Abstract
Twenty-eight spontaneous alterations modifying the hybridization banding pattern of six families of transposable elements (297, Foldback, copia, jockey, P and hobo) have been fixed in a set of mutation-accumulation lines of Drosophila melanogaster. Their effect on fitness has been studied by competition with the original pattern. Most alterations affecting transposable elements were shown to be rearrangements with no detectable effect on fitness, showing that spontaneous transposable element mutations mainly generate minor fitness mutations.
Similar content being viewed by others
Introduction
Transposable elements (TEs) are a common feature of eukaryotic genomes and can comprise a substantial fraction of genomic DNA, but their evolutionary significance is not well understood. Transposition is known to be mutagenic, as the insertion of elements may disrupt a structural locus or alter its regulation, and chromosomal rearrangements may be generated by ectopic exchange between nonhomologous TEs. The population dynamics of TEs is one of the main problems of population genetics because of their importance for genomic diversity. The relative contribution of selective forces to the maintenance of TEs in natural populations depends on the rate at which rearrangements affecting TEs (new insertions and excisions, recombination between elements, deletions and insertions within elements) occur and the consequences of such events for the genome of the host.
Some theoretical models dealing with the consequences on viability of changes affecting TEs have been elaborated (for a discussion see Biémont et al., 1997; Brookfield & Badge, 1997; Charlesworth et al., 1997), but empirical determinations of the significance of the different parameters are partial and scarce. Direct estimations of variation affecting Drosophila melanogaster TEs have been made for the euchromatic section of chromosome X, II or the three major chromosomes (by, respectively, Eggleston et al., 1988; Harada et al., 1990; Nuzhdin & Mackay, 1994, 1995), by in situ hybridization, and for the whole genome using the Southern blot method (Di Franco et al., 1992; Domínguez & Albornoz, 1996). Rates of movement vary widely depending on the elements tested and the stock analysed. Direct determinations of the effects of TEs on fitness in D. melanogaster are limited to the effects of hybrid dysgenesis (Mackay, 1986; Lyman et al., 1996) and show that new P-element insertions predominantly induce lethal or detrimental mutations.
In this paper, we study the nature, and effects on fitness, of spontaneous changes affecting six TE families (297, Foldback, copia, jockey, P and hobo) that have been fixed in a set of mutation-accumulation lines of D. melanogaster derived from an isogenic population. The homogeneous origin of the lines allowed us to estimate the effects that TE mutations have on fitness by competition between the original and the modified insertion patterns. Most of the observed hybridization pattern changes were simultaneous band gains and losses that were consistent with rearrangements (Domínguez & Albornoz, 1996). The study of the segregation of the different hybridization patterns allowed us to confirm this hypothesis.
Materials and methods
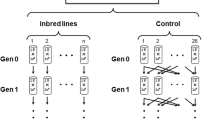
Lines were obtained from a base population which had been made isogenic for all chromosomes following a scheme of crosses with balancer chromosomes. The base population carried the recessive eye-colour marker sepia (se) as an indicator of possible contamination. Starting from the base population, two sets (B and C) of 100 inbred lines each were established (Santiago et al., 1992), either by single brother–sister mating (lines B, denoted B1–B100) or by double-first-cousin mating (lines C, denoted C1–C100). From generation 47, all C lines were also maintained by brother–sister mating. The marker phenotype was verified at each generation. Flies were reared on the standard medium formula of this laboratory (brewer’s yeast–agar–sucrose) at 24 ± 1°C. In a previous study (generation 80, Domínguez & Albornoz, 1996), lines were tested for changes in the insertion pattern of nine TE families (P, hobo, copia, gypsy, blood, Foldback, 412, 297 and jockey). At generation 120, there were 171 surviving lines. The lines that showed alterations in the TE pattern at generation 80 were re-tested by means of Southern blots from pools of flies. One line (C66) that showed the original insertion pattern was selected as a control.
The possibility that TE movements have an effect on fitness was tested by competition between the modified insertion pattern and the original one. Each line was crossed with the control to obtain the two reciprocal F1s. Two independent replicates, with a frequency of 0.5 of the altered pattern, were made. Each replicate was founded with 20 pairs, 10 males and 10 virgin females from each of the two reciprocal F1 offspring. The resultant F2 and successive descendants were kept in one 250 mL glass bottle under crowded conditions for 10 generations. The entire offspring from each generation was transferred to a fresh bottle. In order to avoid generation overlapping, parents were allowed to lay for 7 days and then discarded. After twelve generations (F12), a sample of females from each replicate was tested for their pattern of TEs by means of Southern blots from individual flies. One fly from the tested line and another from the C66 were included as controls. Frequencies of the modified hybridization pattern were compared with 0.5, the expected value if movements have no effect on fitness.
Individual adult flies were homogenized in 100 μL of extraction buffer (0.1 M NaCl; 0.1 M Tris-HCl, pH 8.4; 20 mM EDTA). Proteinase K and SDS were added to a final concentration of 100 μg mL–1 and 1%, respectively, and the mixture was incubated at 65°C for 1 h. After cooling at room temperature for a few minutes, 20 μL of potassium acetate were added. After 30 min on ice, the samples were spun for 5 min in an Eppendorf centrifuge. Ethanol (200 μL) was added to the supernatant, and after 1 h at −70°C, the samples were spun again for 15 min. The pellet was washed twice in 70% ethanol, dried under vacuum and resuspended in TE buffer (Sambrook et al., 1989). The DNA extraction from pools of flies, together with the enzymes used to digest genomic DNA and probes for hobo, copia, jockey, 297, P and Foldback (FB) transposable elements have been previously described in Domínguez & Albornoz (1996). Digested DNA was separated by electrophoresis in 30 cm–0.5% or 0.7% agarose gels and TBE buffer (Sambrook et al., 1989). Gels were run at 30 or 65 V for 48 or 24 h, respectively. DNA was transferred to a charged nylon membrane under saline conditions, following the manufacturer’s recommendations. Probe DNA was 32P-labelled by random priming and hybridized to membranes in 6 × SSC, 0.5% SDS, 100 μg mL–1 denatured herring sperm DNA overnight at 65°C. Membranes were washed twice in 2 × SSC and 0.1% SDS, once in 0.1 × SSC and 0.1% SDS at 65°C for 30 min, briefly washed in 0.1 × SSC at room temperature and autoradiographed for 1–4 days at −70°C with one intensifying screen. Filters were stripped with boiling 0.5% SDS.
Results
Observed differences from the original hybridization pattern are presented in Table 1. Every line conserved the pattern alterations detected at generation 80 and two of them have fixed new changes (lines B78 and C67; compare with Table 1 from Domínguez & Albornoz, 1996). The most frequent change was a band gain and a loss (Fig. 1). This kind of pattern alteration may be the consequence of two transposition events, one insertion plus one excision, but the same would be expected from rearrangements leading to the gain or loss of the TE genetic material. These changes were previously interpreted as rearrangements of the element on the basis of their statistical nonindependence (Domínguez & Albornoz, 1996). Analysis of individual hybridization patterns in the competition tests proved this hypothesis: band gains and losses did not segregate; all the homozygotes for a band loss also showed the band gain, as can be seen in the example of Table 2a. Thus, they were closely linked or, more probably, they were a consequence of a unique event. Rearrangements affecting the element size or their restriction map will be reflected as band shifts. Therefore, all the changes affecting more than one band that did not show segregation at the F12 were classified as rearrangements in Table 2. Most rearrangements were reflected as a band shift, but lines B78 and B96 cosegregated for two gains plus a loss for elements 297 and FB, respectively. Single band gains or losses were classified as insertions or excisions, respectively, but some additional work would be necessary to confirm this.
Three lines carried alterations of the hybridization pattern that could be interpreted as a consequence of more than one change in the same TE family. Line B78 has fixed one 297 insertion and the above-mentioned rearrangement that causes two band gains and one loss (Fig. 2b). In line C72 one FB insertion and one rearrangement (band gain and loss) could be distinguished, and line C65 bore two hobo rearrangements, both reflected as one band shift.
(a) Details of the FB hybridization pattern showing the segregation of the FB rearrangement (r) from line B42 in a sample from the F12 offspring. (b) Details of the 297 hybridization pattern showing the segregation of the 297 insertion (i) and rearrangement (r) from line B78 in a sample from the F12 offspring.
Table 2 also shows the results of competition tests (frequencies attained at the F12) for all TE movements. When only one band gain or loss was segregating, homozygotes were distinguished from heterozygotes by comparing the relative band intensities, determined with the application GEL READER from NCSA (1991). As Southern blots were made with single flies, a band in a heterozygote will be half as intense as the same band in a homozygote.
Transposable element mutations remained segregating at intermediate frequencies with a general mean of 0.48 ± 0.03, a value remarkably close to the 0.5 expected for neutrality of TE movements. The distribution (Fig. 3) was slightly negatively skewed (g1=−0.16 ± 0.44) and slightly leptokurtic (g2=1.07 ± 0.86). The variance between replicates within lines (0.0195) resulted from the joint effects of 10 generations of genetic drift and the sampling error. The variance between line mean frequencies (0.0344), that include effects of TE movements on viability, was compared with the variance within lines by an analysis of variance. The F-value was significant (F27,28=5.35, P=0.0007), denoting that some TE changes affect viability. Individual line frequencies were compared with the 0.5-value. Significance is based on Student’s t-test, after applying the Bonferroni correction (Rice, 1989). Only two FB rearrangements departed from neutrality, one with a positive effect on fitness and the other with a negative effect. Previous tests were based on the normal distribution, which may have some problems because of the small sample sizes. Thus, frequencies were also compared by the nonparametric Kruskal–Wallis test, which gave a P-value of 0.0460, again indicating that some TE changes are not neutral.
Discussion
The most frequent alterations affecting TEs (21 out of the 28 movements studied) were rearrangements that were reflected in the hybridization pattern as nonindependent band gains and losses. Only seven changes could be classified as insertion or excision events, but further additional work is necessary to confirm this, because some deletions or amplifications may also be reflected as single changes, depending on the restriction enzyme and the probe used.
For elements P and hobo, these observations could be related to transposition events, as nonreplicative transposition to a nearby site (Tower et al., 1993). Alternatively, the same pattern differences could be the result of tandem duplications or internal deletions promoted by the element itself, as a consequence of aberrant transposition (Gray et al., 1996; Preston et al., 1996). A prevalence of this kind of rearrangement rather than transposition has been found to be induced by P-element transposase (Delattre et al., 1995). The same events have been described in the phenomenon of hybrid dysgenesis associated with the hobo element, and there is also evidence that they can be promoted by the FB family (Bingham & Zachar, 1989). Hybridization pattern alterations for retrotransposons copia and 297 were not congruent with transposition-related events. These elements transpose via an RNA intermediate and, consequently, in a replicative way. Therefore, our observations are better interpreted as a consequence of internal rearrangements. Assuming that all the band shifts detected in generation 80 were a consequence of rearrangements (three lines showing this kind of change were lost), the mean rate of rearrangements that results from the data of Domínguez & Albornoz (1996) is 8.5 × 10–6 (23/2 693 600) per element per generation.
DNA loss (Petrov et al., 1996; Petrov & Hartl, 1998) may be a factor, common to all the families studied, leading to the observed hybridization pattern alterations. The reported rate of deletion was approximately 0.16 times the nucleotide substitution rate. This rate in Drosophila has been estimated as 8.5 × 10–9 per base per generation (Drake et al., 1998). The mean size of the nine families tested by Domínguez & Albornoz (1996) was 5.3 kb. Thus, the rate of deletion would be of 7.21 × 10–6, a value very close to the above-estimated rate of rearrangement. Unequal crossing-over may be another factor involved in rearrangements. Ectopic exchange between copies of retrotransposons in heterozygosis have been shown to produce rearrangements (Goldberg et al., 1983; Davis et al., 1987), but this can be excluded because the starting population was homozygous.
It has been suggested (Junakovic et al., 1997) that host factors play a significant role as regulators of the activity of TEs. Our experimental material probably reflects a stable genotype where transposition of the different TE families is repressed. There was no P or hobo hybrid dysgenesis during the construction of the base population, and the frequency of movements that could be classified as ‘true’ insertions or excisions was very low: 8.7 × 10–7 and 1.3 × 10–6, respectively (Domínguez & Albornoz, 1996). Consequently, it can be deduced that, in the absence of genomic instability, the main source of spontaneous variation for TEs will be small, probably internal, rearrangements, either promoted by the element itself or by general mechanisms (DNA loss or unequal recombination).
The competition experiment between the original and modified hybridization patterns showed that TE mutations mainly have minor fitness effects. Only two out of the 28 TE mutations analysed have effects on fitness, one negative and the other positive. These two detected effects may not be associated with the TE rearrangements per se, because either the control or the tested lines may have accumulated other mutations. The apparent effect of a particular TE mutation could be caused by a cis-linked mutation with the same effect or a trans-linked one with the opposite effect. Attention must be paid primarily to the distribution of frequencies, which was symmetrical and with a mean very close to 0.5, showing that the contribution of the overall variation of TEs to fitness was small or negligible. As TE movements are a common class of spontaneous mutation event in Drosophila (Green, 1988), our results are consistent with recent observations of small changes in fitness traits in long-term mutation-accumulation experiments using the same set of lines (Fernández & López-Fanjul, 1996, 1997) and other experiments with a similar design (Keightley & Caballero, 1997).
The distribution of spontaneous mutant effects on fitness is highly dependent on the technique used to isolate them. Mutations accumulating against a balancer chromosome are effectively sheltered from natural selection. In inbred lines, however, the fixation probability of unconditionally deleterious mutations will be small and most fixed mutations will be neutral or mildly detrimental, that is, those having a major role on population survival (Lande, 1995). Both methods result in partially overlapping descriptions: accumulation against a balancer chromosome gives a better representation of the original distribution of mutant effects, whereas the distribution obtained from inbreeding is more relevant to the genetic variation in natural populations after purging selection has acted.
References
Biémont, C., Vieira, C., Hoogland, C., Cizeron, G., Lœvenbruck, C., Arnault, C. and Carante, J. (1997). Maintenance of transposable element copy number in natural populations of Drosophila melanogaster and D. simulans. Genetica, 100: 161–166.
Bingham, P. M. and Zachar, Z. (1989). Retrotransposons and the FB transposon from Drosophila Melanogaster. In: Berg, D. and Howe, M., (eds) Mobile DNA, pp. 485–502 American Society for Microbiology, Washington, DC.
Brookfield, J. F. Y. and Badge, R. M. (1997). Population genetics models of transposable elements. Genetica, 100: 281–294.
Charlesworth, B., Langley, C. H. and Sniegowski, P. D. (1997). Transposable element distributions in Drosophila. Genetics, 147: 1993–1995.
Davis, P. S., Shen, M. V. and Judd, B. H. (1987). Asymmetrical pairings of transposons in and proximal to the white locus of Drosophila account for four classes of regulatory occurring exchange products. Proc Natl Acad Sci USA, 84: 174–178.
Delattre, M., Anxolabéhère, D. and Coen, D. (1995). Prevalence of localized rearrangements vs. transpositions among events induced by Drosophila P element transposase on a P transgene. Genetics, 141: 1407–1424.
Di Franco, C., Galuppi, C. and Junakovic, N. (1992). Genomic distribution of transposable elements among individuals of an inbred Drosophila line. Genetica, 86: 1–11.
Domínguez, A. and Albornoz, J. (1996). Rates of movement of transposable elements in Drosophila melanogaster. Mol Gen Genet, 251: 130–138.
Drake, J. W., Charlesworth, B., Charlesworth, D. and Crow, J. F. (1998). Rates of spontaneous mutation. Genetics, 148: 1667–1686.
Eggleston, W. B., Johnson-Schlitz, D. M. and Engels, W. R. (1988). P-M hybrid dysgenesis does not mobilize other transposable element families in D. melanogaster. Nature, 331: 368–370.
Fernández, J. and López-Fanjul, C. (1996). Spontaneous mutational variances and covariances for fitness-related traits in Drosophila melanogaster. Genetics, 143: 829–837.
Fernández, J. and López-Fanjul, C. (1997). Spontaneous mutational genotype–environment interaction for fitness-related traits in Drosophila melanogaster. Evolution, 51: 856–864.
Goldberg, M. L., Sheen, J. -Y., Ghering, W. J. and Green, M. M. (1983). Unequal crossing-over associated with asymmetrical synapsis between nomadic elements in the Drosophila melanogaster genome. Proc Natl Acad Sci USA, 80: 5017–5021.
Gray, Y. H. M., Tanaka, M. M. and Sved, J. A. (1996). P-element-induced recombination in Drosophila melanogaster: hybrid element insertion. Genetics, 144: 1601–1610.
Green, M. M. (1988). Mobile DNA elements and spontaneous gene mutation. Banbury Report, 30: 41–50.
Harada, K., Yukuhiro, K. and Mukai, T. (1990). Transposition rates of movable genetic elements in Drosophila melanogaster. Proc Natl Acad Sci USA, 87: 3248–3252.
Junakovic, N., Difranco, C. and Tierroni, A. (1997). Evidence for a host role in regulating the activity of transposable elements in Drosophila melanogaster: the case of the persistent instability of Bari 1 elements in Charolles stock. Genetica, 100: 149–154.
Keightley, P. D. and Caballero, A. (1997). Genomic mutation rates for lifetime reproductive output and lifespan in Caenorhabditis elegans. Proc Natl Acad Sci USA, 94: 3823–3827.
Lande, R. (1995). Mutation and conservation. Conserv Biol, 9: 782–791.
Lyman, R. F., Lawrence, F., Nuzhdin, S. V. and Mackay, T. F. C. (1996). Effects of single P-elements insertions on bristle number and viability in Drosophila melanogaster. Genetics, 143: 277–292.
Mackay, T. F. C. (1986). Transposable element-induced fitness mutations in Drosophila melanogaster. Genet Res, 48: 77–87.
NCSA (1991). NCSA GelReader for the Macintosh version 2.0.5. University of Illinois, Urbana-Champaign, IL.
Nuzhdin, S. V. and Mackay, T. F. C. (1994). Direct determination of retrotransposon transposition rates in Drosophila melanogaster. Genet Res, 63: 139–144.
Nuzhdin, S. V. and Mackay, T. F. C. (1995). The genomic rate of transposable element movement in Drosophila melanogaster. Mol Biol Evol, 12: 180–181.
Petrov, D. A. and Hartl, D. L. (1998). High rate of DNA loss in the Drosophila melanogaster and Drosophila virilis species groups. Mol Biol Evol, 15: 293–302.
Petrov, D. A., Lozovskaya, E. R. and Hartl, D. L. (1996). High intrinsic rate of DNA loss in Drosophila. Nature, 384: 346–349.
Preston, C. R., Sved, J. A. and Engels, W. R. (1996). Flanking duplications and deletions associated with P-induced male recombination in Drosophila. Genetics, 144: 1623–1638.
Rice, W. R. (1989). Analyzing tables of statistical tests. Evolution, 43: 223–225.
Sambrook, J., Fritsch, E. F. and Maniatis, T. (1989). Molecular Cloning: a Laboratory Manual, 2nd edn. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York, NY.
Santiago, E., Albornoz, J., Domínguez, A., Toro, M. A. and López-fanjul, C. (1992). The distribution of spontaneous mutations on quantitative traits and fitness in Drosophila melanogaster. Genetics, 132: 771–781.
Tower, J., Karpen, G. H., Craig, N. and Spradling, A. C. (1993). Preferential transposition of Drosophila P elements to nearby chromosomal sites. Genetics, 133: 347–359.
Acknowledgements
We thank Dr C. López-Fanjul and Dr C. Biémont for their critical reading of the manuscript and helpful comments. This work was supported by a grant from the Dirección General de Investigación Científica y Técnica (D.G.I.C.Y.T, PB 94–1345 A).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Albornoz, J., Domínguez, A. Spontaneous changes in Drosophila melanogaster transposable elements and their effects on fitness. Heredity 83, 663–670 (1999). https://doi.org/10.1046/j.1365-2540.1999.00590.x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1046/j.1365-2540.1999.00590.x
Keywords
This article is cited by
-
Genomic distribution of retrotransposons 297, 1731, copia, mdg1 and roo in the Drosophila melanogaster species subgroup
Genetica (2010)
-
Direct determination of the influence of extreme temperature on transposition and structural mutation rates of Drosophila melanogaster mobile elements
Genetica (2006)
-
Some Problems of Studies of the Genetic Bases of Speciation on the Example of Drosophila Group virilis
Russian Journal of Developmental Biology (2005)