Abstract
Synthetic genetic circuits are programmed in living cells to perform predetermined cellular functions. However, designing higher-order genetic circuits for sophisticated cellular activities remains a substantial challenge. Here we program a genetic circuit that executes Pavlovian-like conditioning, an archetypical sequential-logic function, in Escherichia coli. The circuit design is first specified by the subfunctions that are necessary for the single simultaneous conditioning, and is further genetically implemented using four function modules. During this process, quantitative analysis is applied to the optimization of the modules and fine-tuning of the interconnections. Analogous to classical Pavlovian conditioning, the resultant circuit enables the cells to respond to a certain stimulus only after a conditioning process. We show that, although the conditioning is digital in single cells, a dynamically progressive conditioning process emerges at the population level. This circuit, together with its rational design strategy, is a key step towards the implementation of more sophisticated cellular computing.
Similar content being viewed by others
Introduction
Synthetic genetic circuits can be programmed in living cells to perform diverse cellular functions1,2 such as the integration of environmental signals3,4,5, the operation of predetermined algorithms6,7,8,9,10 and cellular dynamics control11,12,13,14. Some of these circuits have made possible a large variety of bioengineering applications, including disease therapeutics and diagnostics15,16,17 and process control in biosynthetic pathways18. Despite considerable progress, rationally designing genetic circuits with ever-larger complexity to conduct sophisticated cellular activities remains a substantial challenge2,19. Two recent studies reported significant progress, feasibly integrating logic operation and cellular memory20,21. This capability sets the stage for programming higher-order genetic circuits to endow cells with more complex sequential-logic computing.
The building of synthetic genetic circuits for adaptive learning, a type of complex sequential-logic function that is ubiquitous in organisms that range from microbes22,23 to primates24, has been widely anticipated2,19,25. A central paradigm of adaptive learning is Pavlovian conditioning24, in which an originally unrelated stimulus (for dogs, for example, ringing a bell) can be associated with another stimulus (for example, food), thus to evoke a specific response (for example, salivation) after a learning process. This history-dependent logic (that is, sequential logic) function involves both a combinatorial logic operation and storage/read-out of cellular memory and is therefore a good example to study the programming of complex genetic circuits for sophisticated functions.
Here we take an integrated design strategy to program a genetic circuit that performs a Pavlovian-like conditioning function in Escherichia coli. Analogous to classical Pavlovian conditioning, the circuit executes Learning and Recalling subfunctions: cells that harbour this genetic circuit respond to certain stimuli only after a learning process. To synthesize such a circuit, the paradigm of electronic circuit design is borrowed. The Pavlovian-like conditioning function is first coarsely formulated into a sequential-logic circuit for single simultaneous conditioning using Boolean mathematics. The circuit is then reduced to four genetic modules: two transcriptional AND gates, one genetic Memory device, and one genetic OR gate. Then, these modules are built using readily available genetic components; they are rationally optimized and further bottom–up assembled on the basis of the subfunctions for conditioning. During the module assembly, parameter sensitivity analysis is conducted to guide the module interconnections. The resultant genetic circuit successfully executes the Pavlovian-like conditioning function in living bacterial cells: single simultaneous conditioning can be observed on the single-cell level; on the population level, however, the conditioning process appears to be dynamically progressive.
Results
Circuit specifications for the conditioning function
The single simultaneous conditioning function can be abstracted as follows: initially, the system responds to an unconditioned stimulus (‘Food’) but does not respond to a conditioned stimulus (‘Ring’); however, the system can be trained in a single trial to learn the association between Food and Ring; consequently, the Ring acquires the capacity to evoke the system’s response that was originally evoked only by Food. We borrowed the electronic circuit design to formulate the logic circuit for single simultaneous conditioning (Supplementary Methods). A Memory module was introduced to represent the circuit internal state, with Memory=1 denoting that the circuit has remembered the association between Ring and Food after learning and Memory=0 denoting no association before learning. Then, the Karnaugh maps26 for circuit input/output and input/Memory functions were accordingly generated (Supplementary Figs 1,2). The corresponding simplified expressions are

where either Food alone or the combination of Ring and Memory would trigger the output (Fig. 1a, Supplementary Fig. 3), and
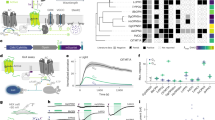
(a) The Karnaugh map for the input/output function of single simultaneous conditioning, visualizing the equation Output=Food | (Ring & Memory). (b) The Karnaugh map for the input/Memory function, which corresponds to the equation Memory=Memory | (Ring & Food). The Memory-ON state arises from either state self-sustainment or an association between Ring and Food. (c) The sequential-logic circuit design for the simultaneous conditioning according to the Karnaugh maps in (a) and (b). This circuit is composed of four modules: Learning AND gate, the Memory module, Recalling AND gate and output OR gate. (d) Genetic implementation of the single simultaneous conditioning circuit based on the sequential-logic design. Boxes represent the inputs, output and individual functional modules that correspond to those in Fig. 1c. Ring, conditioned stimulus; Food, unconditioned stimulus.

where the Memory-ON state can be achieved through either state self-sustainment or the co-occurrence of Ring and Food (Fig. 1b, Supplementary Fig. 3). According to Boolean mathematics26, many logic circuits can perform the two expressions. The simplest one is composed of four modules (Fig. 1c): the Learning AND gate (LAG) to associate Ring with Food by detecting their simultaneous appearance, the Memory module made up of an OR gate and a 0/1 Memory device to memorize the association of two stimuli as two discrete internal states (Memory=1 and Memory=0), the Recalling AND gate (RAG) to give a conditioned response according to the state of the Memory module, and another OR gate to wire conditioned response and unconditioned response together to produce output. The sequential information-processing performance of the overall simplest logic circuit is illustrated in Supplementary Fig. 4.
Genetic implementation of individual modules
To genetically implement the sequential-logic circuit in E. coli., two chemical inducers, salicylate (PSal) and arabinose (PBAD), were, respectively, utilized as Food and Ring (Supplementary Figs 5–7); the output of the overall single simultaneous conditioning circuit was green fluorescence protein (gfp) (Fig. 1d). For the realization of LAG and RAG, a design frame based on the interaction between mRNA and tRNA was adopted3 (Supplementary Fig. 8). In RAG, one input promoter controls the expression of transcription activator δ from phage φR73 (ref. 27), with two amber stop codons inside the coding sequence (φR73δtag), while the other input promoter drives tRNA supD, the amber stop codon suppressor (Fig. 2a). When both inputs appear, active δ protein will be produced and activate the PO promoter. LAG has the same architecture, except that T7ptag, the T7 polymerase bearing amber stop codons inside the coding sequence, was used instead of the activator δ. The output of LAG leads to the activation of the T7 promoter (Fig. 2b). To characterize each AND gate, salicylate (PSal) and arabinose (PBAD) were chosen as inputs, and the output promoter of each AND gate was transcriptionally fused to gfp. RAG showed at least a 31-fold induction (Fig. 2a, Supplementary Fig. 9), but LAG failed. When a single inducer (only arabinose or only salicylate) is applied, the output of LAG should be ‘OFF’, but the corresponding results are apparently more like ‘ON’ (Supplementary Fig. 10). To correct the behaviour of LAG, we exploited a previously developed mathematical model3 to describe the AND gate and parameterized it using the experimental data. The transfer functions of two AND gates are similar because they have the same design framework. Parameter sensitivity analysis revealed that the most sensitive parameter for the LAG is the maximum output of the PSal promoter (Supplementary Fig. 11). Next, eight ribosome-binding site (RBS) sequences preceding T7ptag were predicted to be of different strengths, with 187-fold variation28, and were designed and screened accordingly (Supplementary Methods). A mutant RBS (BBa_B0033) was identified: this mutant decreased the background and increased the dynamic range of LAG to >36-fold (Fig. 2b, Supplementary Figs 12 and 13). In the overall single simultaneous conditioning circuit (Fig. 1d), both PSal and PO drive gfp expression, directly forming the OR gate that wires the conditioned response (the output produced by the RAG) and the unconditioned response (the output produced by Food) together.
(a,b) The architectures and experimentally characterized transfer functions of the Recalling AND gate (a) and Learning AND gate (b). The inducers used were arabinose (10−3, 10−4, 10−5, 10−6, 10−7, 10−8 and 10−9 M) and salicylate (10−4, 10−5, 10−6, 10−7, 10−8, 10−9 and 10−10 M). Data were normalized by dividing the fluorescence values by the minimum fluorescence with both inputs OFF. Ara, arabinose; Sal, salicylate. n=3. (c) The architecture of the GTS and its ability to maintain either of the two distinct stable states (Memory-ON and Memory-OFF) in the long term. Serial dilution was performed on Memory-ON and Memory-OFF cells: every 12 h, cell cultures were used to inoculate the next round and the bistable distribution of the cell population was recorded using flow cytometry. Measurements are from four independent experiments and the error bars represent the s.d. Inset: the toggle switch is composed of the cI and cI434 repressors, which are mutually repressed; a positive feedback on promoter PRM was designed to topologically reinforce the bistability. (d) The switchability of the Memory module. Switchability was characterized by the percentage of cells that were Memory-ON, as assayed using flow cytometry. The cells were treated with a gradient concentration of salicylate for 2 h and then relaxed overnight without inducers. Promoter activity was calculated from the inducer concentration through a mathematical model combined with promoter characterization. Inset: the genetic circuit utilized for serial dilution. Error bars represent s.d.; n=3.
A genetic toggle switch9 (GTS), together with T7-controlled cI, was exploited as the Memory module (Fig. 2c, inset; Supplementary Fig. 14). We defined the cI-dominant state (rfp as an indicator) as a Memory-ON state, and the cI434-dominant state (gfp as an indicator) as the Memory-OFF state. In addition to bistability, there are two more indispensable functional characteristics for the GTS to work as Memory: stability, the ability of GTS to maintain the Memory-ON and Memory-OFF states for a long duration, and switchability, the ability of GTS to flip from Memory-OFF to Memory-ON when triggered by a compatible signal. To better describe the dynamics of Memory flipping, the previous ordinary differential equations model9 is adapted to a stochastic model based on the Gillespie algorithm29. To estimate stability, serial dilution experiments were performed on cells that harboured GTS, starting from either the Memory-ON or Memory-OFF state (Methods). Both the Memory-ON and Memory-OFF states were maintained for a considerably long time. The mean percentage of Memory-ON cells remained >99% for 5.5 days, and no spontaneous Memory-OFF-to-Memory-ON population transition was observed even after a week (Fig. 2c; see Supplementary Fig. 15 and Supplementary Table 8 for the raw data). To validate the switchability, an additional genetic circuit was constructed (Fig. 2d, inset; Supplementary Fig. 16). By applying gradient promoter activities (PSal) to drive the expression of cI, increasing fractions of cells were accordingly triggered to flip, from none to all (Fig. 2d; Supplementary Fig. 17). Therefore, in the overall circuit shown in Figure 1d, the Memory-ON state can be achieved in two ways: state self-sustainment or the stimulus of PT7-controlled cI expression.
Subfunction-based and model-guided module assembly
After defining and characterizing individual function modules, we managed to assemble them together. There are different approaches to assembling these modules. Our method of assembly is specified by the subfunction partitioning of the single simultaneous conditioning: the Learning device (LD) that performs the Learning subfunction (the input/Memory function described in Fig. 1b) and the conditioned response device (CRD) that performs the Recalling subfunction (the input/output function described in Fig. 1a; Supplementary Fig. 3). In the first stage of assembly, LAG coupled with Memory makes the LD, and Memory and RAG form a sub-circuit for Memory reading-out, which is then incorporated with PSal-gfp to form the CRD (Fig. 3, insets; Supplementary Fig. 18). In the second stage, we combine LD with CRD to produce the overall circuit (Supplementary Fig. 18).
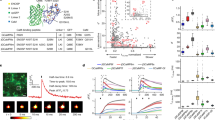
(a) The LD to perform the input/Memory function. Different association degrees of Sal (Food) and Ara (Ring) cause corresponding fractions of Memory-ON cells in a single simultaneous exposure. The flipping fraction tends to decrease when combinations of lower salicylate and/or arabinose concentrations were applied. The exposure induction was saturated (overnight) to maximize the Memory flipping. The inducers were arabinose (Ara; 10−3, 10−4, 10−5 and 10−6 M) and salicylate (Sal; 10−4, 10−5, 10−6 and 10−7 M). The scale bar shows the fraction of Memory-ON cells. (b) The dynamics of the LD under periodic ‘training’ (induction-relaxation treatment). For each session of training, cells were exposed to salicylate and arabinose (Ara+Sal) for 2 h; then, inducers were washed away and the cells were relaxed by at least 10-hour growth in fresh medium. The percentage of Memory-ON cells was assayed after overnight growth. As controls, ‘training’ using arabinose only (Ara), salicylate only (Sal) and water only were also performed. The concentrations of salicylate and arabinose were 10−4 M and 10−3 M, respectively. (c) Memory reading-out performed by the sub-circuit comprising GTS and RAG. Arabinose (Ara) triggers GFP expression if and only if Memory is ON; on the other hand, salicylate (Sal) cannot evoke output regardless of the Memory state. Inset: simplified diagram for the Memory reading-out sub-circuit. (d) CRD to wire outputs of RAG (conditioned response) and PSal-gfp (unconditioned response) together. Salicylate (Sal) evokes the GFP expression regardless of the Memory state, while arabinose (Ara)-induced GFP expression is Memory-dependent. Inset: simplified diagram for the CRD. The inducers in (c) and (d) were arabinose (10−3, 10−4, 10−5, 10−6, 10−7, 10−8 and 10−9 M) and salicylate (10−4, 10−5, 10−6, 10−7, 10−8, 10−9 and 10−10 M). Cells were assayed using flow cytometry in all the above experiments. Measurements are from three independent experiments. Error bars represent s.d.
Utilizing the characterization data of inducible promoters and individual modules, we mathematically simulated the module interconnection and conducted a parameter sensitivity analysis on the related parameters because the fine-tuning of a sensitive parameter(s) is expected to provide an effective result (Supplementary Information, Section IV.F–H). In the analysis, target functions that quantify the performance of modules/sub-circuits were first defined; their values were then calculated using different parameter sets. For each set, the value of a specific parameter moderately varied, while the values of the other parameters remained fixed. The sensitive parameters were computationally determined accordingly; when these parameters moderately varied their values, the outputs of the target functions changed considerably.
The simulation results revealed that appropriate coupling settings between specific modules might be the key to successful circuit assembly. For the LD, the dynamic range of the LAG is the key factor (Supplementary Methods). For the second stage assembly to produce an overall conditioning circuit, the plasmid copy number ratio of PSal-gfp to the LAG is critical: a small ratio guarantees a high signal-to-background performance of the LAG (Supplementary Methods).
Subsequently, module assembly was experimentally conducted. A version of GTS with gfp removed was used because gfp had been designed to be the overall output of the single simultaneous conditioning circuit. Therefore, the Memory state is solely indicated by RFP expression (that is, cells that carry the gfp-deleted GTS in the Memory-ON state produce RFP; however, those in the Memory-OFF state do not) (Supplementary Fig. 19). We expected that combining the modified GTS with LAG would make the LD (Fig. 3a, inset; see Supplementary Fig. 20 for the detailed genetic implementation). As the simulation revealed (Supplementary Methods), the dynamic range of LAG output must match that of GTS input to make a functional LD: either a high basal background or a low induction fold of PT7-controlled cI expression can disrupt the Learning function of the LD (Supplementary Fig. 21). To set an appropriate coupling between LAG and GTS, a saturated mutagenesis library was designed at the RBS of T7-controlled cI and screened for a mutant of LD in which GTS would be turned on by LAG if and only if both salicylate (Food) and arabinose (Ring) were simultaneously presented (Supplementary Fig. 22). In the mutant identified, 98% of the cell population flipped from Memory-OFF to Memory-ON when the mutant was stimulated by a simultaneous saturated exposure to salicylate and arabinose (Fig. 3a), which is consistent with the expectation for the single simultaneous conditioning. Besides, the flipping fractions decreased with weaker association degrees between the two inducers (Fig. 3a; Supplementary Fig. 23). We next examined the Learning dynamics of this intermediate circuit because the coarse Boolean-logic design cannot predict the real-time dynamics. Rather than saturated exposure induction, induction-relaxation treatment (referred to as ‘training’) was periodically applied to the cells harbouring the LD. Each session of training is a 2-hour pulse induction using both salicylate and arabinose, followed by relaxation without inducers. Results shows that the fraction of Memory-ON cells significantly increased along with the number of training sessions (Fig. 3b, Ara+Sal; Supplementary Fig. 24). Following the same protocol, pulse inductions using only salicylate, arabinose or water as the control, as expected, triggered Memory flipping much less effectively (Fig. 3b, Ara, Sal and Blank; Supplementary Fig. 24). We also expected that induction using longer pulses would potentiate the Memory flipping in the population, whereas induction with shorter pulses would weaken this process. These expected scenarios were confirmed by periodic induction-relaxation treatment using 4-hour and 1-hour pulses, respectively (Supplementary Fig. 25). Together, these results validate the proper functioning of the genetic implementation of the Learning subfunction (the input/Memory function described in Fig. 1b). More importantly, we demonstrate that, when incorporated into a genetic circuit, the digital-like Memory device exhibited dynamically progressive Learning on the population level, probably due to the stochasticity in gene expression30. This characteristic of Learning subfunction will make the single simultaneous conditioning circuit behave in a Pavlovian-like conditioning manner.
To construct the CRD, a sub-circuit was first built by integrating GTS and RAG (see Supplementary Fig. 26 for the detailed circuit design) to test whether the state of Memory could be read out: in the Memory-ON state, arabinose (Ring) is expected to trigger the output of RAG, whereas in the Memory-OFF state, the output cannot be triggered at RAG (Fig. 3c). Flow cytometry measurements of cells that harbour this sub-circuit proved that only the cells that express RFP (that is, cells in the Memory-ON state) could be triggered by arabinose to express GFP, which confirms the expectation that the Memory-ON state and arabinose (Ring) stimulus are both necessary to evoke the output (Fig. 3c; Supplementary Fig. 27). Salicylate (Food) cannot trigger any output regardless of the Memory state (Fig. 3c, inset), which proves that the conditioned response is Ring-specific. Then, PSal-gfp was incorporated with the Memory reading-out sub-circuit to produce the CRD (Fig. 3d; Supplementary Fig. 28). This arrangement allows the conditioned response (elicited by Memory and Ring at RAG) and the unconditioned response (directly elicited by Food at PSal-gfp) to be wired together. Cells harbouring the CRD exhibited both Memory-dependent, arabinose-induced and Memory-independent, salicylate-induced GFP expressions (Fig. 3d; Supplementary Fig. 29), which is consistent with the input/output function described in Fig. 1a.
Last, CRD was incorporated with LD to create the overall Pavlovian-like conditioning circuit. Two biomolecular species serve as the interconnections between CRD and LD: the regulatory protein NahR as the salicylate signal receiver (shared by the LAG and PSal-gfp) and supD tRNA as the arabinose signal carrier (shared by the LAG and RAG) (Fig. 1d). Competition for NahR and supD tRNA molecules will thus occur in the conditioning circuit (Supplementary Methods). Computational analysis revealed that the effect of competition for supD tRNA was negligible due to the nature of having a high abundance of tRNA molecules within a cell31; strong competition of PSal-gfp with LAG for NahR protein, however, would attenuate PSal in LAG (Supplementary Fig. 30). The latter prediction was experimentally verified by building two additional genetic circuits that differed in the plasmid–copy-number ratio of PSal-gfp to LAG (Supplementary Figs 31,32). Therefore, for the genetic implementation, PSal-gfp and LAG were placed together on a low-copy-number plasmid backbone (pSB4K5, ~3–5 copies per cell) to minimize the competition for the NahR protein.
Pavlovian-like conditioning function of the overall circuit
To examine the dynamics of the Learning subfunction, ‘training’ was utilized to periodically condition the Memory-OFF cell population: in every session of training, cells were simultaneously exposed to both arabinose (Ring) and salicylate (Food) for 2 h, followed by relaxation through overnight (10-hour) growth without inducers (Fig. 4a). The states of cellular Memory (RFP) and output (GFP) were recorded using flow cytometry for each step of each training session. Results showed that the fraction of Memory-ON cells within the population progressively increased along with the training pulses, until nearly the entire population was Memory-ON (Fig. 4a; see Supplementary Fig. 33 for raw cytometry data and Supplementary Fig. 34 for the mean RFP measurement). In every training session, the significant growth of Memory-ON fraction occurred after the 10-hour relaxation rather than immediately after the 2-hour pulse exposure to inducers. This was probably because that the Memory device (the GTS) was on a high-copy-number plasmid, which caused a low rate of Memory flipping (Supplementary Discussion). After each training session (induction-relaxation), the input/output function of the conditioned cell population was examined by exposure to all four inducer sets (Blank, Ara, Sal and Ara+Sal, respectively; Fig. 4b). When the entire population was Memory-OFF, the GFP expression could be elicited by salicylate (Food) but not by arabinose (Ring) (Fig. 4b, Session 0; see Supplementary Fig. 35 for raw cytometry data). As the fraction of Memory-ON individuals grew, the cell population adapted to express GFP in response to the Ring stimulus (Fig. 4b; see Supplementary Fig. 36 for raw cytometry data). As control experiments, however, periodic ‘control training’ using arabinose (Ring) only, salicylate (Food) only, or water (Blank), could not produce a significant fraction of Memory-ON cells (Fig. 4a; Supplementary Fig. 37). Correspondingly, no significant adaptive change in the input/output function was observed for the control training (Fig. 4b; Supplementary Fig. 38). These results confirmed the expectation that, in the cell population harbouring the Pavlovian-like conditioning circuit, the conditioning of input/output function is the result of the association between the Ring and Food stimuli.
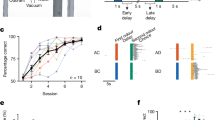
The cell population was progressively conditioned by the ‘training’ with the simultaneous Ring (Ara)+Food (Sal) stimuli but not by the ‘control training’ with Ring (Ara) only. (a) The population-level, Pavlovian-like Learning subfunction of the single simultaneous conditioning circuit. The fraction of Memory-ON cells grew with the sessions of simultaneous conditioning; the ‘control training’ with Ring only, however, produced negligible Memory-ON cells. The 2-hour induction pulse with Food+Ring or Ring only in each session has been shaded in grey; the relaxation of cell populations was performed by washing away the inducer(s) and a 10-hour growth in fresh medium. The line segments in the top indicate the two sessions for fluorescence microscopy. (b) The input/output function of a cell population could be conditioned by the simultaneous training with Food+Ring, rather than the ‘control training’ with Ring. After the induction pulse in each session, the input/output function was examined by exposing the two populations (‘Food+Ring’ training and ‘Ring-only’ control) to the all four inducer sets (Ara+Sal, Ara, Sal and Blank, respectively). The Ara (Ring) progressively gained the ability to evoke GFP expression through simultaneous conditioning. Ara, arabinose in 10−3 M; Sal, salicylate in 10−4 M; Blank, water as a control. The proportions of Memory-ON cells were assayed using cytometry data. Measurements of GFP fluorescence values are based on geometric means. The error bars show s.d. calculated on the basis of three independent replicates.
To elucidate the conditioning process in single cells, fluorescence microscopy and flow cytometry were utilized to record the states of Memory (RFP) and output (GFP) in individual cells. Two periods of ‘training’ (with Food+Ring) and ‘control training’ (with Ring only) were selected (indicated by the line segments in Fig. 4a; Fig. 5 and Supplementary Fig. 39). Before the induction pulse of Session 1, no output (GFP) or Memory (RFP) was observed in the Memory-OFF population (Fig. 5b,c, ‘Memory-OFF’). In the population undergoing the ‘control training’ with Ring only, very few cells flipped Memory-ON after the relaxation (Fig. 5b). Meanwhile, after the 2-hour induction pulse with ‘Food+Ring’, cells exhibited high-level GFP expression, although the Memory remained OFF (Fig. 5c, ‘1st Induction Pulse’). The inducers were then washed away and the cells were relaxed to allow the output to fall back to background (Fig. 5c, ‘Relaxation after 1st pulse’). As expected, after the relaxation a large fraction of cells appeared as Memory-ON in the ‘Food+Ring’-conditioned population; this is also in consistense with the observation in Fig. 4a that the Memory would not flip immediately after the induction pulse but after the relaxation. Next, the two populations were exposed to all four inducer sets (Ring, Food, Ring+Food and Blank, respectively) to examine the input/output function of individual cells (Fig. 5b,c, lower panels). Results showed that the cells in both populations exhibited the expected input/output function: a considerably large fraction of cells in the ‘Food+Ring’ population gained the new ability to respond to the Ring stimulus through registering the association of stimuli in the Memory (Fig. 5c; lower panel); the ‘Ring-only’ population, however, was still predominated by the Memory-OFF cells that could not sense the Ring (Fig. 5b; lower panel). In Session 3, a similar trend of changes in the single-cell Memory and output was observed (Supplementary Fig. 39). These results demonstrate that the progressive Pavlovian-like conditioning on the population level could be interpreted as the population dynamics of simultaneous conditioning in single cells where the storage and reading-out of Memory were digital-like. Moreover, these results, when taken together, demonstrate that the Pavlovian-like conditioning function has been successfully programmed into the bacterial cells.
Fluorescence microscopy and flow cytometry were used to record the Memory (RFP) and output (GFP) states for one typical session (Session 1, indicated by one of the line segments in Fig. 4a) and the following input/output function test. The raw cytometry data and images in bright field, GFP channel and RFP channel for each step are provided. (a) The common thresholds utilized to segregate high/low Memory (RFP) and output (GFP) states across all the cytometry data. (b) The ‘control training’ with Ring only could hardly condition the input/output function of individual cells. Upper panel: the Ring-only induction pulse caused no output during the ‘control training’ (‘1st Induction Pulse’); the Memory-ON cells remained rare after the relaxation (‘Relaxation after 1st pulse’). Lower panel: when exposed to all four inducer sets (Blank, Ara, Sal and Ara+Sal, respectively) to examine the input/output function, few cells exhibited Memory-dependent response to the Ring stimulus because the Memory-OFF cells remained predominant. (c) A fraction of cells adapted to respond to the Ring stimulus through a single-trial simultaneous conditioning. Upper panel: before the induction pulse, no Memory or output was recorded (‘Memory-OFF’); the output of all cells was turned high by the 2-hour induction pulse of simultaneous training with Ara and Sal in Session 1 (‘1st Induction Pulse’); the cells were then relaxed for 10 h to allow the GFP expression to fall back to background, after which a large fraction of cells became Memory-ON (‘Relaxation after 1st pulse’). Lower panel: the conditioned and relaxed cells were exposed to all four inducer sets (Blank, Ara, Sal and Ara+Sal, respectively). The newly conditioned Memory-ON cells showed the ability to respond to Ara (Ring) stimulus. Ara, arabinose in 10−3 M; Sal, salicylate in 10−4 M; Blank, water as a control.
Discussion
Concepts and principles in electronic circuit engineering have been speculated on providing effective guidance in the synthesis of diverse genetic logic gates and function modules32. How electronic circuit engineering can benefit the programming of complex genetic circuits for sophisticated functions in living cells, however, remains largely unexplored33. In this study, we present one of the largest transcriptional-regulator-based sequential-logic circuits developed thus far34, which comprises seven regulatory genes and 32 additional genetic parts. Our workflow of using a top–down design to decompose a complex function into subfunctions and to further divide it into individual function modules, while using bottom–up assembly to implement a genetic circuit utilizing readily available genetic parts, provides a good example of synthesizing complex genetic circuits for sophisticated functions in living cells. This study highlights the importance of engineering principles, including function decoupling, abstraction and modularization, which could reinforce our ability to manage a function’s complexity. Besides, the module-based, model-guided fine-tuning that accompanies the genetic implantation comprises two distinct stages—the optimization of individual modules and the tuning of module interconnections, which avoids ad hoc iterative tweaks.
On the other hand, in this study the single simultaneous conditioning circuit was first formulated by applying the principle of the electronic logic circuit design. In the experimental implementation of the circuit, however, Pavlovian-like conditioning function emerged on the population level, although the Memory device in single cells can store only 1-bit information. This ensemble behaviour, which cannot be predicted by the mean-field theory, is a special feature of the biological systems where stochasticity plays an important role. This reminds us of one of the fundamental differences between the electronic and the biological circuits. The former is totally deterministic, while the latter is basically stochastic that new features can emerge from this stochasticity.
There are a number of ways to expand the cellular memory capacity in the Pavlovian-like conditioning circuit and future sequential-logic genetic circuits. For example, combining libraries of GTSs together can create a cellular memory of multiple bits. Another way would be to incorporate recombinase-based memory units20,21,35, which have been utilized to implement a genetic counter6. This approach is potentially more feasible than using purely transcriptional circuitry because recombinase-based memory allows the implementation of a history-dependent logic operation in a compact manner36.
Methods
Strains and growth conditions
E. coli strain DH5α (TransGen Biotech), Luria–Bertani (LB) medium (10 g l−1 tryptone, 5 g l−1 yeast extract and 10 g l−1 NaCl) and phosphate-buffered saline (PBS) with 2 mg ml−1 kanamycin pre-added were utilized throughout the studies. Kanamycin (25 μg ml−1), ampicillin (50 μg ml−1), tetracycline (7.5 μg ml−1) and chloramphenicol (20 μg ml−1 l−1) were supplemented as needed. The inducers were Sal (sodium salicylate) and Ara (L-arabinose); all of the chemicals were purchased from Sigma-Aldrich unless stated otherwise.
Plasmid construction
BioBrick standard assembly37 was used for plasmid construction throughout the study unless otherwise specified (Supplementary Tables 9–11).
T7ptag coding sequence was PCR-amplified from plasmid pBACr-AraT7940 and supD and nahR-PSal fragments were from plasmid pAC-SalSer914 (ref. 3) using the following primers:
T7ptag-For
5′-CCGGAATTCGCGGCCGCTTCTAGATGACCATGATTACCGTGCACTAGAATACC-3′
T7ptag-Rev
5′-AAACTGCAGCGGCCGCTACTAGTATTATGCGAACGCGAAGTCCGACTCTAAGAT-3′
supD-For
5′-CCGGAATTCGCGGCCGCTTCTAGAGCAATTCGGAGAGATGCCGGAGCGGCTGAAC-3′
supD-Rev
5′-AAACTGCAGCGGCCGCTACTAGTAAGCTTAAAAAAAATCCTTAGCTTTCGCTAA-3′
nahRpSal-For
5′-CCGGAATTCGCGGCCGCTTCTAGAGTCAATCCGTAAACAGGTCAAACATCAGTTG-3′
nahRpSal-Rev
5′-AAACTGCAGCGGCCGCTACTAGTACTATGGTACTCGTGATGGCTTTATTGATGA-3′
The GTS was PCR amplified from plasmid BtoBswitch_pZS9 using the following primers:
Bistable-For
5′-AAGAATTCGCGGCCGCTTCTAGATTATTAAGCACCGGTGGAGT-3′
Bistable-Rev
5′-ATCTGCAGCGGCCGCTACTAGTTTACTTGTACAGCTCGTCCA-3′
The gfp-coding sequence in the GTS was deleted using the following primers:
De-GFP-Rev
5′-ATACTGCAGGTCGACAGATCTTTATTATACGAATTTTACCC-3′
De-GFP-For
5′-AAGAATTCTTATTAAGCACCGGTGGAGT-3′
The PSal promoter prefixed with a 20-nt fragment of the nahR 5′-coding sequence was PCR-amplified using the following primers:
nahRpSal884
5′-GGTTCTAGAGTCCAGGTCACGCAGTTCC-3′
Universal-Rev
5′-AGCCTGCAGCGGCCGCTACTAGTA-3′
Site-directed mutagenesis of genetic parts
Site-directed mutagenesis was conducted using the MutanBEST Kit (Takara) following the manufacturer’s protocol. To create amber mutations within the φR73δ-coding sequence (BBa_I746352 from in Registry of Standard Biological Parts), we conducted site-directed mutagenesis on the original serine residues. To avoid unexpected functions of truncated φR73δ (such as a dominant negative effect), the first two serine codons in the coding sequence were chosen. The primers used were:
PhiRMut-Rev
5′-TATGCGCCTAATGACGACAGAAAGGGC-3′
PhiRMut-For
5′-CCCGCACCTAGCGGTATGTGAGTGAC-3′
For the directed evolution of the LD, a saturated mutagenesis library was designed at the RBS of T7-controlled cI using the following primers:
Mut-Lib-For
5′-TTTTGCATACTAGAGNNNgNNNAGAAATACTAGATGAG-3′
Mut-Lib-Rev
5′-AGAACAAGTAGCTTGTATTCCCTATAGTGAGTCG-3′
The parental sequence was BBa_B0034. The sequence of RBS obtained from screening was 5′-TGGGGTGAGAAA-3′.
Serial dilution to validate the stability of the GTS
The plasmid harbouring the GTS was transformed into DH5α cells. Eight single colonies growing Memory-ON or Memory-OFF cells (four colonies for each Memory state) were selected using fluorescence stereomicroscopy (Leica). The colonies were inoculated into 1 ml of LB broth and grown overnight (37 °C, 250 r.p.m.) in 96-deep-well plates. A serial dilution was performed as follows.
A 2 μl aliquot of the overnight culture was diluted into 200 μl PBS for cytometry analysis (BD LSRII flow cytometer). Meanwhile, 1:1,000 dilutions were created by inoculating 1 μl of each overnight culture into 1 ml of fresh LB broth in wells of 96-deep-well plate, and these cultures were incubated overnight (37 °C, 1,000 r.p.m., mB100-40 Thermo Shaker, AOSHENG). The cytometry-dilution-regrowth process was cycled for the duration of the experiment.
In the analysis of cytometry data, for the Memory-OFF to Memory-ON transition (starting from Memory-OFF), the fraction of cells in the Memory-ON state was directly obtained through gating for mCherry and FITC. For the Memory-ON-to-Memory-OFF transition (starting from Memory-ON), the fraction of cells in the Memory-OFF state (x) was primarily obtained through gating and then converted into the fraction of cells in the Memory-ON state through the following equation: 100%−x.
Validation of the GTS switchability
Six single colonies growing Memory-ON or Memory-OFF cells (three colonies for each Memory state) were selected using fluorescence stereomicroscopy (Leica), inoculated in LB broth at 37 °C overnight in culture tubes and then diluted 1,000-fold into fresh LB broth in beaker flasks. Once the OD600 reached 0.3–0.4 (Spectrumlab 721 N), each culture was transferred into 96-deep-well plates (1 ml culture per well), supplemented with an appropriate concentration of salicylate and incubated for additional 2 h (1,000 r.p.m., 37 °C, mB100-40 Thermo Shaker, AOSHENG). The inducers were then washed away and the cultures were diluted 1,000-fold using fresh LB broth and cultivated overnight to relax the cells. Finally, 2 μl of each cell culture was diluted into 200 μl of PBS for the cytometry recording (BD LSRII flow cytometer).
Measurement of the AND gate transfer function
Cells that harbour the appropriate plasmids were incubated in LB broth at 37 °C overnight in culture tubes and then diluted 1,000-fold into fresh LB broth in beaker flasks. Once the OD600 reached 0.3–0.4 (Spectrumlab 721 N), each culture was transferred into 96-deep-well plates (1 ml culture per well), supplemented with appropriate inducer sets and incubated for 2 h (1,000 r.p.m., 37 °C, mB100-40 Thermo Shaker, AOSHENG). Finally, 20 μl of each cell culture was diluted 10-fold into 200 μl of PBS for a microplate reader recording of the fluorescence and OD600 values (Thermo Scientific Varioskan Flash).
Measurement of the Learning subfunction
The experiment described in Fig. 3b was performed as follows.
Three colonies growing Memory-OFF cells were selected using fluorescence stereomicroscopy (Leica) and were inoculated in LB broth at 37 °C overnight in culture tubes.
The cell cultures were diluted 1,000-fold using fresh LB broth in beaker flasks and incubated at 37 °C (250 r.p.m., 37 °C). Once the OD600 reached 0.3–0.4 (Spectrumlab 721 N), each culture was transferred into 96-deep-well plates (1 ml culture per well); supplemented with appropriate inducer sets (salicylate+arabinose for the training of stimulus association; salicylate only, arabinose only or water for the ‘control training’) and incubated for additional 1, 2 or 4 h (1,000 r.p.m., 37 °C, mB100-40 Thermo Shaker, AOSHENG). The inducer(s) was then washed away and the cultures were diluted 1,000-fold into fresh LB broth and cultivated overnight to relax the cells.
A 2 μl aliquot of each overnight culture was finally diluted into 200 μl PBS for a cytometry recording (BD LSRII flow cytometer). Dilutions of 1:1,000 were then created by inoculating 1 μl of each overnight culture into 1 ml of fresh LB broth in 96-deep-well plates and incubating these cultures overnight (37 °C, 1000, r.p.m., mB100-40 Thermo Shaker, AOSHENG).
The ‘training’ (or ‘control training’) relaxation process was cycled for the duration of the experiment. Each cycle counts as one session in Fig. 3b. The experiment of single simultaneous saturated induction described in Fig. 3a followed the same protocol except that the inductions for ‘training’ or ‘control training’ were overnight, thus guaranteeing inductions to be saturated.
Measurement of the Recalling subfunction
The experiments described in Fig. 3c,d were performed as follows. Six colonies growing Memory-ON or Memory-OFF cells (three colonies for each Memory state) were selected using fluorescence stereomicroscopy (Leica) and inoculated in LB broth at 37 °C overnight in culture tubes.
The cell cultures were diluted 1,000-fold using fresh LB broth in the beaker flasks. Once the OD600 reached 0.3–0.4 (Spectrumlab 721 N), each culture was transferred into 96-deep-well plates (1 ml culture per well) and supplemented with appropriate concentrations of salicylate or arabinose. The cell cultures were incubated overnight (1,000 r.p.m., 37 °C, mB 100-40 Thermo Shaker, AOSHENG). A 2-μl aliquot of each overnight cell culture was finally diluted 100-fold into 200 μl of PBS for the cytometry analysis (BD LSRII flow cytometer).
Measurement of the Pavlovian-like conditioning function
The experiments described in Figs 4 and 5 were performed as follows. Three colonies growing cells harbouring the Pavlovian-like conditioning circuit in Memory-OFF state were selected using fluorescence stereomicroscopy (Leica) and were inoculated in LB broth at 37 °C overnight in culture tubes. Through a 2-μl aliquot of each culture diluted into 200 μl PBS, the distribution of Memory states within cell populations was examined using flow cytometry (BD LSRII flow cytometer) and fluorescence microscopy (Nikon TE2000E Inverted Microscope). The cell cultures were then diluted 1,000-fold using fresh LB broth in beaker flasks and incubated at 37 °C (250 r.p.m., 37 °C).
Once the OD600 reached 0.3–0.4 (Spectrumlab 721 N), each culture was transferred into 96-deep-well plates (1 ml culture per well), supplemented with appropriate inducer sets (salicylate+arabinose for the simultaneous conditioning; salicylate only, arabinose only, or water for the ‘control training’), and incubated for an additional 2 h (1,000 r.p.m., 37 °C, mB100-40 Thermo Shaker, AOSHENG). After the 2-hour pulse induction, a 2 μl aliquot of each culture was diluted into 200 μl PBS for cytometry (BD LSRII flow cytometer) and microscopy recording (Nikon TE2000E Inverted Microscope) as needed.
The inducer(s) was then washed away and the cultures were diluted 1,000-fold using fresh LB broth and cultivated overnight (10 h at least) to relax the cells. A 2-μl aliquot of each overnight culture was diluted into 200 μl PBS for a cytometry recording (BD LSRII flow cytometer) and microscopy recording (Nikon TE2000E Inverted Microscope) as needed. Next, 1:1,000 dilutions were created by inoculating 1 μl of each overnight-relaxed cell culture into 1 ml of fresh LB broth in 96-deep-well plates, and further incubated (37 °C, 1,000 r.p.m., mB100-40 Thermo Shaker, AOSHENG).
The ‘training’ (or ‘control training’)-relaxation process was cycled for the duration of the experiment. Each cycle (induction-relaxation) counts as one session in Fig. 4. After the relaxation in each session, the cell cultures were diluted 1,000-fold using fresh LB broth in the beaker flasks. Once the OD600 reached 0.3–0.4 (Spectrumlab 721 N), each culture was transferred into 96-deep-well plates (1 ml culture per well) and supplemented with appropriate inducer sets (arabinose+salicylate, arabinose only, salicylate only and water) to examine the input–output function of the conditioned but relaxed cell population. The cell cultures were incubated for 2 h (1,000 r.p.m., 37 °C, mB 100-40 Thermo Shaker, AOSHENG), followed by 2 μl aliquot of each overnight cell culture diluted 100-fold into 200 μl of PBS for the cytometry recording (BD LSRII flow cytometer) and microscopy recording (Nikon TE2000E Inverted Microscope) as needed.
Fluorescence microscopy and image analysis
The expression of fluorescence proteins in E. coli colonies was examined using a Leica S8APO stereomicroscope (Leica Kubler CODIX fluorescence illuminator), and photographs were taken using a Leica DFC420C camera (Leica Application Suite V3 software, Leica). The GFP channel (excitation wavelength 470 nm) and RFP channel (excitation wavelength 580 nm) signals were recorded with two different sets of camera settings. Fluorescence and bright-field microscopies were performed on Nikon Eclipse Ti inverted microscope using a × 60 PlanApo total internal reflection fluorescence, 1.49 numerical aperture objective, an Evolve electron-multiplying charge-coupled device camera (Photometrics) and NIS-Elements Advanced Research software. Images were segmented and processed using Adobe Photoshop CS5 software (Adobe) and ImageJ.
Flow cytometry analysis
All data were recorded with appropriate voltage settings, gated by forward and side scatter, and contained at least 10,000 cells. Geometric mean fluorescence values were calculated using FlowJo (v7.6). The auto-fluorescence value of E. coli DH5α cells harbouring no plasmid was subtracted before reporting the fluorescence values.
Translation strength prediction
The translation strength of RBS sequences was predicted using the ‘Reverse Engineer RBS’ function of the online RBS calculator v1.128: ( https://salis.psu.edu/software/reverse). RBS translation strength output from the RBS calculation was rounded up to an integral value.
Additional information
How to cite this article: Zhang, H. et al. Programming a Pavlovian-like conditioning circuit in Escherichia coli. Nat. Commun. 5:3102 doi: 10.1038/ncomms4102 (2014).
References
Clancy, K. & Voigt, C. A. Programming cells: towards an automated 'Genetic Compiler'. Curr. Opin. Biotechnol. 21, 572–581 (2010).
Slusarczyk, A. L., Lin, A. & Weiss, R. Foundations for the design and implementation of synthetic genetic circuits. Nat. Rev. Genet. 13, 406–420 (2012).
Anderson, J. C., Voigt, C. A. & Arkin, A. P. Environmental signal integration by a modular AND gate. Mol. Syst. Biol. 3, 133 (2007).
Moon, T. S., Lou, C., Tamsir, A., Stanton, B. C. & Voigt, C. A. Genetic programs constructed from layered logic gates in single cells. Nature 491, 249–253 (2012).
Lucks, J. B., Qi, L., Mutalik, V. K., Wang, D. & Arkin, A. P. Versatile RNA-sensing transcriptional regulators for engineering genetic networks. Proc. Natl Acad. Sci. USA 108, 8617–8622 (2011).
Friedland, A. E. et al. Synthetic gene networks that count. Science 324, 1199–1202 (2009).
Liu, C. et al. Sequential establishment of stripe patterns in an expanding cell population. Science 334, 238–241 (2011).
Tabor, J. J. et al. A synthetic genetic edge detection program. Cell 137, 1272–1281 (2009).
Lou, C. et al. Synthesizing a novel genetic sequential logic circuit: a push-on push-off switch. Mol. Syst. Biol. 6, 350 (2010).
Daniel, R., Rubens, J. R., Sarpeshkar, R. & Lu, T. K. Synthetic analog computation in living cells. Nature 497, 619–623 (2013).
Chau, A. H., Walter, J. M., Gerardin, J., Tang, C. & Lim, W. A. Designing synthetic regulatory networks capable of self-organizing cell polarization. Cell 151, 320–332 (2012).
Elowitz, M. B. & Leibler, S. A synthetic oscillatory network of transcriptional regulators. Nature 403, 335–338 (2000).
Tigges, M., Marquez-Lago, T. T., Stelling, J. & Fussenegger, M. A tunable synthetic mammalian oscillator. Nature 457, 309–312 (2009).
Danino, T., Mondragon-Palomino, O., Tsimring, L. & Hasty, J. A synchronized quorum of genetic clocks. Nature 463, 326–330 (2010).
Lu, T. K. & Collins, J. J. Engineered bacteriophage targeting gene networks as adjuvants for antibiotic therapy. Proc. Natl Acad. Sci. USA. 106, 4629–4634 (2009).
Ye, H., Daoud-El Baba, M., Peng, R. W. & Fussenegger, M. A synthetic optogenetic transcription device enhances blood-glucose homeostasis in mice. Science 332, 1565–1568 (2011).
Xie, Z., Wroblewska, L., Prochazka, L., Weiss, R. & Benenson, Y. Multi-input RNAi-based logic circuit for identification of specific cancer cells. Science 333, 1307–1311 (2011).
Zhang, F., Carothers, J. M. & Keasling, J. D. Design of a dynamic sensor-regulator system for production of chemicals and fuels derived from fatty acids. Nat. Biotechnol. 30, 354–359 (2012).
Lu, T. K., Khalil, A. S. & Collins, J. J. Next-generation synthetic gene networks. Nat. Biotechnol. 27, 1139–1150 (2009).
Bonnet, J., Yin, P., Ortiz, M. E., Subsoontorn, P. & Endy, D. Amplifying genetic logic gates. Science 340, 599–603 (2013).
Siuti, P., Yazbek, J. & Lu, T. K. Synthetic circuits integrating logic and memory in living cells. Nat. Biotechnol. 31, 448–452 (2013).
Tagkopoulos, I., Liu, Y. C. & Tavazoie, S. Predictive behavior within microbial genetic networks. Science 320, 1313–1317 (2008).
Mitchell, A. et al. Adaptive prediction of environmental changes by microorganisms. Nature 460, 220–224 (2009).
Pavlov, I. P. & Anrep, G. V. Conditioned reflexes; An Investigation of the Physiological Activity of the Cerebral Cortex Oxford University Press: Humphrey Milford (1927).
Fritz, G., Buchler, N. E., Hwa, T. & Gerland, U. Designing sequential transcription logic: a simple genetic circuit for conditional memory. Systems Synthet Biol 1, 89–98 (2007).
Vingron, S. P. Logic Circuit Design: Selected Methods Springer (2012).
Julien, B. & Calendar, R. Bacteriophage PSP3 and phiR73 activator proteins: analysis of promoter specificities. J. Bacteriol. 178, 5668–5675 (1996).
Salis, H. M., Mirsky, E. A. & Voigt, C. A. Automated design of synthetic ribosome binding sites to control protein expression. Nat. Biotechnol. 27, 946–950 (2009).
Gillespie, D. T. Exact stochastic simulation of coupled chemical-reactions. J. Phys. Chem. 81, 2340–2361 (1977).
Eldar, A. & Elowitz, M. B. Functional roles for noise in genetic circuits. Nature 467, 167–173 (2010).
Deutscher, M. P. Degradation of RNA in bacteria: comparison of mRNA and stable RNA. Nucleic Acids Res. 34, 659–666 (2006).
Miyamoto, T., Razavi, S., DeRose, R. & Inoue, T. Synthesizing biomolecule-based Boolean logic gates. ACS Synth. Biol. 2, 72–82 (2013).
Kittleson, J. T., Wu, G. C. & Anderson, J. C. Successes and failures in modular genetic engineering. Curr. Opin. Chem. Biol. 16, 329–336 (2012).
Purnick, P. E. & Weiss, R. The second wave of synthetic biology: from modules to systems. Nat. Rev. Mol. Cell Biol. 10, 410–422 (2009).
Bonnet, J., Subsoontorn, P. & Endy, D. Rewritable digital data storage in live cells via engineered control of recombination directionality. Proc. Natl Acad. Sci. U. S. A. 109, 8884–8889 (2012).
Benenson, Y. Engineering. Recombinatorial logic. Science 340, 554–555 (2013).
Canton, B., Labno, A. & Endy, D. Refinement and standardization of synthetic biological parts and devices. Nat. Biotechnol. 26, 787–793 (2008).
Acknowledgements
We would like to thank C. Lou and Y.P. Wang for their discussions; L. Jiang, W. Wei, C. Xu, L. Liu, Y. Qu and L. Ji for their technical assistance; the BioBrick Foundation for providing BioBricks; H.M. Salis for providing the online RBS Calculator v1.1 and C.A. Voigt for supplying plasmids pAC-SalSer914 and pBACr-AraT7940. This work was supported by the Teaching Centre for Experimental Biology, the National Undergraduate Innovation Experiment Program, the NSF of China (11074009 and 10721463), the NFFTBS of China (J1030623, J1103505, J1030310 and J1103205) and the MOST of China (2009CB918500 and 2012AA02A702).
Author information
Authors and Affiliations
Contributions
H.Z. and M.L. designed and performed the experiments, analysed the data, developed the computational models and wrote the manuscript. H.S., X.Z. and C.T. analysed the data and developed the computational models. L.H. designed the experiments. S.S., R.G., S.W., Z.Y., G.Z., S.H., H.W., T.S. and Y.C. performed the experiments. W.J. analysed the data and wrote the manuscript. Q.O. designed the experiments, developed the computational models and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Figures 1-39, Supplementary Tables 1-11, Supplementary Discussion, Supplementary Methods and Supplementary References (PDF 6261 kb)
Rights and permissions
About this article
Cite this article
Zhang, H., Lin, M., Shi, H. et al. Programming a Pavlovian-like conditioning circuit in Escherichia coli. Nat Commun 5, 3102 (2014). https://doi.org/10.1038/ncomms4102
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/ncomms4102
This article is cited by
-
Programmable responsive hydrogels inspired by classical conditioning algorithm
Nature Communications (2019)
-
Computing with biological switches and clocks
Natural Computing (2018)
-
Layering genetic circuits to build a single cell, bacterial half adder
BMC Biology (2015)
-
Optogenetic characterization methods overcome key challenges in synthetic and systems biology
Nature Chemical Biology (2014)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.