Featured
-
-
Article
| Open Access ID1 expressing macrophages support cancer cell stemness and limit CD8+ T cell infiltration in colorectal cancer
ID1 expressing macrophages support cancer cell stemness and limit CD8+ T cell infiltration in colorectal cancer
Inhibitor of differentiation 1 (ID1) has been described as a cancer-promoting factor and also involved in the formation of an immunosuppressive tumor microenvironment. Here the authors report that ID1-expressing tumor associated macrophages favor colorectal cancer progression by promoting cancer cell stemness and CD8+ T cell exclusion.
- Shuang Shang
- , Chen Yang
- & Fang Hua
-
Article
| Open Access c-di-GMP inhibits the DNA binding activity of H-NS in Salmonella
c-di-GMP inhibits the DNA binding activity of H-NS in Salmonella
H-NS is a global regulatory protein that represses expression of many genes in bacteria. Here, Li et al. show that a second messenger, cyclic di-GMP, binds to H-NS and inhibits its binding to DNA, thus relieving H-NS-mediated transcriptional silencing.
- Shuyu Li
- , Qinmeng Liu
- & Lei Zhang
-
Article
| Open Access Synthetic genetic oscillators demonstrate the functional importance of phenotypic variation in pneumococcal-host interactions
Synthetic genetic oscillators demonstrate the functional importance of phenotypic variation in pneumococcal-host interactions
Here, Rueff et al engineered a CRISPRi-based oscillator to rewire capsule production in Streptococcus pneumoniae from its native control. They show that heterogeneity in capsule production is beneficial for fitness in several virulence associated traits.
- Anne-Stéphanie Rueff
- , Renske van Raaphorst
- & Jan-Willem Veening
-
Article
| Open Access The environmentally-regulated interplay between local three-dimensional chromatin organisation and transcription of proVWX in E. coli
The environmentally-regulated interplay between local three-dimensional chromatin organisation and transcription of proVWX in E. coli
Here, the authors use the proVWXoperon of Escherichia coli as a model system to show how the nucleoid associated protein H-NS regulates gene expression in vivo by local chromatin remodelling.
- Fatema-Zahra M. Rashid
- , Frédéric G. E. Crémazy
- & Remus T. Dame
-
Article
| Open Access Transposable element-initiated enhancer-like elements generate the subgenome-biased spike specificity of polyploid wheat
Transposable element-initiated enhancer-like elements generate the subgenome-biased spike specificity of polyploid wheat
The direct impacts of transposable element dynamics on polyploid regulation and developmental specificity remain unclear. Here, the authors show that a large proportion of enhancer-like elements (ELEs) are mainly originated from RLG_famc7.3 specifically expanded in subgenome A, producing active nascent transcripts and influencing wheat spike development.
- Yilin Xie
- , Songbei Ying
- & Yijing Zhang
-
Article
| Open Access Transcriptional responses of cancer cells to heat shock-inducing stimuli involve amplification of robust HSF1 binding
Transcriptional responses of cancer cells to heat shock-inducing stimuli involve amplification of robust HSF1 binding
The authors compare the heat shock response between different cell lines and stimuli and reveal the genome-wide binding of its master transcription factor HSF1 as a platform for context-specific transcription activation.
- Sayantani Ghosh Dastidar
- , Bony De Kumar
- & Sergei Nechaev
-
Article
| Open Access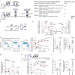 Themis2 regulates natural killer cell memory function and formation
Themis2 regulates natural killer cell memory function and formation
Innate immunity represents the first line of defence against pathogens, but certain innate cells are capable of memory formation, albeit with different and lesser-known mechanisms than adoptive immune cells. Here authors show that Themis2 regulates both memory NK cell development and function, via distinct downstream pathways.
- Tsukasa Nabekura
- , Elfira Amalia Deborah
- & Akira Shibuya
-
Article
| Open Access Transcriptional reprogramming by mutated IRF4 in lymphoma
Transcriptional reprogramming by mutated IRF4 in lymphoma
Cancer is often associated with mutant transcription factors (TFs) but their functional characterization is challenging. Here, the authors describe a recurrent mutation within TF IRF4 in human lymphomas and they show how it causes a complex switch in TF specificity and functionality.
- Nikolai Schleussner
- , Pierre Cauchy
- & Stephan Mathas
-
Article
| Open Access Paired yeast one-hybrid assays to detect DNA-binding cooperativity and antagonism across transcription factors
Paired yeast one-hybrid assays to detect DNA-binding cooperativity and antagonism across transcription factors
Combinations of transcription factors (TFs) bind DNA to fine-tune gene expression. Here, the authors map cooperative and antagonistic DNA binding across hundreds of TF-pairs. TF-TF relationships vary depending on DNA targets and TF isoforms.
- Anna Berenson
- , Ryan Lane
- & Juan I. Fuxman Bass
-
Article
| Open Access Structure of the transcription open complex of distinct σI factors
Structure of the transcription open complex of distinct σI factors
Here the authors show that σI factors encompass a unique, hitherto-unknown recognition mode of bacterial transcriptional promoters and represent a new distinctive class of σ70-family σ factors for bacterial transcription.
- Jie Li
- , Haonan Zhang
- & Ping Zhu
-
Article
| Open Access Transcriptional repression by a secondary DNA binding surface of DNA topoisomerase I safeguards against hypertranscription
Transcriptional repression by a secondary DNA binding surface of DNA topoisomerase I safeguards against hypertranscription
Aberrant hypertranscription in cells can affect development and disease. Here, the authors show that DNA topoisomerase I prevents hypertranscription; not through its catalytic function, but through a DNA binding mechanism.
- Mei Sheng Lau
- , Zhenhua Hu
- & Wee-Wei Tee
-
Article
| Open Access Acetylation reprograms MITF target selectivity and residence time
Acetylation reprograms MITF target selectivity and residence time
The microphthalmia-associated transcription factor MITF is a lineage-survival oncogene that plays a crucial role in melanocyte development and melanoma. Here, the authors reveal that MITF has a very long chromatin-bound half-life, and that MITF target selectivity is regulated by K206 acetylation, a residue linked to Waardenburg syndrome.
- Pakavarin Louphrasitthiphol
- , Alessia Loffreda
- & Colin R. Goding
-
Article
| Open Access Genome-wide probing of eukaryotic nascent RNA structure elucidates cotranscriptional folding and its antimutagenic effect
Genome-wide probing of eukaryotic nascent RNA structure elucidates cotranscriptional folding and its antimutagenic effect
Here, the authors present eSPET-seq a method to measure cotranscriptional RNA folding in eukaryotes. Further analysis reveals an antimutagenic effect of nascent RNA folding and contribution to the variability of local mutation rates across the yeast genome.
- Gongwang Yu
- , Yao Liu
- & Jian-Rong Yang
-
Article
| Open Access Activator-blocker model of transcriptional regulation by pioneer-like factors
Activator-blocker model of transcriptional regulation by pioneer-like factors
How gene expression timing is regulated during development remains a key area of research. Here they show that zebrafish genome activators Pou5f3 and Nanog block each other’s activity on the enhancers of differentiation genes, preventing their premature expression.
- Aileen Julia Riesle
- , Meijiang Gao
- & Daria Onichtchouk
-
Article
| Open Access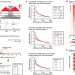 Multi-feature clustering of CTCF binding creates robustness for loop extrusion blocking and Topologically Associating Domain boundaries
Multi-feature clustering of CTCF binding creates robustness for loop extrusion blocking and Topologically Associating Domain boundaries
Most mammalian TAD boundaries, which separate functional chromosomal domains, bind the CTCF protein. Here, the authors identify multi-level clustering of CTCF binding sites at TAD boundaries and confirm their individual contribution to TAD formation.
- Li-Hsin Chang
- , Sourav Ghosh
- & Daan Noordermeer
-
Article
| Open Access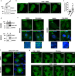 Cyclic AMP induces reversible EPAC1 condensates that regulate histone transcription
Cyclic AMP induces reversible EPAC1 condensates that regulate histone transcription
Spatial compartmentalization is central to nuclear function. Here, the authors demonstrate that EPAC1 can enter the nucleus and regulate the transcription of a histone cluster by forming biomolecular condensates in its proximity in response to cAMP.
- Liliana Felicia Iannucci
- , Anna Maria D’Erchia
- & Konstantinos Lefkimmiatis
-
Article
| Open Access Structural convergence endows nuclear transport receptor Kap114p with a transcriptional repressor function toward TATA-binding protein
Structural convergence endows nuclear transport receptor Kap114p with a transcriptional repressor function toward TATA-binding protein
Nuclear transport receptors mediate nucleocytoplasmic transport, collectively termed karyopherin-β (Kap-β) in yeast. Here, the authors present a cryo-EM structure of Kap114p, one of the Kap-βs, revealing a non-canonical function beyond nuclear transport that modulates yTBP-dependent transcription.
- Chung-Chi Liao
- , Yi-Sen Wang
- & Kuo-Chiang Hsia
-
Article
| Open Access MLL-AF4 cooperates with PAF1 and FACT to drive high-density enhancer interactions in leukemia
MLL-AF4 cooperates with PAF1 and FACT to drive high-density enhancer interactions in leukemia
Previous studies have reported MLL-AF4 binding at intragenic and intergenic enhancers, however, the role of MLL-AF4 in enhancer function remains to be investigated. Here, the authors show that MLL-AF4 cooperates with PAF1 and FACT at enhancers to promote high-density interactions with oncogene promoters in leukemia.
- Nicholas T. Crump
- , Alastair L. Smith
- & Thomas A. Milne
-
Article
| Open Access RNAPII-dependent ATM signaling at collisions with replication forks
RNAPII-dependent ATM signaling at collisions with replication forks
Deregulation of transcription by oncogenes leads to collisions of RNA Polymerase II (RNAPII) with DNA replication machinery (transcription-replication conflicts, TRCs). This study shows that RNAPII activates ATM kinase at TRCs providing a mechanism for replication fork stalling and ATM activation at TRCs.
- Elias Einig
- , Chao Jin
- & Nikita Popov
-
Article
| Open Access Redox driven B12-ligand switch drives CarH photoresponse
Redox driven B12-ligand switch drives CarH photoresponse
CarH is a bacterial B12-binding photoreceptor involved in transcriptional regulation. Here, the authors provide insights into B12 dynamics and associated cobalt redox changes following light activation. These demonstrate the CarH response integrates light and oxygen sensing.
- Harshwardhan Poddar
- , Ronald Rios-Santacruz
- & David Leys
-
Article
| Open Access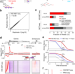 High-sensitive nascent transcript sequencing reveals BRD4-specific control of widespread enhancer and target gene transcription
High-sensitive nascent transcript sequencing reveals BRD4-specific control of widespread enhancer and target gene transcription
Here the authors reveal that high-sensitive nascent transcript sequencing provides an extended high-resolution view on transcription, including lowly transcribed enhancers. Widespread transcription at enhancers and their target genes depends on the BET family protein BRD4.
- Annkatrin Bressin
- , Olga Jasnovidova
- & Andreas Mayer
-
Article
| Open Access Structural basis for negative regulation of the Escherichia coli maltose system
Structural basis for negative regulation of the Escherichia coli maltose system
MalY represses the E. coli maltose system by direct interaction with MalT that blocks its oligomerization. Maltotriose-binding leads to conformational remodelling of MalT and stabilizes the C-terminal domains required for downstream signalling.
- Yuang Wu
- , Yue Sun
- & Jijie Chai
-
Article
| Open Access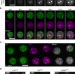 Stepwise modifications of transcriptional hubs link pioneer factor activity to a burst of transcription
Stepwise modifications of transcriptional hubs link pioneer factor activity to a burst of transcription
Eukaryotic transcription involves the formation of subnuclear hubs that enrich transcriptional machinery. Here the authors show that the hubs undergo stepwise modifications to fuel a burst of transcription rather than having a stable composition.
- Chun-Yi Cho
- & Patrick H. O’Farrell
-
Article
| Open Access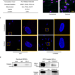 TRIM28 modulates nuclear receptor signaling to regulate uterine function
TRIM28 modulates nuclear receptor signaling to regulate uterine function
This paper identifies Tripartite motif-containing 28 (TRIM28) as a novel modulator of steroid hormone signaling. TRIM28 deficiency disrupts uterine cell functions and composition leading to fertility defects
- Rong Li
- , Tianyuan Wang
- & Francesco J. DeMayo
-
Article
| Open Access The HSV-1 ICP22 protein selectively impairs histone repositioning upon Pol II transcription downstream of genes
The HSV-1 ICP22 protein selectively impairs histone repositioning upon Pol II transcription downstream of genes
Herpes simplex virus 1 (HSV-1) infection disrupts transcription termination by RNA Polymerase II. Here, Djakovic et al. identify the immediate-early protein ICP22 protein of HSV-1 to induce open chromatin downstream of genes upon read-through transcription involving the histone chaperone FACT.
- Lara Djakovic
- , Thomas Hennig
- & Lars Dölken
-
Article
| Open Access Multifactor transcriptional control of alternative oxidase induction integrates diverse environmental inputs to enable fungal virulence
Multifactor transcriptional control of alternative oxidase induction integrates diverse environmental inputs to enable fungal virulence
Metabolic flexibility allows fungi to invade hostile niches. Here, Liu et al. dissect the molecular mechanisms by which Candida albicans upregulates virulence-enabling alternative oxidase expression in response to host-relevant respiratory stresses.
- Zhongle Liu
- , Pauline Basso
- & Leah E. Cowen
-
Article
| Open Access MOF-mediated histone H4 Lysine 16 acetylation governs mitochondrial and ciliary functions by controlling gene promoters
MOF-mediated histone H4 Lysine 16 acetylation governs mitochondrial and ciliary functions by controlling gene promoters
Here the authors show that epigenetic regulation through the histone acetyltransferase MOF and the acetylation of histone H4 lysine 16 affect essential functions in the mitochondria and primary cilia. This regulation is important to promote epidermal differentiation and hair follicle formation.
- Dongmei Wang
- , Haimin Li
- & Rui Yi
-
Article
| Open Access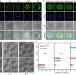 Maternal TDP-43 interacts with RNA Pol II and regulates zygotic genome activation
Maternal TDP-43 interacts with RNA Pol II and regulates zygotic genome activation
Zygotic genome activation is crucial for mammalian embryonic development. Here, the authors find that TDP-43 is indispensable for mouse embryonic development and mediates zygotic genome activation through Pol II configuration.
- Xiaoqing Nie
- , Qianhua Xu
- & Lei Li
-
Article
| Open Access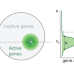 A model for organization and regulation of nuclear condensates by gene activity
A model for organization and regulation of nuclear condensates by gene activity
Through a physics-based model framework, the authors propose a central role for the nonequilibrium processes underling gene activity in shaping morphology, dynamics, and regulation of diverse nuclear condensates.
- Halima H. Schede
- , Pradeep Natarajan
- & Krishna Shrinivas
-
Article
| Open Access Selective binding of retrotransposons by ZFP352 facilitates the timely dissolution of totipotency network
Selective binding of retrotransposons by ZFP352 facilitates the timely dissolution of totipotency network
During zygotic genome activation the embryo must re-wire the regulatory network that sustains totipotency earlier during development. Here they identify ZFP352 as an essential factor that targets retrotransposon families to facilitate dissolution of the totipotency network and enable ZGA.
- Zhengyi Li
- , Haiyan Xu
- & Hongqing Liang
-
Article
| Open Access The NAD salvage pathway in mesenchymal cells is indispensable for skeletal development in mice
The NAD salvage pathway in mesenchymal cells is indispensable for skeletal development in mice
Deficiency in NAD+ has been implicated in skeletal deformities during development in both humans and mice. Here, the authors use mice that lack the critical enzyme of the NAD+ salvage pathway Nampt in mesenchymal lineage cells to show that the NAD salvage pathway is indispensable for endochondral but not intramembranous bone development.
- Aaron Warren
- , Ryan M. Porter
- & Maria Almeida
-
Article
| Open Access Chromatin remodeling by Pol II primes efficient Pol III transcription
Chromatin remodeling by Pol II primes efficient Pol III transcription
Transcription of RNA polymerase II is coupled with remodeling of chromatin. This study reports that transcription of RNA polymerase II is also required to prime and maintain nucleosome depletion at RNA polymerase III loci.
- Carlo Yague-Sanz
- , Valérie Migeot
- & Damien Hermand
-
Article
| Open Access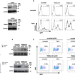 TRIM5α recruits HDAC1 to p50 and Sp1 and promotes H3K9 deacetylation at the HIV-1 LTR
TRIM5α recruits HDAC1 to p50 and Sp1 and promotes H3K9 deacetylation at the HIV-1 LTR
TRIM5α is known to restrict HIV-1 reverse transcription. Here, Ran et al. report that TRIM5α recruits HDAC1 to NF-κB p50, Sp1, and HIV-1 LTR, and promotes H3K9 deacetylation at the HIV-1 LTR, leading to TNFα-induced HIV-1 LTR-driven expression.
- Xiang-Hong Ran
- , Jia-Wu Zhu
- & Dan Mu
-
Article
| Open Access A multiple super-enhancer region establishes inter-TAD interactions and controls Hoxa function in cranial neural crest
A multiple super-enhancer region establishes inter-TAD interactions and controls Hoxa function in cranial neural crest
The authors discovered a far distant genomic region containing multiple clusters of regulatory elements that drive coordinated Hoxa expression across chromatin topologically associating domains in cranial neural crest, and are required for patterning of facial structures.
- Sandra Kessler
- , Maryline Minoux
- & Filippo M. Rijli
-
Article
| Open Access M1BP is an essential transcriptional activator of oxidative metabolism during Drosophila development
M1BP is an essential transcriptional activator of oxidative metabolism during Drosophila development
The transcriptional regulation of mitochondrial oxidative phosphorylation gene expression is poorly understood. Using the developing Drosophila flight muscle, the authors identify the transcription factor M1BP as a new major regulator of this process.
- Gabriela Poliacikova
- , Marine Barthez
- & Andrew J. Saurin
-
Article
| Open Access Transcriptional coactivation by EHMT2 restricts glucocorticoid-induced insulin resistance in a study with male mice
Transcriptional coactivation by EHMT2 restricts glucocorticoid-induced insulin resistance in a study with male mice
Glucocorticoids are known to induce insulin resistance via transcriptional activation of genes related to liver gluconeogenesis and insulin resistance. Here the authors report that in male mice treated with glucocorticoids, the transcriptional co-regulator EHMT2 is involved in the induction of Irs2 (a gene promoting insulin action) to restrict the extent of insulin resistance in the liver.
- Rebecca A. Lee
- , Maggie Chang
- & Jen-Chywan Wang
-
Article
| Open Access The conserved RNA-binding protein Seb1 promotes cotranscriptional ribosomal RNA processing by controlling RNA polymerase I progression
The conserved RNA-binding protein Seb1 promotes cotranscriptional ribosomal RNA processing by controlling RNA polymerase I progression
Ribosome biogenesis is influenced by the rate of RNAPI progression. Yet, mechanisms that control RNAPI elongation have remained elusive. Here, the authors show that the conserved protein Seb1 promotes cotranscriptional rRNA processing by controlling RNAPI progression.
- Maxime Duval
- , Carlo Yague-Sanz
- & François Bachand
-
Article
| Open Access The NELF pausing checkpoint mediates the functional divergence of Cdk9
The NELF pausing checkpoint mediates the functional divergence of Cdk9
Promoter-proximal pausing by RNA Pol II is a key aspect of how gene expression is transcriptionally regulated in higher eukaryotes. Here the authors show that only NELF-mediated pausing enforces a strict early checkpoint for Cdk9 by efficiently shutting down gene transcription following loss of Cdk9.
- Michael DeBerardine
- , Gregory T. Booth
- & John T. Lis
-
Article
| Open Access Double DAP-seq uncovered synergistic DNA binding of interacting bZIP transcription factors
Double DAP-seq uncovered synergistic DNA binding of interacting bZIP transcription factors
Here, the authors describe a new method to study how some proteins work together to control gene activity. They show that certain protein pairs can recognize new DNA sequences that they can’t recognize individually and control a wider range of genes.
- Miaomiao Li
- , Tao Yao
- & Shao-shan Carol Huang
-
Article
| Open Access Lung endothelial cells regulate pulmonary fibrosis through FOXF1/R-Ras signaling
Lung endothelial cells regulate pulmonary fibrosis through FOXF1/R-Ras signaling
Pulmonary fibrosis results from dysregulated lung repair, but the role of endothelial cells (EC) in fibrosis is unclear. Here, the authors show that FOXF1/R-Ras signalling in EC inhibits profibrotic mediators and that ECspecific nanoparticle FOXF1 gene therapy decreases lung fibrosis in mice.
- Fenghua Bian
- , Ying-Wei Lan
- & Tanya V. Kalin
-
Article
| Open Access Dependency of NELF-E-SLUG-KAT2B epigenetic axis in breast cancer carcinogenesis
Dependency of NELF-E-SLUG-KAT2B epigenetic axis in breast cancer carcinogenesis
Transcriptional dysregulation contributes to tumor progression. Here the authors show that transcriptional complex NELF interacts with SLUG, and co-opts KAT2B, to promote the expression of epithelial-mesenchymal transition (EMT) and stemness-associated genes in breast cancer.
- Jieqiong Zhang
- , Zhenhua Hu
- & Wee-Wei Tee
-
Article
| Open Access Transcription factor binding site orientation and order are major drivers of gene regulatory activity
Transcription factor binding site orientation and order are major drivers of gene regulatory activity
Gene regulatory grammar remains difficult to decipher, hindering our ability to link genotype to phenotype. Here they use massively parallel reporter assays to test over 200,000 synthetic sequences, finding that transcription factor binding site order and orientation have a major effect on gene regulatory activity.
- Ilias Georgakopoulos-Soares
- , Chengyu Deng
- & Nadav Ahituv
-
Article
| Open Access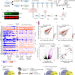 Buffering of transcription rate by mRNA half-life is a conserved feature of Rett syndrome models
Buffering of transcription rate by mRNA half-life is a conserved feature of Rett syndrome models
Rett syndrome (RTT) is a neurodevelopmental disorder caused by mutations in the transcriptional modulator MECP2. Here, the authors measured transcription rate and mRNA half-life changes in RTT patient-derived neurons to show transcription rate buffered by mRNA half-life changes.
- Deivid C. Rodrigues
- , Marat Mufteev
- & James Ellis
-
Article
| Open Access Structural basis of Ty1 integrase tethering to RNA polymerase III for targeted retrotransposon integration
Structural basis of Ty1 integrase tethering to RNA polymerase III for targeted retrotransposon integration
Cryo-EM structures of Ty1 integrase-Pol III complexes reveal determinants of Ty1 targeting upstream of Pol III-transcribed genes, and a functional impact of the integrase on Pol III activity that may increase the probability of Ty1 integration.
- Phong Quoc Nguyen
- , Sonia Huecas
- & Carlos Fernández-Tornero
-
Article
| Open Access The RRM-mediated RNA binding activity in T. brucei RAP1 is essential for VSG monoallelic expression
The RRM-mediated RNA binding activity in T. brucei RAP1 is essential for VSG monoallelic expression
Monoallelic VSG expression is essential for Trypanosoma brucei survival. Competition between TbRAP1’s RNA and dsDNA binding activities ensures that TbRAP1 sustains a high level expression of the active VSG while silencing other VSGs globally.
- Amit Kumar Gaurav
- , Marjia Afrin
- & Bibo Li
-
Article
| Open Access Dynamic interplay between non-coding enhancer transcription and gene activity in development
Dynamic interplay between non-coding enhancer transcription and gene activity in development
Non-coding transcription at the intergenic regulatory regions is a prevalent feature of metazoan genomes, but its function remains uncertain. Here the authors show that enhancer function is flexibly tunable through the modulation of hub formation via surrounding non-coding transcription.
- Kota Hamamoto
- , Yusuke Umemura
- & Takashi Fukaya
-
Article
| Open Access CvkR is a MerR-type transcriptional repressor of class 2 type V-K CRISPR-associated transposase systems
CvkR is a MerR-type transcriptional repressor of class 2 type V-K CRISPR-associated transposase systems
RNA-guided, CRISPR-associated transposons hold great promise for precision genome editing. Here, the authors provide genetic, biochemical and structural data how their activity is regulated in situ by CvkR, an unusual MerR family regulator.
- Marcus Ziemann
- , Viktoria Reimann
- & Wolfgang R. Hess
-
Article
| Open Access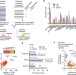 Widespread perturbation of ETS factor binding sites in cancer
Widespread perturbation of ETS factor binding sites in cancer
Few cancer drivers in non-coding regions have been identified so far. Here, the authors develop a transcription factor-aware burden test to predict non-coding variants and analyze the impact on transcription factor binding - especially ETS factors - as well as their impact on transcriptional activity.
- Sebastian Carrasco Pro
- , Heather Hook
- & Juan Ignacio Fuxman Bass
-
Article
| Open Access Angiotensin-converting enzyme inhibitor promotes angiogenesis through Sp1/Sp3-mediated inhibition of notch signaling in male mice
Angiotensin-converting enzyme inhibitor promotes angiogenesis through Sp1/Sp3-mediated inhibition of notch signaling in male mice
ACE inhibitors are widely used to treat cardiovascular diseases and promote angiogenesis. Here, the authors show a central role for endothelial USP7-Sp1/Sp3-Notch1 signalling in pathophysiological angiogenesis in response to ACE inhibitor treatment.
- Hanlin Lu
- , Peidong Yuan
- & Wencheng Zhang
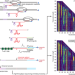
 Observation of coordinated RNA folding events by systematic cotranscriptional RNA structure probing
Observation of coordinated RNA folding events by systematic cotranscriptional RNA structure probing