Article
|
Open Access
Featured
-
-
Article
| Open Access Multi-batch single-cell comparative atlas construction by deep learning disentanglement
Multi-batch single-cell comparative atlas construction by deep learning disentanglement
Comparing single-cell RNA-seq and ATAC-seq data from multiple batches is challenging due to technical artifacts. Here, the authors propose a method that disentangles technical and biological effects, facilitating batch-confounded chromatin and gene expression state discovery and enhancing the analysis of perturbation effects on cell populations.
- Allen W. Lynch
- , Myles Brown
- & Clifford A. Meyer
-
Article
| Open Access The role of vaccination and public awareness in forecasts of Mpox incidence in the United Kingdom
The role of vaccination and public awareness in forecasts of Mpox incidence in the United Kingdom
An outbreak of Mpox in the UK began in May 2022 and peaked in July. In this modelling study, the authors show that the decline in cases was likely due to behavioural changes among high-risk populations, whilst vaccination could prevent a rebound.
- Samuel P. C. Brand
- , Massimo Cavallaro
- & Matt J. Keeling
-
Article
| Open Access Genome-wide association analysis and Mendelian randomization proteomics identify drug targets for heart failure
Genome-wide association analysis and Mendelian randomization proteomics identify drug targets for heart failure
Here, the authors perform a large-scale meta-analysis of genome-wide association studies and cis-MR proteomics to identify protein biomarkers and drug targets for heart failure.
- Danielle Rasooly
- , Gina M. Peloso
- & Juan P. Casas
-
Article
| Open Access nnSVG for the scalable identification of spatially variable genes using nearest-neighbor Gaussian processes
nnSVG for the scalable identification of spatially variable genes using nearest-neighbor Gaussian processes
The identification of top spatially variable genes is a key step in the analysis of spatially-resolved transcriptomics data. Here, the authors develop a scalable method based on nearest-neighbor Gaussian processes and evaluate performance compared to existing and baseline methods.
- Lukas M. Weber
- , Arkajyoti Saha
- & Stephanie C. Hicks
-
Article
| Open Access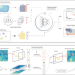 Leveraging spatial transcriptomics data to recover cell locations in single-cell RNA-seq with CeLEry
Leveraging spatial transcriptomics data to recover cell locations in single-cell RNA-seq with CeLEry
Cell location information is important for understanding how tissue is spatially organized. Here, the authors develop CeLEry, a machine learning method that aims to recover cell locations for single-cell RNA-seq data by leveraging information learned from spatial transcriptomics.
- Qihuang Zhang
- , Shunzhou Jiang
- & Mingyao Li
-
Article
| Open Access SpatialDM for rapid identification of spatially co-expressed ligand–receptor and revealing cell–cell communication patterns
SpatialDM for rapid identification of spatially co-expressed ligand–receptor and revealing cell–cell communication patterns
Spatial omics are increasingly being recognised to study cell-cell communications. Here, the authors present a bioinformatics toolbox for rapid identification of spatially co-expressed ligand-receptor and revealing cell-cell communication patterns.
- Zhuoxuan Li
- , Tianjie Wang
- & Yuanhua Huang
-
Article
| Open Access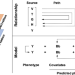 Joint analysis of phenotype-effect-generation identifies loci associated with grain quality traits in rice hybrids
Joint analysis of phenotype-effect-generation identifies loci associated with grain quality traits in rice hybrids
Genetic dissection of hybrids is more difficult than inbreds as nonadditive effects are involved. Here, the authors report a pipeline for joint analysis of phenotypes, effects, and generations and demonstrate its usefulness in identification of loci associated with quality traits and improving predict accuracy in genomic selection of hybrid rice.
- Lanzhi Li
- , Xingfei Zheng
- & Zhongli Hu
-
Article
| Open Access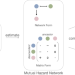 Joint inference of exclusivity patterns and recurrent trajectories from tumor mutation trees
Joint inference of exclusivity patterns and recurrent trajectories from tumor mutation trees
Understanding cancer evolution is crucial for developing effective therapies. Here, authors present TreeMHN, a probabilistic model for inferring exclusivity patterns of genomic events and evolutionary trajectories from intra-tumor phylogenetic trees.
- Xiang Ge Luo
- , Jack Kuipers
- & Niko Beerenwinkel
-
Article
| Open Access grandR: a comprehensive package for nucleotide conversion RNA-seq data analysis
grandR: a comprehensive package for nucleotide conversion RNA-seq data analysis
Nucleotide conversion approaches facilitate metabolic RNA labelling experiments but complicate computational analysis. Here, the authors develop a methodology and software package to enable specific analysis methods for nucleotide conversion RNA-seq data.
- Teresa Rummel
- , Lygeri Sakellaridi
- & Florian Erhard
-
Article
| Open Access A computational method for cell type-specific expression quantitative trait loci mapping using bulk RNA-seq data
A computational method for cell type-specific expression quantitative trait loci mapping using bulk RNA-seq data
Detecting cell type-specific genetic effects on gene expression is challenging in bulk RNA-seq data. Here, the authors develop a method to increase power which incorporates allele-specific expression and does not transform the gene expression data.
- Paul Little
- , Si Liu
- & Wei Sun
-
Article
| Open Access A positive statistical benchmark to assess network agreement
A positive statistical benchmark to assess network agreement
In the variety of biological and social networks, the validation of experimental data is done by comparing an overlap with reference networks. The authors introduce a positive statistical benchmark corresponding to the best possible overlap between two networks to threshold and validate new experimental datasets.
- Bingjie Hao
- & István A. Kovács
-
Article
| Open Access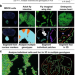 The PECAn image and statistical analysis pipeline identifies Minute cell competition genes and features
The PECAn image and statistical analysis pipeline identifies Minute cell competition genes and features
The 3D nature of clones makes sample image analysis challenging. Here the authors report PECAn, a pipeline for image processing and statistical analysis of complex multi-genotype 3D images, and apply this to the study of Minute cell competition in drosophila.
- Michael E. Baumgartner
- , Paul F. Langton
- & Eugenia Piddini
-
Article
| Open Access Reconstruction of the cell pseudo-space from single-cell RNA sequencing data with scSpace
Reconstruction of the cell pseudo-space from single-cell RNA sequencing data with scSpace
Methods to reanalyze scRNA-seq data in a spatial perspective are vital but lacking. Here, the authors develop scSpace, an integrative method that uses ST data as spatial reference to reconstruct the pseudo-space of scRNA-seq data and identify spatially variable cell subpopulations, providing insights into spatial heterogeneity from scRNA-seq data.
- Jingyang Qian
- , Jie Liao
- & Xiaohui Fan
-
Article
| Open Access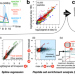 Virome-wide detection of natural infection events and the associated antibody dynamics using longitudinal highly-multiplexed serology
Virome-wide detection of natural infection events and the associated antibody dynamics using longitudinal highly-multiplexed serology
Methods to detect infections are often limited to specific viruses or do not yield information on the immune response. Here, the authors analyse temporal changes in high-dimensional antibody profiles and chart more than 650 infection events across the human virome providing a high-resolution view of host-virus dynamics.
- Erin J. Kelley
- , Sierra N. Henson
- & John A. Altin
-
Article
| Open Access Benchmarking integration of single-cell differential expression
Benchmarking integration of single-cell differential expression
Integration of single-cell RNA sequencing data between different samples has been a major challenge for analyzing cell populations. Here the authors benchmark 46 workflows for differential expression analysis of single-cell data with multiple batches and suggest several high-performance methods under different conditions based on simulation and real data analyses.
- Hai C. T. Nguyen
- , Bukyung Baik
- & Dougu Nam
-
Article
| Open Access Reciprocal causation mixture model for robust Mendelian randomization analysis using genome-scale summary data
Reciprocal causation mixture model for robust Mendelian randomization analysis using genome-scale summary data
Mendelian randomization methods are prone to produce false positive results when assumptions are violated. Here, the authors propose a statistical model that offers good power to detect causation between traits while controlling the false positive rate.
- Zipeng Liu
- , Yiming Qin
- & Pak Chung Sham
-
Article
| Open Access Reconstructing clonal tree for phylo-phenotypic characterization of cancer using single-cell transcriptomics
Reconstructing clonal tree for phylo-phenotypic characterization of cancer using single-cell transcriptomics
The functional changes of individual clones in single cell RNA sequencing (scRNA-seq) data remain elusive. Here, the authors develop PhylEx that integrates bulk genomics data with co-occurrences of mutations revealed by scRNA-seq data and apply it to high-grade serous ovarian cancer cell line and breast cancer datasets.
- Seong-Hwan Jun
- , Hosein Toosi
- & Jens Lagergren
-
Article
| Open Access Transcription factor binding sites are frequently under accelerated evolution in primates
Transcription factor binding sites are frequently under accelerated evolution in primates
Characterizing genomic elements under accelerated evolution is crucial for understanding the genomic basis of human evolution and disease. Here, Zhang et al. introduce GroupAcc, a collection of two pooling-based phylogenetic methods with enhanced sensitivity to examine accelerated evolution in transcription factor binding sites.
- Xinru Zhang
- , Bohao Fang
- & Yi-Fei Huang
-
Article
| Open Access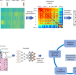 Cartography of Genomic Interactions Enables Deep Analysis of Single-Cell Expression Data
Cartography of Genomic Interactions Enables Deep Analysis of Single-Cell Expression Data
Existing genomic data analysis methods tend to not take full advantage of underlying biological characteristics. Here, the authors leverage the inherent interactions of scRNA-seq data and develop a cartography strategy to contrive the data into a spatially configured genomap for accurate deep pattern discovery.
- Md Tauhidul Islam
- & Lei Xing
-
Article
| Open Access MiXcan: a framework for cell-type-aware transcriptome-wide association studies with an application to breast cancer
MiXcan: a framework for cell-type-aware transcriptome-wide association studies with an application to breast cancer
Conventional transcriptome-wide association study (TWAS) approaches predict genetically regulated gene expression at the tissue level. Here, the authors develop a framework for cell-type-aware TWAS that predicts cell-type level expression from genotype data and identifies disease-associated genes with cell-type-specific effects.
- Xiaoyu Song
- , Jiayi Ji
- & Weiva Sieh
-
Article
| Open Access Probabilistic embedding, clustering, and alignment for integrating spatial transcriptomics data with PRECAST
Probabilistic embedding, clustering, and alignment for integrating spatial transcriptomics data with PRECAST
Methods that perform data integration are needed to analyse spatial transcriptomics data from multiple tissue slides. Here, the authors present PRECAST, an efficient data integration method for multiple spatial transcriptomics datasets with complex batch or biological effects between slides.
- Wei Liu
- , Xu Liao
- & Jin Liu
-
Article
| Open Access Direct and indirect effects of the COVID-19 pandemic on mortality in Switzerland
Direct and indirect effects of the COVID-19 pandemic on mortality in Switzerland
COVID-19-releated public health measures may have indirectly impacted mortality rates by causing or averting deaths. Here, the authors use data from Switzerland until April 2022 and estimate that, after accounting for deaths directly related to COVID-19, mortality was lower than expected, indicating some evidence of an overall positive impact of control measures.
- Julien Riou
- , Anthony Hauser
- & Garyfallos Konstantinoudis
-
Matters Arising
| Open AccessReply to: A balanced measure shows superior performance of pseudobulk methods in single-cell RNA-sequencing analysis
- Kip D. Zimmerman
- , Ciaran Evans
- & Carl D. Langefeld
-
Matters Arising
| Open Access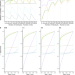 A balanced measure shows superior performance of pseudobulk methods in single-cell RNA-sequencing analysis
A balanced measure shows superior performance of pseudobulk methods in single-cell RNA-sequencing analysis
- Alan E. Murphy
- & Nathan G. Skene
-
Article
| Open Access Benchmarking tools for detecting longitudinal differential expression in proteomics data allows establishing a robust reproducibility optimization regression approach
Benchmarking tools for detecting longitudinal differential expression in proteomics data allows establishing a robust reproducibility optimization regression approach
Longitudinal proteomics holds great promise for biomarker discovery, but the data interpretation has remained a challenge. Here, the authors evaluate several tools to detect longitudinal differential expression in proteomics data and introduce RolDE, a robust reproducibility optimization approach.
- Tommi Välikangas
- , Tomi Suomi
- & Laura L. Elo
-
Article
| Open Access Estimating diagnostic uncertainty in artificial intelligence assisted pathology using conformal prediction
Estimating diagnostic uncertainty in artificial intelligence assisted pathology using conformal prediction
Artificial intelligence prediction accuracy can be reduced with new data. Here, the authors utilise conformal prediction to reduce incorrect predictions in histopathological analysis of prostate cancer biopsies.
- Henrik Olsson
- , Kimmo Kartasalo
- & Martin Eklund
-
Article
| Open Access Inferring time-varying generation time, serial interval, and incubation period distributions for COVID-19
Inferring time-varying generation time, serial interval, and incubation period distributions for COVID-19
The generation time (interval between successive infections in a transmission chain) is an important parameter for epidemiological modeling. Here, the authors develop a framework for estimating this parameter and how it changes over time and apply it to data from China in the first months of the pandemic.
- Dongxuan Chen
- , Yiu-Chung Lau
- & Sheikh Taslim Ali
-
Article
| Open Access Causal inference in medical records and complementary systems pharmacology for metformin drug repurposing towards dementia
Causal inference in medical records and complementary systems pharmacology for metformin drug repurposing towards dementia
Previous observational studies of the diabetes drugs metformin vs. sulfonylureas have yielded mixed results about whether metformin reduces the risk of dementia, relative to the sulfonylureas. Here, the authors apply a novel competing risks approach to emulate dementia-related target trials in electronic health records of diabetic patients and a complementary systems pharmacology evaluation on human neural cells.
- Marie-Laure Charpignon
- , Bella Vakulenko-Lagun
- & Mark W. Albers
-
Article
| Open Access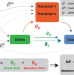 Quantifying the role of transcript levels in mediating DNA methylation effects on complex traits and diseases
Quantifying the role of transcript levels in mediating DNA methylation effects on complex traits and diseases
The mechanism by which DNA methylation might affect complex traits is not well understood. Here, the authors use Mendelian randomization to reveal a substantial role of transcript levels in mediating DNA methylation effects on complex traits and diseases.
- Marie C. Sadler
- , Chiara Auwerx
- & Zoltán Kutalik
-
Article
| Open Access Spatially aware dimension reduction for spatial transcriptomics
Spatially aware dimension reduction for spatial transcriptomics
Spatial transcriptomics analyses can be affected by noise and spatial correlation across tissue locations. Here, the authors develop SpatialPCA, a spatially-aware dimensionality reduction method that explicitly models spatial correlation structures, and show its application to the analysis of healthy and tumour tissues.
- Lulu Shang
- & Xiang Zhou
-
Article
| Open Access Leveraging data-driven self-consistency for high-fidelity gene expression recovery
Leveraging data-driven self-consistency for high-fidelity gene expression recovery
Recovering dropout-affected gene expression values is a challenging problem in bioinformatics. Here, the authors propose a data-driven framework, that first learns the underlying data distribution and then recovers the expression values by imposing a self-consistency on the expression matrix.
- Md Tauhidul Islam
- , Jen-Yeu Wang
- & Lei Xing
-
Article
| Open Access Integrating transcription factor occupancy with transcriptome-wide association analysis identifies susceptibility genes in human cancers
Integrating transcription factor occupancy with transcriptome-wide association analysis identifies susceptibility genes in human cancers
Transcriptome-wide association studies can uncover genes involved in disease. Here, the authors extend the framework with a transcriptome-wide association study approach which incorporates transcription factor occupancy, adding tissue-specific mechanistic support to associations.
- Jingni He
- , Wanqing Wen
- & Xingyi Guo
-
Article
| Open Access Deep learning to decompose macromolecules into independent Markovian domains
Deep learning to decompose macromolecules into independent Markovian domains
Modeling the dynamics of large proteins reveals a fundamental scaling problem. Here, the authors tackle this challenge by decomposing a large system into smaller independent subsystems, simultaneously modeling each subsystem’s kinetics and ensuring their mutual independence.
- Andreas Mardt
- , Tim Hempel
- & Frank Noé
-
Article
| Open Access CLIMB: High-dimensional association detection in large scale genomic data
CLIMB: High-dimensional association detection in large scale genomic data
Comparisons among experimental results with large amounts of data can be more precise and meaningful when done across multiple different conditions simultaneously. Koch et al. introduce a method, called CLIMB, that does this, and captures interpretable and biologically meaningful information.
- Hillary Koch
- , Cheryl A. Keller
- & Qunhua Li
-
Article
| Open Access dcHiC detects differential compartments across multiple Hi-C datasets
dcHiC detects differential compartments across multiple Hi-C datasets
The organisation of mammalian genomes plays a role in many biological processes. Here the authors report dcHiC, a tool which uses a multivariate distance measure to identify changes in compartmentalisation among multiple genome-wide chromatin contact maps, and apply this to different human and mouse datasets.
- Abhijit Chakraborty
- , Jeffrey G. Wang
- & Ferhat Ay
-
Article
| Open Access UniTVelo: temporally unified RNA velocity reinforces single-cell trajectory inference
UniTVelo: temporally unified RNA velocity reinforces single-cell trajectory inference
RNA velocity can detect the differentiation directionality by modelling sparse unspliced RNAs, but suffers from high estimation errors. Here, the authors develop a computational method called UniTVelo to reinforce the velocity estimation by introducing a unified time and a top-down model design.
- Mingze Gao
- , Chen Qiao
- & Yuanhua Huang
-
Article
| Open Access Tempo: an unsupervised Bayesian algorithm for circadian phase inference in single-cell transcriptomics
Tempo: an unsupervised Bayesian algorithm for circadian phase inference in single-cell transcriptomics
Previous efforts to study the circadian clock using scRNA-seq have relied on time course designs that treat cell collection time as a proxy for circadian time. Here, the authors introduce a statistical method to infer circadian timing directly from expression, enabling researchers to study circadian phase heterogeneity.
- Benjamin J. Auerbach
- , Garret A. FitzGerald
- & Mingyao Li
-
Article
| Open Access Cost-effective methylome sequencing of cell-free DNA for accurately detecting and locating cancer
Cost-effective methylome sequencing of cell-free DNA for accurately detecting and locating cancer
Early cancer detection by cell-free DNA (cfDNA) is challenged by the low amount of tumour DNA in cfDNA, tumour heterogeneity and the small patient cohorts. Here, the authors develop a method, cfMethyl-Seq, for cost-effective methylome profiling of cfDNA and for detecting and locating cancer.
- Mary L. Stackpole
- , Weihua Zeng
- & Xianghong Jasmine Zhou
-
Article
| Open Access Tracking changes in SARS-CoV-2 transmission with a novel outpatient sentinel surveillance system in Chicago, USA
Tracking changes in SARS-CoV-2 transmission with a novel outpatient sentinel surveillance system in Chicago, USA
In this study, the authors develop a method for estimation of SARS-CoV-2 community transmission rates based on a sentinel population of people seeking outpatient testing with recent symptom onset. This method has fewer operational delays than methods based on hospital data, and may be subject to fewer biases.
- Reese Richardson
- , Emile Jorgensen
- & Jaline Gerardin
-
Article
| Open Access Efficient and accurate frailty model approach for genome-wide survival association analysis in large-scale biobanks
Efficient and accurate frailty model approach for genome-wide survival association analysis in large-scale biobanks
The proliferation of large biobanks necessitates statistical methods designed for genetic analysis on biobank data. Here, the authors have developed a frailty model-based method for GWAS analysis of time-to-event phenotypes in large biobanks that accounts for relatedness in samples and censoring of phenotypes.
- Rounak Dey
- , Wei Zhou
- & Xihong Lin
-
Article
| Open Access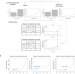 Batch effects removal for microbiome data via conditional quantile regression
Batch effects removal for microbiome data via conditional quantile regression
Here, the authors present ConQuR, a conditional quantile regression method that removes microbiome batch effects through non-parametric modeling of complex microbial read counts, while preserving the signals of interest.
- Wodan Ling
- , Jiuyao Lu
- & Michael C. Wu
-
Article
| Open Access A blind benchmark of analysis tools to infer kinetic rate constants from single-molecule FRET trajectories
A blind benchmark of analysis tools to infer kinetic rate constants from single-molecule FRET trajectories
The ability to infer quantitative kinetic information from single-molecule FRET (smFRET) data can be challenging. Here the authors perform a blind benchmark study assessing different analysis tools used to infer kinetic rate constants from smFRET trajectories, testing on simulated and experimental data.
- Markus Götz
- , Anders Barth
- & Sonja Schmid
-
Article
| Open Access The impact of repeated rapid test strategies on the effectiveness of at-home antiviral treatments for SARS-CoV-2
The impact of repeated rapid test strategies on the effectiveness of at-home antiviral treatments for SARS-CoV-2
Antiviral treatments for SARS-CoV-2 infection are only beneficial when used early in infection, so early case detection is important. Here, the authors assess the frequency of testing needed to achieve population-level benefits and demonstrate the importance of high coverage and short delays from test to treatment.
- Tigist F. Menkir
- & Christl A. Donnelly
-
Article
| Open Access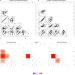 Linear and nonlinear correlation estimators unveil undescribed taxa interactions in microbiome data
Linear and nonlinear correlation estimators unveil undescribed taxa interactions in microbiome data
Here, the authors present Sparse Estimation of Correlations among Microbiomes (SECOM), a tool devised to characterize both linear and nonlinear relationships in microbiome data. When applied to human skin and infant gut microbiome data, SECOM is able to retrieve taxa interactions undescribed by previous methods.
- Huang Lin
- , Merete Eggesbø
- & Shyamal Das Peddada
-
Article
| Open Access Differential analysis of RNA structure probing experiments at nucleotide resolution: uncovering regulatory functions of RNA structure
Differential analysis of RNA structure probing experiments at nucleotide resolution: uncovering regulatory functions of RNA structure
The authors present DiffScan, an advanced tool for normalization and differential analysis of RNA structure probing experiments, combining their power in deciphering the dynamic RNA structurome and facilitating the discovery of RNA regulatory functions.
- Bo Yu
- , Pan Li
- & Lin Hou
-
Article
| Open Access Deep learning from phylogenies to uncover the epidemiological dynamics of outbreaks
Deep learning from phylogenies to uncover the epidemiological dynamics of outbreaks
Widely applicable, accurate and fast inference methods in phylodynamics are needed to fully profit from the richness of genetic data in uncovering the dynamics of epidemics. Here, the authors develop a likelihood-free, simulation-based deep learning approach.
- J. Voznica
- , A. Zhukova
- & O. Gascuel
-
Article
| Open Access Forest Fire Clustering for single-cell sequencing combines iterative label propagation with parallelized Monte Carlo simulations
Forest Fire Clustering for single-cell sequencing combines iterative label propagation with parallelized Monte Carlo simulations
In the era of single-cell sequencing, there is a growing need to extract insights from data with clustering methods. Here, inspired by forest fire dynamics, the authors devise an algorithm that can cluster single-cell data with minimal prior assumptions and can compute a non-parametric posterior probability for each data point.
- Zhanlin Chen
- , Jeremy Goldwasser
- & Mark Gerstein
-
Article
| Open Access Identifying colorectal cancer caused by biallelic MUTYH pathogenic variants using tumor mutational signatures
Identifying colorectal cancer caused by biallelic MUTYH pathogenic variants using tumor mutational signatures
Germline biallelic pathogenic MUTYH variants predispose patients to colorectal cancer (CRC); however, approaches to identify MUTYH variant carriers are lacking. Here, the authors evaluated mutational signatures that could distinguish MUTYH carriers in large CRC cohorts, and found MUTYH-associated somatic mutations.
- Peter Georgeson
- , Tabitha A. Harrison
- & Daniel D. Buchanan
-
Article
| Open Access Estimating tumor mutational burden from RNA-sequencing without a matched-normal sample
Estimating tumor mutational burden from RNA-sequencing without a matched-normal sample
The identification of somatic point mutations in tumor samples is of high clinical value, such as for the development of targeted therapies. Here the authors develop a machine learning pipeline for detecting somatic point mutations from RNA sequencing without a matched-normal sample, and utilize the model's prediction for computing the tumor mutational burden.
- Rotem Katzir
- , Noam Rudberg
- & Keren Yizhak

 Atlas-scale single-cell multi-sample multi-condition data integration using scMerge2
Atlas-scale single-cell multi-sample multi-condition data integration using scMerge2