. Rev. 117, 12584–12640 (2017).
DeLuca, M., Shi, Z., Castro, C. E. & Arya, G. Dynamic DNA nanotechnology: toward functional nanoscale devices. Nanoscale Horiz. 5, 182–201 (2020).
Zhang, Y. et al. Dynamic DNA structures. Small 15, e1900228 (2019).
Wang, F., Liu, X. & Willner, I. DNA switches: from principles to applications. Angew. Chem. Int. Ed. Engl. 54, 1098–1129 (2015).
Zhang, D. Y. & Seelig, G. Dynamic DNA nanotechnology using strand displacement reactions. Nat. Chem. 3, 103–113 (2011).
Simmel, F. C., Yurke, B. & Singh, H. R. Principles and applications of nucleic acid strand displacement reactions. Chem. Rev. 119, 6326–6369 (2019).
Mattath, M. N., Ghosh, D., Pratihar, S., Shi, S. & Govindaraju, T. Nucleic acid architectonics for pH-responsive DNA systems and devices. ACS Omega 7, 3167–3176 (2022).
Wang, C. et al. Photoresponsive DNA materials and their applications. Chem. Soc. Rev. 51, 720–760 (2022).
Lubbe, A. S., Szymanski, W. & Feringa, B. L. Recent developments in reversible photoregulation of oligonucleotide structure and function. Chem. Soc. Rev. 46, 1052–1079 (2017).
Kamiya, Y. & Asanuma, H. Light-driven DNA nanomachine with a photoresponsive molecular engine. Acc. Chem. Res. 47, 1663–1672 (2014).
Yang, H., Liu, H., Kang, H. & Tan, W. Engineering target-responsive hydrogels based on aptamer-target interactions. J. Am. Chem. Soc. 130, 6320–6321 (2008).
Zhang, Q.-L., Wang, L.-L., Liu, Y., Lin, J. & Xu, L. A kinetically controlled platform for ligand-oligonucleotide transduction. Nat. Commun. 12, 4654 (2021).
Madsen, M. & Gothelf, K. V. Chemistries for DNA nanotechnology. Chem. Rev. 119, 6384–6458 (2019).
Krissanaprasit, A., Key, C. M., Pontula, S. & LaBean, T. H. Self-assembling nucleic acid nanostructures functionalized with aptamers. Chem. Rev. 121, 13797–13868 (2021).
Ranallo, S., Sorrentino, D. & Ricci, F. Orthogonal regulation of DNA nanostructure self-assembly and disassembly using antibodies. Nat. Commun. 10, 5509 (2019).
McConnell, A. J., Wood, C. S., Neelakandan, P. P. & Nitschke, J. R. Stimuli-responsive metal-ligand assemblies. Chem. Rev. 115, 7729–7793 (2015).
Champin, B., Mobian, P. & Sauvage, J.-P. Transition metal complexes as molecular machine prototypes. Chem. Soc. Rev. 36, 358–366 (2007).
Durot, S., Heitz, V., Sour, A. & Sauvage, J.-P. Transition-metal-complexed catenanes and rotaxanes: from dynamic systems to functional molecular machines. Top. Curr. Chem. 354, 35–70 (2014).
Goswami, A., Saha, S., Biswas, P. K. & Schmittel, M. (Nano)mechanical motion triggered by metal coordination: from functional devices to networked multicomponent catalytic machinery. Chem. Rev. 120, 125–199 (2020).
Mergny, J.-L. & Sen, D. DNA quadruple helices in nanotechnology. Chem. Rev. 119, 6290–6325 (2019).
Tanaka, Y. et al. Structures, physicochemical properties, and applications of T–HgII–T, C–AgI–C, and other metallo-base-pairs. Chem. Commun. 51, 17343–17360 (2015).
Takezawa, Y., Müller, J. & Shionoya, M. Artificial DNA base pairing mediated by diverse metal ions. Chem. Lett. 46, 622–633 (2017).
Takezawa, Y. & Shionoya, M. Metal-mediated DNA base pairing: alternatives to hydrogen-bonded Watson–Crick base pairs. Acc. Chem. Res. 45, 2066–2076 (2012).
Müller, J. Nucleic acid duplexes with metal-mediated base pairs and their structures. Coord. Chem. Rev. 393, 37–47 (2019).
Yurke, B., Turberfield, A. J., Mills, A. P., Simmel, F. C. & Neumann, J. L. A DNA-fuelled molecular machine made of DNA. Nature 406, 605–608 (2000).
Ma, L. & Liu, J. Catalytic nucleic acids: biochemistry, chemical biology, biosensors, and nanotechnology. iScience 23, 100815 (2020).
Takezawa, Y., Nishiyama, K., Mashima, T., Katahira, M. & Shionoya, M. Bifacial base-pairing behaviors of 5-hydroxyuracil DNA bases through hydrogen bonding and metal coordination. Chem. Eur. J. 21, 14713–14716 (2015).
Nishiyama, K., Mori, K., Takezawa, Y. & Shionoya, M. Metal-responsive reversible binding of triplex-forming oligonucleotides with 5-hydroxyuracil nucleobases. Chem. Commun. 57, 2487–2490 (2021).
Nishiyama, K., Takezawa, Y. & Shionoya, M. pH-dependence of the thermal stability of metallo-DNA duplexes containing ligand-type 5-hydroxyuracil nucleobases. Inorg. Chim. Acta 452, 176–180 (2016).
Amodio, A. et al. Rational design of pH-controlled DNA strand displacement. J. Am. Chem. Soc. 136, 16469–16472 (2014).
Morihiro, K., Kodama, T., Waki, R. & Obika, S. Light-triggered strand exchange reaction using the change in the hydrogen bonding pattern of a nucleobase analogue. Chem. Sci. 5, 744–750 (2014).
Kou, B., Zhang, J., Huai, X., Liang, X. & Xiao, S.-J. Light-driven reversible strand displacement using glycerol azobenzene inserted DNA. RSC Adv. 5, 5055–5058 (2015).
Wang, L.-L. et al. Controllable DNA strand displacement by independent metal–ligand complexation. Chem. Sci. 12, 8698–8705 (2021).
Ding, W., Deng, W., Zhu, H. & Liang, H. Metallo-toeholds: controlling DNA strand displacement driven by Hg(II) ions. Chem. Commun. 49, 9953–9955 (2013).
Liu, M. et al. A DNA tweezer-actuated enzyme nanoreactor. Nat. Commun. 4, 2127 (2013).
Liang, X., Nishioka, H., Takenaka, N. & Asanuma, H. A DNA nanomachine powered by light irradiation. ChemBioChem 9, 702–705 (2008).
Elbaz, J., Wang, Z.-G., Orbach, R. & Willner, I. pH-stimulated concurrent mechanical activation of two DNA “tweezers”. A “SET-RESET” logic gate system. Nano Lett. 9, 4510–4514 (2009).
Zhou, C., Yang, Z. & Liu, D. Reversible regulation of protein binding affinity by a DNA machine. J. Am. Chem. Soc. 134, 1416–1418 (2012).
Hu, Y. et al. Switchable enzyme/DNAzyme cascades by the reconfiguration of DNA nanostructures. Chem. Eur. J. 20, 16203–16209 (2014).
Liu, M., Chang, D. & Li, Y. Discovery and biosensing applications of diverse RNA-cleaving DNAzymes. Acc. Chem. Res. 50, 2273–2283 (2017).
Orbach, R., Willner, B. & Willner, I. Catalytic nucleic acids (DNAzymes) as functional units for logic gates and computing circuits: from basic principles to practical applications. Chem. Commun. 51, 4144–4160 (2015).
Huang, Z., Wang, X., Wu, Z. & Jiang, J.-H. Recent advances on DNAzyme-based sensing. Chem. Asian J. 17, e2021014 (2022).
Takezawa, Y., Nakama, T. & Shionoya, M. Enzymatic synthesis of Cu(II)-responsive deoxyribozymes through polymerase incorporation of artificial ligand-type nucleotides. J. Am. Chem. Soc. 141, 19342–19350 (2019).
Nakama, T., Takezawa, Y., Sasaki, D. & Shionoya, M. Allosteric regulation of DNAzyme activities through intrastrand transformation induced by Cu(II)-mediated artificial base pairing. J. Am. Chem. Soc. 142, 10153–10162 (2020).
Takezawa, Y., Hu, L., Nakama, T. & Shionoya, M. Sharp switching of DNAzyme activity through the formation of a Cu(II)-mediated carboxyimidazole base pair. Angew. Chem. Int. Ed. Engl. 59, 21488–21492 (2020).
Nakama, T., Takezawa, Y. & Shionoya, M. Site-specific polymerase incorporation of consecutive ligand-containing nucleotides for multiple metal-mediated base pairing. Chem. Commun. 57, 1392–1395 (2021).
Rajasree, S. C., Takezawa, Y. & Shionoya, M. Cu(II)-mediated stabilisation of DNA duplexes bearing consecutive ethenoadenine lesions and its application to a metal-responsive DNAzyme. Chem. Commun. 59, 1006–1009 (2023).
Torabi, S.-F. et al. In vitro selection of a sodium-specific DNAzyme and its application in intracellular sensing. Proc. Natl. Acad. Sci. USA 112, 5903–5908 (2015).
Wang, Z. G., Elbaz, J., Remacle, F., Levine, R. D. & Willner, I. All-DNA finite-state automata with finite memory. Proc. Natl. Acad. Sci. USA 107, 21996–22001 (2010).
Wang, Z. G., Elbaz, J. & Willner, I. DNA machines: bipedal walker and stepper. Nano Lett. 11, 304–309 (2011).
Thomas, J. M., Yu, H. Z. & Sen, D. A mechano-electronic DNA switch. J. Am. Chem. Soc. 134, 13738–13748 (2012).
Lu, C.-H., Cecconello, A., Elbaz, J., Credi, A. & Willner, I. A three-station DNA catenane rotary motor with controlled directionality. Nano Lett. 13, 2303–2308 (2013).
Murata, S., Toyota, T., Nomura, S. M., Nakakuki, T. & Kuzuya, A. Molecular cybernetics: challenges toward cellular chemical artificial intelligence. Adv. Funct. Mater. 32, 2201866 (2022).
McGhee, C. E., Lake, R. J. & Lu, Y. Preparation of metalloDNAzymes. Artificial Metalloenzymes and Metallo DNAzymes in Catalysis: From Design to Applications (eds Diéguez, M. et al.) 41–68 (Wiley-VCH, 2018).
Takezawa, Y., Suzuki, A., Nakaya, M., Nishiyama, K. & Shionoya, M. Metal-dependent DNA base pairing of 5‑carboxyuracil with itself and all four canonical nucleobases. J. Am. Chem. Soc. 142, 21640–21644 (2020).
Mori, K., Takezawa, Y. & Shionoya, M. Metal-dependent base pairing of bifacial iminodiacetic acid-modified uracil bases for switching DNA hybridization partner. Chem. Sci. 14, 1082–1088 (2023).
Fujimoto, J., Tran, L. & Sowers, L. C. Synthesis and cleavage of oligodeoxynucleotides containing a 5-hydroxyuracil residue at a defined site. Chem. Res. Toxicol. 10, 1254–1258 (1997).
Morningstar, M. L., Kreutzer, D. A. & Essigmann, J. M. Synthesis of oligonucleotides containing two putatively mutagenic DNA lesions: 5-hydroxy-2′-deoxyuridine and 5-hydroxy-2′-deoxycytidine. Chem. Res. Toxicol. 10, 1345–1350 (1997).
Acknowledgements
This work was supported by JSPS KAKENHI Grant Numbers JP18H02081, JP21H02055, JP22K19100 to Y.T., JP21J11325 (Grant-in-Aid for JSPS Fellows) to K.M., and JP21H05022 to M.S. and MEXT KAKENHI Grant Numbers JP21H00384 (Molecular Engine), JP21H05866, JP23H04399 (Molecular Cybernetics) to Y.T., and JP16H06509 (Coordination Asymmetry) to M.S., Japan.
Author information
Authors and Affiliations
Contributions
Y.T., K.M., and M.S. conceived and designed the project. K.M., W.-E.H., K.N., and T.X. carried out the experiments with assistance from Y.T. and T.N. Y.T., K.M., and W.-E. H. analyzed the data. Y.T., K.M., and M.S. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Julian Tanner and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Takezawa, Y., Mori, K., Huang, WE. et al. Metal-mediated DNA strand displacement and molecular device operations based on base-pair switching of 5-hydroxyuracil nucleobases. Nat Commun 14, 4759 (2023). https://doi.org/10.1038/s41467-023-40353-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-023-40353-3
This article is cited by
-
Rational design of metal-responsive functional DNA supramolecules
Journal of Inclusion Phenomena and Macrocyclic Chemistry (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.