Article
|
Open Access
Featured
-
-
Letter |
 The architecture of human general transcription factor TFIID core complex
The architecture of human general transcription factor TFIID core complex
The structures of three distinct human transcription factor IID (TFIID) protein assemblies are solved using cryo-electron microscopy; by incorporating TAF8 and TAF10, the key structural changes that remodel TFIID during assembly are determined, particularly the transition from a symmetric core-TFIID to an asymmetric holo-complex.
- Christoph Bieniossek
- , Gabor Papai
- & Imre Berger
-
Letter |
 Structure and function of the initially transcribing RNA polymerase II–TFIIB complex
Structure and function of the initially transcribing RNA polymerase II–TFIIB complex
Crystal structures of the Pol II–TFIIB complex in free form and bound by the DNA template and a short RNA product are reported; the latter complex represents an initially transcribing complex, a critical transient state in the pathway from transcription initiation to elongation.
- Sarah Sainsbury
- , Jürgen Niesser
- & Patrick Cramer
-
Article |
 Novel Foxo1-dependent transcriptional programs control Treg cell function
Novel Foxo1-dependent transcriptional programs control Treg cell function
The results of a series of genetic experiments indicate that Foxo1 has a pivotal, Foxp3-independent role controlling regulatory T-cell function.
- Weiming Ouyang
- , Will Liao
- & Ming O. Li
-
Letter |
 BTB-ZF factors recruit the E3 ligase cullin 3 to regulate lymphoid effector programs
BTB-ZF factors recruit the E3 ligase cullin 3 to regulate lymphoid effector programs
The E3 ubiquitin ligase cullin 3 is shown to bind BTB-zinc finger transcription factors to direct the ubiquitination of nuclear chromatin-associated factors to control transcription and cell-fate decisions in B- and T-cell populations.
- Rebecca Mathew
- , Michael P. Seiler
- & Albert Bendelac
-
Article |
 Generation of functional thyroid from embryonic stem cells
Generation of functional thyroid from embryonic stem cells
Transient overexpression of the transcription factors NKX2-1 and PAX8 in a murine cell model is shown to direct the differentiation of embryonic stem cells towards a thyroid follicular cell lineage; the resulting three-dimensional thyroid follicles created by subsequent thyrotropin treatment show hallmarks of thyroid function in vitro and rescue thyroid function in vivo when transplanted into athyroid mice, adding to our understanding of the molecular mechanisms underlying thyroid development.
- Francesco Antonica
- , Dominika Figini Kasprzyk
- & Sabine Costagliola
-
Article |
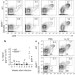 Compensatory dendritic cell development mediated by BATF–IRF interactions
Compensatory dendritic cell development mediated by BATF–IRF interactions
The roles of BATF transcription factors in dendritic cell differentiation are studied, providing evidence for molecular compensation by related family members; compensation is based on the interaction of the BATF leucine zipper domains with IRF factors to mediate cooperative gene activation.
- Roxane Tussiwand
- , Wan-Ling Lee
- & Kenneth M. Murphy
-
Article |
 Heart repair by reprogramming non-myocytes with cardiac transcription factors
Heart repair by reprogramming non-myocytes with cardiac transcription factors
A combination of four transcription factors, GATA4, HAND2, MEF2C and TBX5, can reprogram fibroblasts into cardiac-like myocytes in vitro and in vivo; expression of these factors ameliorated cardiac function in mice that had suffered myocardial infarction.
- Kunhua Song
- , Young-Jae Nam
- & Eric N. Olson
-
News & Views |
 Running to stand still
Running to stand still
Transcription factors regulate the expression of genes by binding to certain DNA sequences. But the outcome can be markedly different, depending on whether the binding is stable or short-lived. See Letter p.251
- Tommy Kaplan
- & Nir Friedman
-
Letter |
 Genome-wide protein–DNA binding dynamics suggest a molecular clutch for transcription factor function
Genome-wide protein–DNA binding dynamics suggest a molecular clutch for transcription factor function
Competition ChIP with a sequence-specific S. cerevisiae transcription factor, Rap1, reveals that long Rap1 residence is coupled to transcriptional activation, whereas fast binding turnover is linked to low transcriptional output, suggesting that DNA-binding events that appear identical by conventional ChIP may have different underlying modes of interaction, leading to opposing functional outcomes.
- Colin R. Lickwar
- , Florian Mueller
- & Jason D. Lieb
-
Letter |
 Transcription factor PIF4 controls the thermosensory activation of flowering
Transcription factor PIF4 controls the thermosensory activation of flowering
A novel mechanism by which warming temperatures can directly activate flowering in the model plant Arabidopsis thaliana.
- S. Vinod Kumar
- , Doris Lucyshyn
- & Philip A. Wigge
-
Letter |
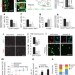 Ars2 maintains neural stem-cell identity through direct transcriptional activation of Sox2
Ars2 maintains neural stem-cell identity through direct transcriptional activation of Sox2
Mammalian zinc finger protein Ars2 is revealed as a sequence-specific transcription factor that promotes the self-renewal of postnatal and adult neural stem cells by directly activating transcription of the pluripotency factor Sox2.
- Celia Andreu-Agullo
- , Thomas Maurin
- & Eric C. Lai
-
Letter |
 T-cell differentiation factor CBF-β regulates HIV-1 Vif-mediated evasion of host restriction
T-cell differentiation factor CBF-β regulates HIV-1 Vif-mediated evasion of host restriction
CBF-β is shown to regulate the ability of HIV-1 to evade host restriction mediated by the deaminase APOBEC3.
- Wenyan Zhang
- , Juan Du
- & Xiao-Fang Yu
-
Letter |
 Spalt mediates an evolutionarily conserved switch to fibrillar muscle fate in insects
Spalt mediates an evolutionarily conserved switch to fibrillar muscle fate in insects
- Cornelia Schönbauer
- , Jutta Distler
- & Frank Schnorrer
-
Letter |
 Regulatory evolution through divergence of a phosphoswitch in the transcription factor CEBPB
Regulatory evolution through divergence of a phosphoswitch in the transcription factor CEBPB
- Vincent J. Lynch
- , Gemma May
- & Günter P. Wagner
-
Letter |
 Control of Drosophila endocycles by E2F and CRL4CDT2
Control of Drosophila endocycles by E2F and CRL4CDT2
- Norman Zielke
- , Kerry J. Kim
- & Bruce A. Edgar
-
Article |
 A critical role for TCF-1 in T-lineage specification and differentiation
A critical role for TCF-1 in T-lineage specification and differentiation
- Brittany Nicole Weber
- , Anthony Wei-Shine Chi
- & Avinash Bhandoola
-
Letter |
 DMRT1 prevents female reprogramming in the postnatal mammalian testis
DMRT1 prevents female reprogramming in the postnatal mammalian testis
- Clinton K. Matson
- , Mark W. Murphy
- & David Zarkower
-
Letter |
 Direct reprogramming of somatic cells is promoted by maternal transcription factor Glis1
Direct reprogramming of somatic cells is promoted by maternal transcription factor Glis1
- Momoko Maekawa
- , Kei Yamaguchi
- & Shinya Yamanaka
-
Letter |
 Induction of human neuronal cells by defined transcription factors
Induction of human neuronal cells by defined transcription factors
- Zhiping P. Pang
- , Nan Yang
- & Marius Wernig
-
Letter |
 A cis-regulatory map of the Drosophila genome
A cis-regulatory map of the Drosophila genome
As part of the modENCODE initiative, which aims to characterize functional DNA elements in D. melanogaster and C. elegans, this study created a map of the regulatory part of the fruitfly genome. On the basis of the developmental dynamics of chromatin modifications, polymerase and transcription factor occupancy this work defines a vast array of putative regulatory elements, such as enhancers, promoters, insulators and silencers. This resource represents the first attempt at a comprehensive annotation of cis-regulatory elements in a metazoan genome.
- Nicolas Nègre
- , Christopher D. Brown
- & Kevin P. White
-
Article |
 CRTC3 links catecholamine signalling to energy balance
CRTC3 links catecholamine signalling to energy balance
β-adrenergic receptor signalling in adipocytes stimulates energy expenditure via cAMP-dependent increases in lipolysis and fatty-acid oxidation, and this signalling mechanism is thought to be disrupted in obesity. Here, the cAMP-responsive CREB coactivator Crtc3 is shown to promote obesity in mice by attenuating β adrenergic receptor signalling in adipose tissue.
- Youngsup Song
- , Judith Altarejos
- & Marc Montminy
-
Article |
 Heterochromatin silencing of p53 target genes by a small viral protein
Heterochromatin silencing of p53 target genes by a small viral protein
Adenovirus E1B-55k targets transcription factor p53 for degradation and is thought to be critical for p53 inactivation during adenovirus replication. Indeed, mutant viruses lacking E1B-55k have been tested as viral cancer therapies selective for p53-positive tumours. These authors find another adenoviral protein, E4-ORF3, to inactivate p53 independently of E1B-55k by means of a chromatin-silencing mechanism that prevents access of p53 to its DNA target sites.
- Conrado Soria
- , Fanny E. Estermann
- & Clodagh C. O’Shea
-
Letter |
 Histone H4K20/H3K9 demethylase PHF8 regulates zebrafish brain and craniofacial development
Histone H4K20/H3K9 demethylase PHF8 regulates zebrafish brain and craniofacial development
PHF8 is a JmjC domain-containing protein, the gene for which has been linked to X-linked mental retardation (XLMR). These authors demonstrate PHF8 to be a histone demethylase with activity against H4K20me1. It has a role in regulating gene expression as well as in neuronal cell survival and craniofacial development in zebrafish. The results suggest there may be a link between histone methylation dynamics and XLMR.
- Hank H. Qi
- , Madathia Sarkissian
- & Yang Shi
-
Letter |
 A new DAF-16 isoform regulates longevity
A new DAF-16 isoform regulates longevity
The insulin/IGF-1 signalling (IIS) pathway is involved in various biological processes, including regulation of longevity. In the worm Caenorhabditis elegans, the transcription factor DAF-16a, one of two isoforms, has a major role in this pathway, regulating longevity, stress response and dauer diapause. These authors describe a new isoform, DAF-16d/f, which is also important in the regulation of lifespan. The DAF-16 isoforms functionally cooperate to fine-tune IIS-mediated processes in the context of a whole organism.
- Eun-Soo Kwon
- , Sri Devi Narasimhan
- & Heidi A. Tissenbaum
-
News & Views |
 Roots respond to an inner calling
Roots respond to an inner calling
In plant roots, patterning of two types of water-conducting xylem tissue is determined by a signalling system that involves the reciprocal dance of a mobile transcription factor and mobile microRNAs.
- Ben Scheres
-
Letter |
 IκBζ regulates TH17 development by cooperating with ROR nuclear receptors
IκBζ regulates TH17 development by cooperating with ROR nuclear receptors
Interleukin-17-producing helper T (TH17) cells are a distinct T-cell subset characterized by its role in autoimmune disease. Here it is shown that the development of TH17 cells requires the transcription factor IκBζ, as well as nuclear receptors of the ROR family. Mice lacking IκBζ have a defect in TH17 development and are resistant to the induction of experimental autoimmune encephalomyelitis. The study points to some new potential molecular targets for drugs to treat autoimmune disease.
- Kazuo Okamoto
- , Yoshiko Iwai
- & Hiroshi Takayanagi
-
Letter |
 NINJA connects the co-repressor TOPLESS to jasmonate signalling
NINJA connects the co-repressor TOPLESS to jasmonate signalling
In plants, the hormone jasmonoyl-isoleucine (JA-Ile) regulates growth, development and defence against pathogens. Proteins of the JAZ family repress JA-Ile-dependent gene expression, but the mechanism has been unclear. Here, an adaptor protein, NINJA, has been identified, which recruits co-repressor proteins that are known to mediate auxin-responsive gene expression as well. Hence these co-repressors are part of general repression complexes that are recruited to several different signalling pathways.
- Laurens Pauwels
- , Gemma Fernández Barbero
- & Alain Goossens
-
Letter |
 Primary contribution to zebrafish heart regeneration by gata4+ cardiomyocytes
Primary contribution to zebrafish heart regeneration by gata4+ cardiomyocytes
Zebrafish are able to replace lost heart muscle efficiently, and are used as a model to understand why natural heart regeneration — after a heart attack, for instance — is blocked in mammals. Here, and in an accompanying paper, genetic fate-mapping approaches reveal which cell population contributes prominently to cardiac muscle regeneration after an injury approximating myocardial infarction. The results show that cardiac muscle regenerates through activation and expansion of existing cardiomyocytes, without involving a stem-cell population.
- Kazu Kikuchi
- , Jennifer E. Holdway
- & Kenneth D. Poss
-
Letter |
 Genetic analysis of variation in transcription factor binding in yeast
Genetic analysis of variation in transcription factor binding in yeast
Variation in the regulation of gene transcription between individuals is thought to be a major cause of phenotypic diversity. Here, individual differences in the binding of transcription-factor proteins are studied. A well-known transcription factor in the yeast pheromone pathway is used as an example, and the underlying genetic loci responsible for variation in its binding are mapped. The study reveals new insights into the mechanisms of gene regulation, and new regulators of the yeast pheromone pathway.
- Wei Zheng
- , Hongyu Zhao
- & Michael Snyder
-
Letter |
 MONOPTEROS controls embryonic root initiation by regulating a mobile transcription factor
MONOPTEROS controls embryonic root initiation by regulating a mobile transcription factor
During Arabidopsis embryogenesis, a single cell is specified to become the founder cell of the root meristem — the hypophysis — in response to signals from adjacent cells. Hypophysis specification requires an auxin-responsive transcription factor, MONOPTEROS (MP), which promotes transport of auxin from the embryo to the hypophysis precursor. Here, MP target genes are identified and the means by which they mediate root formation is shown.
- Alexandra Schlereth
- , Barbara Möller
- & Dolf Weijers
-
Letter |
 Control of Arabidopsis apical–basal embryo polarity by antagonistic transcription factors
Control of Arabidopsis apical–basal embryo polarity by antagonistic transcription factors
During development in Arabidopsis plants, populations of shoot stem cells and root stem cells are established at the embryo's apical and basal poles, respectively. PLETHORA genes are master regulators of root fate, but the regulators of shoot fate were unknown. Here, CLASS III HOMEODOMAIN-LEUCINE ZIPPER genes are identified as master regulators of apical/shoot fate, and are shown to be sufficient to convert the embryonic root pole into a second shoot pole.
- Zachery R. Smith
- & Jeff A. Long
-
Article |
 Rfx6 directs islet formation and insulin production in mice and humans
Rfx6 directs islet formation and insulin production in mice and humans
Pancreatic β-cells release insulin, which controls energy homeostasis in vertebrates, and its lack causes diabetes mellitus. The transcription factor neurogenin 3 (Neurog3) initiates differentiation of β-cells and other islet cell types from pancreatic endoderm; here, the transcription factor Rfx6 is shown to direct islet cell differentiation downstream of Neurog3 in mice and humans. This may be useful in efforts to generate β-cells for patients with diabetes.
- Stuart B. Smith
- , Hui-Qi Qu
- & Michael S. German
-
Letter |
 Tbx3 improves the germ-line competency of induced pluripotent stem cells
Tbx3 improves the germ-line competency of induced pluripotent stem cells
The transcription factor Tbx3 is shown to significantly improve the quality of induced pluripotent stem (iPS) cells. Tbx3 binding sites in embryonic stem cells are present in genes involved in pluripotency and reprogramming factors. Furthermore, there are intrinsic qualitative differences in iPS cells generated by different methods in terms of their pluripotency, thus highlighting the need to rigorously characterize iPS cells beyond in vitro studies.
- Jianyong Han
- , Ping Yuan
- & Bing Lim
-
Letter |
 TGF-β–FOXO signalling maintains leukaemia-initiating cells in chronic myeloid leukaemia
TGF-β–FOXO signalling maintains leukaemia-initiating cells in chronic myeloid leukaemia
Chronic myeloid leukaemia is caused by a BCR-ABL fusion, a constitutively active tyrosine kinase that, it is believed, leads to suppression of the forkhead O transcription factors (FOXO). Although the use of the tyrosine kinase inhibitor imatinib is a breakthrough for CML therapy, imatinib does not deplete the leukaemia-initiating cells (LICs) that drive the recurrence of CML. Foxo3a is now shown to have an essential role in the maintenance of CML LICs in a mouse model.
- Kazuhito Naka
- , Takayuki Hoshii
- & Atsushi Hirao
-
Research Highlights |
Cancer biology: Weighted cancer risk
-
Article |
 Direct conversion of fibroblasts to functional neurons by defined factors
Direct conversion of fibroblasts to functional neurons by defined factors
Mouse and human fibroblasts can be reprogrammed to a pluripotent state with a combination of four transcription factors. Here, mature differentiated cells are directed, via a combination of a few transcription factors (distinct from those described for generating iPS cells), to form functional neurons in vitro, without having to revert the fibroblasts to an embryonic state.
- Thomas Vierbuchen
- , Austin Ostermeier
- & Marius Wernig
-
Letter |
 FOXO-dependent regulation of innate immune homeostasis
FOXO-dependent regulation of innate immune homeostasis
Antimicrobial peptides (AMPs) are an important class of immune effector molecules which fight pathogen infections. AMP induction in Drosophila is regulated through the activation of the Toll and immune deficiency pathways; it is now shown that AMP activation can be achieved independently of these pathways by the transcription factor FOXO. In non-infected animals, AMP genes are activated in response to nuclear FOXO activity when induced by starvation.
- Thomas Becker
- , Gerrit Loch
- & Michael Hoch
-
Letter |
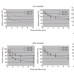 An allosteric mechanism of Rho-dependent transcription termination
An allosteric mechanism of Rho-dependent transcription termination
Rho is a general transcription termination factor in bacteria, but the mechanism by which it disrupts the RNA polymerase (RNAP) elongation complex is unknown. Here, Rho is shown to bind tightly to the RNAP throughout the transcription cycle, with the formation of the RNAP–Rho complex being crucial for termination. Furthermore, RNAP is proposed to have an active role in Rho termination through an allosteric mechanism.
- Vitaly Epshtein
- , Dipak Dutta
- & Evgeny Nudler

 A NPAS4–NuA4 complex couples synaptic activity to DNA repair
A NPAS4–NuA4 complex couples synaptic activity to DNA repair