Featured
-
-
Article |
 Isomerization of BRCA1–BARD1 promotes replication fork protection
Isomerization of BRCA1–BARD1 promotes replication fork protection
BRCA1–BARD1 has a role in replication fork protection that is mediated by a mechanism of phosphorylation-targeted isomerization of BRCA1 and is independent of the canonical interaction between BRCA1 and PALB2.
- Manuel Daza-Martin
- , Katarzyna Starowicz
- & Joanna R. Morris
-
Letter |
 High speed of fork progression induces DNA replication stress and genomic instability
High speed of fork progression induces DNA replication stress and genomic instability
Inhibition of PARP is shown to accelerate the speed of replication fork elongation, which prevents fork stalling and induces DNA damage, with implications for genomic instability and cancer treatment.
- Apolinar Maya-Mendoza
- , Pavel Moudry
- & Jiri Bartek
-
Article |
 SAMHD1 acts at stalled replication forks to prevent interferon induction
SAMHD1 acts at stalled replication forks to prevent interferon induction
SAMHD1 has an essential role in the replication stress response and prevents inflammation by activating the MRE11 nuclease to degrade nascent DNA strands at stalled replication forks, thus enabling replication.
- Flavie Coquel
- , Maria-Joao Silva
- & Philippe Pasero
-
Article |
 Replication fork stability confers chemoresistance in BRCA-deficient cells
Replication fork stability confers chemoresistance in BRCA-deficient cells
Protection of nascent DNA from degradation provides a mechanism that can promote synthetic viability and drug resistance in Brca-deficient cells without restoring homologous recombination at double-strand breaks.
- Arnab Ray Chaudhuri
- , Elsa Callen
- & André Nussenzweig
-
Letter |
 BRCA1 controls homologous recombination at Tus/Ter-stalled mammalian replication forks
BRCA1 controls homologous recombination at Tus/Ter-stalled mammalian replication forks
Direct evidence for the role of BRCA1 in controlling homologous recombination at stalled replication forks has been obtained in mammalian cells using the bacterial Tus/Ter system.
- Nicholas A. Willis
- , Gurushankar Chandramouly
- & Ralph Scully
-
Letter |
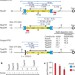 Recombination-restarted replication makes inverted chromosome fusions at inverted repeats
Recombination-restarted replication makes inverted chromosome fusions at inverted repeats
A new mechanism of chromosomal rearrangement is identified through the observation that broken or collapsed DNA replication forks restarted by homologous recombination have a high propensity for U-turns at short inverted repeats; the error-prone nature of this mechanism is suggested to contribute to gross chromosomal rearrangements and copy-number variations present in cancer and other genomic disorders.
- Ken’Ichi Mizuno
- , Izumi Miyabe
- & Johanne M. Murray

 Alcohol-derived DNA crosslinks are repaired by two distinct mechanisms
Alcohol-derived DNA crosslinks are repaired by two distinct mechanisms