Featured
-
-
Article
| Open Access Seasonal antigenic prediction of influenza A H3N2 using machine learning
Seasonal antigenic prediction of influenza A H3N2 using machine learning
This study presents a machine learning model that accurately predicts seasonal antigenic changes of influenza A H3N2 using genetic data. The model’s predictions can aid influenza surveillance, vaccine strain selection, and public health management.
- Syed Awais W. Shah
- , Daniel P. Palomar
- & Matthew R. McKay
-
Article
| Open Access Dynamic diversity of SARS-CoV-2 genetic mutations in a lung transplantation patient with persistent COVID-19
Dynamic diversity of SARS-CoV-2 genetic mutations in a lung transplantation patient with persistent COVID-19
In this study, the authors report the case of a patient who underwent lung transplantation and subsequently developed COVID-19 that resulted in persistent infection. Following antiviral treatment, SARS-CoV-2 (BA.5) showed dynamic genetic diversity with remdesivir resistant mutations leading to enhanced fusogenicity.
- Hidetoshi Igari
- , Seiichiro Sakao
- & Eiji Ido
-
Article
| Open Access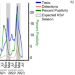 Deviations in RSV epidemiological patterns and population structures in the United States following the COVID-19 pandemic
Deviations in RSV epidemiological patterns and population structures in the United States following the COVID-19 pandemic
Non-pharmaceutical interventions for COVID-19 also impacted the transmission of other viruses including respiratory syncytial virus (RSV). Here the authors describe the changing epidemiology, clinical severity, and genetic diversity of RSV in Chicago, Illinois, from July 2010 to April 2023.
- Estefany Rios-Guzman
- , Lacy M. Simons
- & Judd F. Hultquist
-
Article
| Open Access Evolution and neutralization escape of the SARS-CoV-2 BA.2.86 subvariant
Evolution and neutralization escape of the SARS-CoV-2 BA.2.86 subvariant
The Omicron BA.2.86 subvariant differs from previous variants by over 30 spike mutations. Here, the authors report that BA.2.86 likely evolved in Southern Africa and that its immune escape is not larger than recently circulating SARS-CoV-2 strains. Neither its replication nor its pathogenicity are enhanced in vitro.
- Khadija Khan
- , Gila Lustig
- & Alex Sigal
-
Article
| Open Access Co-option of a non-retroviral endogenous viral element in planthoppers
Co-option of a non-retroviral endogenous viral element in planthoppers
Non-retroviral endogenous viral elements are widely dispersed in eukaryotic genomes, but their functions remain largely unknown. Here, Huang et al show that one such element in planthoppers has been co-opted and contributes to insect fitness..
- Hai-Jian Huang
- , Yi-Yuan Li
- & Jun-Min Li
-
Article
| Open Access Genomic adaptation of giant viruses in polar oceans
Genomic adaptation of giant viruses in polar oceans
This study examines the biogeography and functional gene repertoires of marine eukaryote-infecting large and giant DNA viruses. It shows a clear divide in the viral communities between polar and nonpolar environments, with recurrent evolutionary adaptations to the polar environment likely driven by alterations of their genomic functions.
- Lingjie Meng
- , Tom O. Delmont
- & Hiroyuki Ogata
-
Article
| Open Access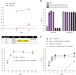 An Influenza A virus can evolve to use human ANP32E through altering polymerase dimerization
An Influenza A virus can evolve to use human ANP32E through altering polymerase dimerization
Despite their essentiality, human ANP32A and ANP32B are redundant host factors for influenza virus genome replication. In this work, authors show that an influenza virus grown in cells lacking ANP32A and ANP32B evolved to use ANP32E. They explore the polymerase mutations that enable this, and demonstrate increased virulence in mice.
- Carol M. Sheppard
- , Daniel H. Goldhill
- & Wendy S. Barclay
-
Article
| Open Access Panoramic analysis of coronaviruses carried by representative bat species in Southern China to better understand the coronavirus sphere
Panoramic analysis of coronaviruses carried by representative bat species in Southern China to better understand the coronavirus sphere
In this study, Han, Xu, and Wang et al. probe the diversity of bat coronaviruses (CoVs), revealing their evolutionary pattern with hosts. It underscores the evolutionary processes of CoVs, including SARS-CoV-2, and emphasizes the urgency of ongoing bat CoV surveillance.
- Yelin Han
- , Panpan Xu
- & Zhiqiang Wu
-
Article
| Open Access Genome mining shows that retroviruses are pervasively invading vertebrate genomes
Genome mining shows that retroviruses are pervasively invading vertebrate genomes
Ongoing retroviral invasion into vertebrates has been rarely documented. Here the authors have identified 412 endogenous retroviruses that are invading the genomes of over a hundred vertebrate species. This may be relevant to conservation of threatened species, zoonoses in the wild, and emerging infectious diseases in humans.
- Jianhua Wang
- & Guan-Zhu Han
-
Article
| Open Access Bipartite genome and structural organization of the parvovirus Acheta domesticus segmented densovirus
Bipartite genome and structural organization of the parvovirus Acheta domesticus segmented densovirus
Parvoviruses have been reported to carry a linear monopartite ssDNA genome. In this study, the authors isolated an insect-infecting parvovirus with a bipartite genome from house crickets and discovered a genome packaging strategy distinct from other parvoviruses.
- Judit J. Pénzes
- , Hanh T. Pham
- & Peter Tijssen
-
Article
| Open Access Virological characteristics of the SARS-CoV-2 XBB variant derived from recombination of two Omicron subvariants
Virological characteristics of the SARS-CoV-2 XBB variant derived from recombination of two Omicron subvariants
XBB is the first recombinant, globally dominant variant of SARS-CoV-2. Here, the authors examine the variant’s origins and virological properties, showing it is the first example of SARS-CoV-2 improving its fitness through recombination.
- Tomokazu Tamura
- , Jumpei Ito
- & Kei Sato
-
Article
| Open Access Hybrids of RNA viruses and viroid-like elements replicate in fungi
Hybrids of RNA viruses and viroid-like elements replicate in fungi
RNA viruses are defined by linear RNA genomes encoding an RNA-dependent RNA polymerase, while viroid-like elements consist of small, single-stranded, circular RNA genomes that, in some cases, encode self-cleaving catalytic RNAs. Here, the authors identify over 20,000 candidate viroid-like elements, and show that infectious agents of fungi display hybrid features of viroid-like RNAs and RNA viruses.
- Marco Forgia
- , Beatriz Navarro
- & Marcos de la Peña
-
Article
| Open Access Total escape of SARS-CoV-2 from dual monoclonal antibody therapy in an immunocompromised patient
Total escape of SARS-CoV-2 from dual monoclonal antibody therapy in an immunocompromised patient
Antibodies against SARS-CoV-2 can be used to treat infections but there is a risk of driving viral resistance to antibodies. Here the authors characterise SARS-CoV-2 escape mutants from an immunocompromised patient treated with anti-SARS-CoV-2 antibodies using mouse protection studies and structural prediction.
- Lena Jaki
- , Sebastian Weigang
- & Jonas Fuchs
-
Article
| Open Access Within-host genetic diversity of SARS-CoV-2 lineages in unvaccinated and vaccinated individuals
Within-host genetic diversity of SARS-CoV-2 lineages in unvaccinated and vaccinated individuals
There is limited data on within-host SARS-CoV-2 genetic diversity and how it is affected by vaccination. The authors analysed intra-host sequence diversity and found that VOCs may have more sequence variations than non-VOCs and that breakthrough infections in vaccinated individuals do not seem to increase non-silent mutations.
- Haogao Gu
- , Ahmed Abdul Quadeer
- & Leo L. M. Poon
-
Article
| Open Access Evolution of giant pandoravirus revealed by CRISPR/Cas9
Evolution of giant pandoravirus revealed by CRISPR/Cas9
Until today, genetic tools have been lacking to enable manipulation of amoebal giant viruses (GVs) by CRISPR/Cas9 technology. Here, Bisio et al. apply S. pyogenes Cas9 together with pU6- driven guide RNAs to investigate the replication of pandoravirus, a GV replication in the nucleus. Using this tool, they provide evidence for stepwise evolution and genetic expansion of viral gigantism.
- Hugo Bisio
- , Matthieu Legendre
- & Chantal Abergel
-
Article
| Open Access Cryo-EM structure of ssDNA bacteriophage ΦCjT23 provides insight into early virus evolution
Cryo-EM structure of ssDNA bacteriophage ΦCjT23 provides insight into early virus evolution
Structural biology investigation of conserved capsid proteins facilitates the study of virus evolution. Here, characterization of the lipid-containing ssDNA bacteriophage ΦCjT23 suggests that this phage may serve as a model for the last common ancestor between ssDNA and dsDNA viruses in the Bamfordvirae.
- Nejc Kejzar
- , Elina Laanto
- & Juha T. Huiskonen
-
Article
| Open Access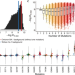 Compensatory epistasis maintains ACE2 affinity in SARS-CoV-2 Omicron BA.1
Compensatory epistasis maintains ACE2 affinity in SARS-CoV-2 Omicron BA.1
Evolution of the SARS-CoV-2 spike protein is likely driven by many factors, including immune escape and receptor binding. Here, by measuring the binding affinity of more than 30,000 variants of the SARS-CoV-2 RBD to its receptor ACE2, Moulana et al. show that the evolution of the Omicron BA.1 variant was driven by interactions between mutations.
- Alief Moulana
- , Thomas Dupic
- & Michael M. Desai
-
Article
| Open Access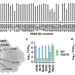 Prevalence and mechanisms of evolutionary contingency in human influenza H3N2 neuraminidase
Prevalence and mechanisms of evolutionary contingency in human influenza H3N2 neuraminidase
Lei et al. systematically characterized the epistasis among natural mutations in the neuraminidase of human influenza H3N2 virus, which provide insights into the biophysical constraints that shaped its evolution trajectory over the past half-century.
- Ruipeng Lei
- , Timothy J. C. Tan
- & Nicholas C. Wu
-
Matters Arising
| Open AccessMachine-learning prediction of hosts of novel coronaviruses requires caution as it may affect wildlife conservation
- Sophie Lund Rasmussen
- , Cino Pertoldi
- & David W. Macdonald
-
Matters Arising
| Open AccessReply to: Machine-learning prediction of hosts of novel coronaviruses requires caution as it may affect wildlife conservation
- Marcus S. C. Blagrove
- , Matthew Baylis
- & Maya Wardeh
-
Article
| Open Access Dynamics of competing SARS-CoV-2 variants during the Omicron epidemic in England
Dynamics of competing SARS-CoV-2 variants during the Omicron epidemic in England
This study presents data from the REACT-1 SARS-CoV-2 community sampling study in England from November 2021 to March 2022. They show that the Omicron variant peaked in January with a prevalence of ~7% and that the BA.2 sublineage had a 1.5x higher reproduction number compared to other Omicron sublineages.
- Oliver Eales
- , Leonardo de Oliveira Martins
- & Marc Chadeau-Hyam
-
Article
| Open Access A Bayesian approach to infer recombination patterns in coronaviruses
A Bayesian approach to infer recombination patterns in coronaviruses
Genetic recombination can confound standard phylogenetic approaches. Here, the authors present a method to reconstruct virus recombination networks, and show the importance of recombination in shaping the ongoing evolution of SARS-like, MERS and 3 human seasonal coronaviruses.
- Nicola F. Müller
- , Kathryn E. Kistler
- & Trevor Bedford
-
Article
| Open Access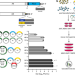 Influenza A virus undergoes compartmentalized replication in vivo dominated by stochastic bottlenecks
Influenza A virus undergoes compartmentalized replication in vivo dominated by stochastic bottlenecks
Transmission of influenza A viruses (IAV) between hosts and replication within host impose genetic bottlenecks, constraining viral diversity and adaptation. Here, Amato et al. perform site-specific inoculation of barcoded IAV of ferrets and track viral diversity as infection spreads to the lower respiratory tract and conclude that narrow population bottlenecks are an important feature of the within-host infection dynamics.
- Katherine A. Amato
- , Luis A. Haddock III
- & Andrew Mehle
-
Article
| Open Access Archival influenza virus genomes from Europe reveal genomic variability during the 1918 pandemic
Archival influenza virus genomes from Europe reveal genomic variability during the 1918 pandemic
For archival pathogens, like pH1N1 Influenza A virus the causative agent of 1918/19 pandemic, only few whole genome sequences exist. Here, Patrono et al. provide one complete and two partial genomes from Germany and find variation in two sites in the nucleoprotein gene in pandemic samples compared to pre-pandemic samples, that are associated with resistance to host antiviral response, pointing at a possible viral adaptation to humans.
- Livia V. Patrono
- , Bram Vrancken
- & Sébastien Calvignac-Spencer
-
Article
| Open Access De novo emergence of a remdesivir resistance mutation during treatment of persistent SARS-CoV-2 infection in an immunocompromised patient: a case report
De novo emergence of a remdesivir resistance mutation during treatment of persistent SARS-CoV-2 infection in an immunocompromised patient: a case report
Here, the authors identify and validate the emergence of a SARS-CoV-2 resistance mutation to Remdesivir, associated with virological recrudesce in an immunocompromised patient with persistent COVID-19.
- Shiv Gandhi
- , Jonathan Klein
- & Albert I. Ko
-
Article
| Open Access Evolution of the SARS-CoV-2 spike protein in the human host
Evolution of the SARS-CoV-2 spike protein in the human host
The SARS-CoV-2 spike has been evolving in the human population. The variants of concern alpha and beta evolved to optimise spike openness and so ability to bind its receptor ACE2, the affinity towards the receptor, and stability upon receptor binding.
- Antoni G. Wrobel
- , Donald J. Benton
- & Steven J. Gamblin
-
Article
| Open Access Functional comparison of MERS-coronavirus lineages reveals increased replicative fitness of the recombinant lineage 5
Functional comparison of MERS-coronavirus lineages reveals increased replicative fitness of the recombinant lineage 5
MERS-CoV is enzootic in dromedary camels, can spread to humans but undergoes limited onward transmission. Here, Schroeder et al. compare clinical isolates of MERS-CoV in vitro and show that the predominantly circulating recombinant lineage 5 possess a fitness advantage over parental lineage 3 and 4 due to reduced activation of innate immune signaling.
- Simon Schroeder
- , Christin Mache
- & Christian Drosten
-
Article
| Open Access Distinct patterns of within-host virus populations between two subgroups of human respiratory syncytial virus
Distinct patterns of within-host virus populations between two subgroups of human respiratory syncytial virus
Respiratory syncytial virus (RSV) is a common infection in children and older adults but little is known about within-host viral population diversity. Here, the authors perform deep sequencing and find that RSV subgroup B exhibited more diversity than subgroup A, with implications for development of therapeutics and vaccines.
- Gu-Lung Lin
- , Simon B. Drysdale
- & Andrew J. Pollard
-
Article
| Open Access Mapping mutations to the SARS-CoV-2 RBD that escape binding by different classes of antibodies
Mapping mutations to the SARS-CoV-2 RBD that escape binding by different classes of antibodies
Emerging SARS-CoV-2 mutants may escape neutralization by antibodies. Here, the authors use deep mutational scanning to identify mutations in the RBD that escape human monoclonal antibodies or convalescent plasmas.
- Allison J. Greaney
- , Tyler N. Starr
- & Jesse D. Bloom
-
Article
| Open Access Genomic epidemiology of the early stages of the SARS-CoV-2 outbreak in Russia
Genomic epidemiology of the early stages of the SARS-CoV-2 outbreak in Russia
The COVID-19 epidemic began later in Russia than many European countries, possibly due to restrictions on travel from China. Here, the authors analyze whole genome sequences sampled early in the epidemic in Russia, and find that most strains were not linked to China.
- Andrey B. Komissarov
- , Ksenia R. Safina
- & Georgii A. Bazykin
-
Article
| Open Access Parallel evolution in the emergence of highly pathogenic avian influenza A viruses
Parallel evolution in the emergence of highly pathogenic avian influenza A viruses
Highly pathogenic avian influenza viruses (HPAIV) can evolve via acquisition of polybasic cleavage sites, but the contribution of other mutations remains unclear. Here, the authors combine phylogenetic, statistical and structural approaches, and identify parallel mutations that are associated with HPAIV phenotype.
- Marina Escalera-Zamudio
- , Michael Golden
- & Oliver G. Pybus
-
Article
| Open Access Cryo-EM structure of coronavirus-HKU1 haemagglutinin esterase reveals architectural changes arising from prolonged circulation in humans
Cryo-EM structure of coronavirus-HKU1 haemagglutinin esterase reveals architectural changes arising from prolonged circulation in humans
Human coronavirus-HKU1 contains two surface projections called spike and haemagglutinin-esterase (HE), with the latter acting as a receptor-destroying enzyme. Here, the authors use cryo-EM and mass spectrometry to characterise the small, heavily glycosylated HKU1 HE, revealing a vestigial lectin domain covered with a putative glycan shield; and they discuss these features in the context of host adaptation.
- Daniel L. Hurdiss
- , Ieva Drulyte
- & Raoul J. de Groot
-
Article
| Open Access Entamoeba and Giardia parasites implicated as hosts of CRESS viruses
Entamoeba and Giardia parasites implicated as hosts of CRESS viruses
Metagenomics allows virus genome discovery, but the viral hosts are often not identified. Here, Kinsella et al. use recombination events between virus genomes, statistical association of viruses to hosts in clinical samples, and analysis of endogenous viral elements in host genomes to identify probable hosts of three CRESS virus families.
- Cormac M. Kinsella
- , Aldert Bart
- & Lia van der Hoek
-
Article
| Open Access A distinct lineage of Caudovirales that encodes a deeply branching multi-subunit RNA polymerase
A distinct lineage of Caudovirales that encodes a deeply branching multi-subunit RNA polymerase
Viruses have been difficult to position in the Tree of Life using phylogenetic methods. This study uses an ancient enzyme multi-subunit RNA polymerase (RNAP) to reveal a novel viral group, the Caudovirales, and to suggest an ancient origin of RNAP in this group.
- Alaina R. Weinheimer
- & Frank O. Aylward
-
Article
| Open Access Flexible genes establish widespread bacteriophage pan-genomes in cryoconite hole ecosystems
Flexible genes establish widespread bacteriophage pan-genomes in cryoconite hole ecosystems
Bacteriophages and their hosts are involved in a constant evolutionary arms race that should lead to divergence between phage genes over time. Here, the authors recruit metagenomic reads to virus reference genomes and genome fragments in samples from cryoconite holes and show that phages with near-identical core genomes maintain diversity by possession of numerous flexible gene modules, where homologous genes present in the pan-genome interchange to create new phage variants.
- Christopher M. Bellas
- , Declan C. Schroeder
- & Alexandre M. Anesio
-
Article
| Open Access Selective flexible packaging pathways of the segmented genome of influenza A virus
Selective flexible packaging pathways of the segmented genome of influenza A virus
The mechanism underlying packaging of the 8 segments of the influenza virus genome into virions is not well understood. Here, the authors use a multiplexed FISH assay to monitor the 8 segments in parallel in infected cells suggesting bundling routes during the packaging process.
- Ivan Haralampiev
- , Simon Prisner
- & Andreas Herrmann
-
Article
| Open Access Origin and cross-species transmission of bat coronaviruses in China
Origin and cross-species transmission of bat coronaviruses in China
Bats are a likely reservoir of zoonotic coronaviruses (CoVs). Here, analyzing bat CoV sequences in China, the authors find that alpha-CoVs have switched hosts more frequently than betaCoVs, identify a bat family and genus that are highly involved in host-switching, and define hotspots of CoV evolutionary diversity.
- Alice Latinne
- , Ben Hu
- & Peter Daszak
-
Article
| Open Access A hidden gene in astroviruses encodes a viroporin
A hidden gene in astroviruses encodes a viroporin
Astroviruses are common human pathogens and their genomes contain three known protein-coding genes. Here, Lulla et al. report a fourth, previously overlooked gene encoding protein XP which has a viroporin-like activity that is important for efficient production and/or release of virus particles.
- Valeria Lulla
- & Andrew E. Firth
-
Article
| Open Access Cryo-EM structures of HKU2 and SADS-CoV spike glycoproteins provide insights into coronavirus evolution
Cryo-EM structures of HKU2 and SADS-CoV spike glycoproteins provide insights into coronavirus evolution
Several coronaviruses infecting humans and animals have emerged in recent years. Here, the authors provide structures of the spike proteins of the porcine coronavirus SADS-CoV and closely related bat coronavirus HKU2, providing insights into evolution of coronavirus spike proteins.
- Jinfang Yu
- , Shuyuan Qiao
- & Xinquan Wang
-
Article
| Open Access The DNA methylation landscape of giant viruses
The DNA methylation landscape of giant viruses
DNA methylation is an epigenetic marker in all domains of life. Here, Jeudy et al., using single-molecule realtime sequencing, determine DNA methylation patterns in giant viruses and evolutionary analysis of virus encoded DNA methyltransferases suggests that they affect viral fitness.
- Sandra Jeudy
- , Sofia Rigou
- & Matthieu Legendre
-
Article
| Open Access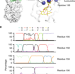 Major antigenic site B of human influenza H3N2 viruses has an evolving local fitness landscape
Major antigenic site B of human influenza H3N2 viruses has an evolving local fitness landscape
Antigenic site B in influenza A virus hemagglutinin (HA) is immunodominant in circulating human H3N2 strains. Using deep mutational scanning, Wu et al. here define the local fitness landscapes of HA antigenic site B in six human H3N2 strains, providing insights into evolvability of influenza antigenicity.
- Nicholas C. Wu
- , Jakub Otwinowski
- & Ian A. Wilson
-
Article
| Open Access Deconvolving mutational patterns of poliovirus outbreaks reveals its intrinsic fitness landscape
Deconvolving mutational patterns of poliovirus outbreaks reveals its intrinsic fitness landscape
Poliovirus has a higher mutation rate than HIV, yet has been almost eradicated by vaccination while an effective vaccine against HIV does not exist. Here, the authors develop a fitness model for poliovirus viral protein 1 to show that it is subject to stringent evolutionary constraints that limit its ability to avoid vaccine-induced immune responses.
- Ahmed A. Quadeer
- , John P. Barton
- & Matthew R. McKay
-
Article
| Open Access Incomplete influenza A virus genomes occur frequently but are readily complemented during localized viral spread
Incomplete influenza A virus genomes occur frequently but are readily complemented during localized viral spread
The genome of influenza is often incomplete in infected cells, but the implications for infection remain unclear. Here, Jacobs et al. show that an average of 3.6 particles is necessary for productive infection and that coinfection supports efficient complementation within a host but not upon transmission to a new host.
- Nathan T. Jacobs
- , Nina O. Onuoha
- & Anice C. Lowen
-
Article
| Open Access Profiling host ANP32A splicing landscapes to predict influenza A virus polymerase adaptation
Profiling host ANP32A splicing landscapes to predict influenza A virus polymerase adaptation
Polymorphisms in the avian influenza A virus (IAV) polymerase restrict its host range during transmission from birds to mammals. Here, the authors investigate differences in the host chromatin regulator ANP32A regarding IAV polymerase adaptation, and profile ANP32A splicing to predict avian species associated with pre-adaptive human-signatures in the virus.
- Patricia Domingues
- , Davide Eletto
- & Benjamin G. Hale
-
Article
| Open Access HIV-1 DNA sequence diversity and evolution during acute subtype C infection
HIV-1 DNA sequence diversity and evolution during acute subtype C infection
The dynamics of HIV-1 DNA sequences early after HIV-1 transmission remains poorly characterized. Here, the authors perform a longitudinal evaluation of HIV-1 DNA sequences in subtype C-infected individuals during acute infection, providing a landscape of the nature and evolution of the very early viral genome.
- Guinevere Q. Lee
- , Kavidha Reddy
- & Mathias Lichterfeld
-
Article
| Open Access Assembly of complex viruses exemplified by a halophilic euryarchaeal virus
Assembly of complex viruses exemplified by a halophilic euryarchaeal virus
Here, the authors present the cryo-EM structure of the archaeal virus SH1 at 3.8 Å resolution and show how the major capsid proteins assemble into hetero-hexamers, providing insights into the assembly process of this and related PRD1-adeno lineage viruses.
- Luigi De Colibus
- , Elina Roine
- & David I. Stuart
-
Article
| Open Access Virus-mediated export of chromosomal DNA in plants
Virus-mediated export of chromosomal DNA in plants
Viruses are potential vectors for horizontal gene transfer. Here, studying viral infection of sugar beet plants, the authors report the generation of virus-host circular DNA hybrids and provide a picture of the initial steps in virus-mediated horizontal transfer of chromosomal DNA between plant species.
- Marco Catoni
- , Emanuela Noris
- & Gian Paolo Accotto
-
Article
| Open Access A naturally protective epitope of limited variability as an influenza vaccine target
A naturally protective epitope of limited variability as an influenza vaccine target
Current influenza vaccine approaches largely focus on highly variable epitopes with high immunogenicity or epitopes of low variability that often have low immunogenicity. Here, Thompson et al. identify a highly immunogenic epitope of limited variability in the head domain of the H1 haemagglutinin and show protection from diverse H1N1 strains in mice.
- Craig P. Thompson
- , José Lourenço
- & Sunetra Gupta
-
Article
| Open Access Diversity and evolution of the emerging Pandoraviridae family
Diversity and evolution of the emerging Pandoraviridae family
Giant viruses are visible by light microscopy and have unusually long genomes. Here, the authors report three new members of the Pandoraviridae family and investigate their evolution and diversity.
- Matthieu Legendre
- , Elisabeth Fabre
- & Jean-Michel Claverie

 Genetic determinants of host- and virus-derived insertions for hepatitis E virus replication
Genetic determinants of host- and virus-derived insertions for hepatitis E virus replication