Article
|
Open Access
Featured
-
-
Article
| Open Access replicAnt: a pipeline for generating annotated images of animals in complex environments using Unreal Engine
replicAnt: a pipeline for generating annotated images of animals in complex environments using Unreal Engine
Deep learning-based computer vision tools are transforming animal behavioural research; however, many challenges remain. Here, Plum et al. present replicAnt, a novel tool for generating synthetic data to train computer vision models for animal behaviour studies, reducing the need for manual annotation.
- Fabian Plum
- , René Bulla
- & David Labonte
-
Article
| Open Access Systematic analysis of paralogous regions in 41,755 exomes uncovers clinically relevant variation
Systematic analysis of paralogous regions in 41,755 exomes uncovers clinically relevant variation
Chameleolyser enables the accurate identification of genetic variants hidden within complex regions of the genome. Its application uncovers the disease-explanatory variant in 25 previously undiagnosed patients.
- Wouter Steyaert
- , Lonneke Haer-Wigman
- & Christian Gilissen
-
Article
| Open Access Identification of errors in draft genome assemblies at single-nucleotide resolution for quality assessment and improvement
Identification of errors in draft genome assemblies at single-nucleotide resolution for quality assessment and improvement
A high-quality genome assembly is essential for various genomic studies in life sciences. Here the authors develop CRAQ, a reference-free method that facilitates the evaluation and improvement of any de novo genome assembly with single nucleotide resolution.
- Kunpeng Li
- , Peng Xu
- & Yuannian Jiao
-
Article
| Open Access Deep-LASI: deep-learning assisted, single-molecule imaging analysis of multi-color DNA origami structures
Deep-LASI: deep-learning assisted, single-molecule imaging analysis of multi-color DNA origami structures
Analysis of single-molecule experiments remains time-consuming and prone to human bias. Here, the authors propose Deep-Learning Assisted Single-molecule Imaging analysis, a tool to rapidly analyse single-, two- and three-color single-molecule FRET data.
- Simon Wanninger
- , Pooyeh Asadiatouei
- & Don C. Lamb
-
Article
| Open Access Streamlined structure determination by cryo-electron tomography and subtomogram averaging using TomoBEAR
Streamlined structure determination by cryo-electron tomography and subtomogram averaging using TomoBEAR
Cryo-electron tomography (cryo-ET) enables structural analysis of molecules in situ, but the process is demanding. Here, authors report a software package, TomoBEAR, that streamlines data processing yielding high resolution structures with minimal user input.
- Nikita Balyschew
- , Artsemi Yushkevich
- & Mikhail Kudryashev
-
Article
| Open Access Benchmarking strategies for cross-species integration of single-cell RNA sequencing data
Benchmarking strategies for cross-species integration of single-cell RNA sequencing data
The growing number of available single-cell RNA-sequencing datasets from different species creates opportunities to explore evolutionary relationships between cell types across species. Here, the authors compare different strategies for cross-species integration of these data and offer guidelines for effective integration.
- Yuyao Song
- , Zhichao Miao
- & Irene Papatheodorou
-
Article
| Open Access Predicting discrete-time bifurcations with deep learning
Predicting discrete-time bifurcations with deep learning
Critical transitions and qualitative changes of dynamics in cardiac, ecological, and economical systems, can be characterized by discrete-time bifurcations. The authors propose a deep learning framework that provides early warning signals for critical transitions in discrete-time experimental data.
- Thomas M. Bury
- , Daniel Dylewsky
- & Gil Bub
-
Article
| Open Access Spectroscape enables real-time query and visualization of a spectral archive in proteomics
Spectroscape enables real-time query and visualization of a spectral archive in proteomics
Proteomics data repositories are deluged with data that is scarcely reused. Here, the authors developed Spectroscape, an interactive web-based tool for efficient similarity search of a query spectrum against a repository-scale spectral archive, and real-time visualization of its neighborhood.
- Long Wu
- , Ayman Hoque
- & Henry Lam
-
Article
| Open Access MetaCC allows scalable and integrative analyses of both long-read and short-read metagenomic Hi-C data
MetaCC allows scalable and integrative analyses of both long-read and short-read metagenomic Hi-C data
The authors develop an integrative and scalable framework to eliminate systematic biases and retrieve high-quality metagenome-assembled genomes using either long-read or short-read metagenomic Hi-C data.
- Yuxuan Du
- & Fengzhu Sun
-
Article
| Open Access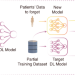 A unified method to revoke the private data of patients in intelligent healthcare with audit to forget
A unified method to revoke the private data of patients in intelligent healthcare with audit to forget
Revoking personal private data is one of the basic human rights. Here, the authors show AFS, a unified method to revoke patients’ private data from pre-trained deep learning models.
- Juexiao Zhou
- , Haoyang Li
- & Xin Gao
-
Article
| Open Access Ictogenesis proceeds through discrete phases in hippocampal CA1 seizures in mice
Ictogenesis proceeds through discrete phases in hippocampal CA1 seizures in mice
Predicting seizure onsets may allow for seizure prevention in patients. Here, authors show two distinct phases that always preceded temporal lobe seizures in mice, with activity confined within these two phases failing to progress into a seizure.
- John-Sebastian Mueller
- , Fabio C. Tescarollo
- & Hai Sun
-
Article
| Open Access TAGET: a toolkit for analyzing full-length transcripts from long-read sequencing
TAGET: a toolkit for analyzing full-length transcripts from long-read sequencing
Accurate long-read RNA sequencing facilitates analysis of full-length transcripts. Here the authors develop an integrative toolkit, optimised for Iso-Seq data analysis, that includes transcript alignment, annotation, quantification and gene fusion detection.
- Yuchao Xia
- , Zijie Jin
- & Ruibin Xi
-
Article
| Open Access epiAneufinder identifies copy number alterations from single-cell ATAC-seq data
epiAneufinder identifies copy number alterations from single-cell ATAC-seq data
'Here the authors present epiAneufinder, an algorithm for the identification of single-cell copy number alterations from scATAC-seq data, and explore the clonal heterogeneity in cell populations.
- Akshaya Ramakrishnan
- , Aikaterini Symeonidi
- & Maria Colomé-Tatché
-
Article
| Open Access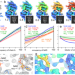 A robust normalized local filter to estimate compositional heterogeneity directly from cryo-EM maps
A robust normalized local filter to estimate compositional heterogeneity directly from cryo-EM maps
Heterogeneity in structural biology data includes potentially valuable information about binding and dynamics. Here, the authors devise, validate and demonstrate a method to quantify local heterogeneity in 3D reconstructions.
- Björn O. Forsberg
- , Pranav N. M. Shah
- & Alister Burt
-
Article
| Open Access Context-dependent perturbations in chromatin folding and the transcriptome by cohesin and related factors
Context-dependent perturbations in chromatin folding and the transcriptome by cohesin and related factors
Enhancer–promoter looping and topologically associating domain are at the base of chromatin structures. Here the authors present a computational workflow in which multi-omics datasets are compared systematically to explore how three-dimensional (3D) structure and gene expression are regulated by cohesin and related factors.
- Ryuichiro Nakato
- , Toyonori Sakata
- & Katsuhiko Shirahige
-
Article
| Open Access Performance of tumour microenvironment deconvolution methods in breast cancer using single-cell simulated bulk mixtures
Performance of tumour microenvironment deconvolution methods in breast cancer using single-cell simulated bulk mixtures
Multiple computational approaches have been developed for the deconvolution of cells in the tumour microenvironment (TME) using bulk RNA-seq data. Here, the authors use breast cancer single-cell RNA-seq data to produce simulated bulk data, with which they compare the performance of nine TME deconvolution methods.
- Khoa A. Tran
- , Venkateswar Addala
- & Nicola Waddell
-
Article
| Open Access SiGra: single-cell spatial elucidation through an image-augmented graph transformer
SiGra: single-cell spatial elucidation through an image-augmented graph transformer
Recent advances have pushed spatial transcriptomics to subcellular resolution. Here, the authors propose SiGra, a graph artificial intelligence model designed for high-throughput spatial molecular imaging.
- Ziyang Tang
- , Zuotian Li
- & Qianqian Song
-
Article
| Open Access Deciphering complex breakage-fusion-bridge genome rearrangements with Ambigram
Deciphering complex breakage-fusion-bridge genome rearrangements with Ambigram
Breakage-fusion-bridge (BFB) is a mechanism that leads to complex genome rearrangements in multiple cancers. Here, the authors develop a computational method for identifying these events, even when further complicated by additional structural variations.
- Chaohui Li
- , Lingxi Chen
- & Shuai Cheng Li
-
Article
| Open Access A deep learning-based stripe self-correction method for stitched microscopic images
A deep learning-based stripe self-correction method for stitched microscopic images
Image stitching in fluorescence microscopy can be a hindrance to image quality and to downstream quantitative analyses. Here, the authors propose a deep learning-based stripe self-correction method that corrects diverse stripes and artifacts for stitched microscopic images.
- Shu Wang
- , Xiaoxiang Liu
- & Jianxin Chen
-
Article
| Open Access CRUSTY: a versatile web platform for the rapid analysis and visualization of high-dimensional flow cytometry data
CRUSTY: a versatile web platform for the rapid analysis and visualization of high-dimensional flow cytometry data
CRUSTY is an interactive webtool for flow cytometry data analysis, offering popular algorithms and visualizations, and generating publication-quality figures in minutes. It enables users without bioinformatics expertize to mine complex datasets, supports real-time exploration, and is freely available online.
- Simone Puccio
- , Giorgio Grillo
- & Enrico Lugli
-
Article
| Open Access The Oncology Biomarker Discovery framework reveals cetuximab and bevacizumab response patterns in metastatic colorectal cancer
The Oncology Biomarker Discovery framework reveals cetuximab and bevacizumab response patterns in metastatic colorectal cancer
Identifying actionable biomarkers remains a challenge. Here, the authors develop a framework Oncology Biomarker Discovery (OncoBird), apply it to a phase III trial and investigate the molecular and biomarker landscape of metastatic colorectal carcinoma patients.
- Alexander J. Ohnmacht
- , Arndt Stahler
- & Michael P. Menden
-
Article
| Open Access Interpreting biologically informed neural networks for enhanced proteomic biomarker discovery and pathway analysis
Interpreting biologically informed neural networks for enhanced proteomic biomarker discovery and pathway analysis
Deep neural networks hold significant promise in capturing the complexity of biological systems. However, they suffer from a lack of interpretability. Here, authors present a generalizable method for developing, interpreting, and visualizing biologically informed neural networks for proteomics data.
- Erik Hartman
- , Aaron M. Scott
- & Johan Malmström
-
Article
| Open Access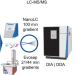 Deep and fast label-free Dynamic Organellar Mapping
Deep and fast label-free Dynamic Organellar Mapping
Regulated subcellular localization changes control protein function. Here, the authors provide a seamless spatial proteomics pipeline for mapping whole-cell protein localization dynamics, which includes a scalable workflow and a software suite for automated data analysis and visualization.
- Julia P. Schessner
- , Vincent Albrecht
- & Georg H. H. Borner
-
Article
| Open Access iCLOTS: open-source, artificial intelligence-enabled software for analyses of blood cells in microfluidic and microscopy-based assays
iCLOTS: open-source, artificial intelligence-enabled software for analyses of blood cells in microfluidic and microscopy-based assays
Microscopy has undoubtedly advanced biomedical research, but novel hypotheses are often lost to a lack of analytical tools. Here authors propose iCLOTS, a freely-available software that allows researchers to apply image processing and artificial intelligence algorithms to their own data.
- Meredith E. Fay
- , Oluwamayokun Oshinowo
- & Wilbur A. Lam
-
Article
| Open Access A deep learning method for replicate-based analysis of chromosome conformation contacts using Siamese neural networks
A deep learning method for replicate-based analysis of chromosome conformation contacts using Siamese neural networks
Siamese neural networks are a powerful deep learning approach for image analysis. Here, the authors adapt this method to the replicate-based analysis of Hi-C data and find that it successfully discriminates technical noise from biological variation.
- Ediem Al-jibury
- , James W. D. King
- & Daniel Rueckert
-
Article
| Open Access Characterizing cancer metabolism from bulk and single-cell RNA-seq data using METAFlux
Characterizing cancer metabolism from bulk and single-cell RNA-seq data using METAFlux
Metabolic reprogramming is a common indicator of the tumour microenvironment. Here the authors develop the METAflux framework to predict metabolic fluxes from single cell RNA-seq data.
- Yuefan Huang
- , Vakul Mohanty
- & Ken Chen
-
Article
| Open Access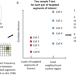 SnapFISH: a computational pipeline to identify chromatin loops from multiplexed DNA FISH data
SnapFISH: a computational pipeline to identify chromatin loops from multiplexed DNA FISH data
Multiplexed DNA FISH technologies are powerful tools to reveal chromatin spatial organisation. Here, the authors developed SnapFISH, a computational pipeline to identify chromatin loops from multiplexed DNA FISH data.
- Lindsay Lee
- , Hongyu Yu
- & Ming Hu
-
Article
| Open Access Cell-type-specific co-expression inference from single cell RNA-sequencing data
Cell-type-specific co-expression inference from single cell RNA-sequencing data
Inferring co-expressions with scRNA-seq data is challenging, and existing methods suffer from inflated false positives and biases. Here, the authors proposed CS-CORE, which yields unbiased estimates and identifies co-expressions that are more reproducible and biologically relevant for scRNA-seq data.
- Chang Su
- , Zichun Xu
- & Jingfei Zhang
-
Article
| Open Access Segmenting functional tissue units across human organs using community-driven development of generalizable machine learning algorithms
Segmenting functional tissue units across human organs using community-driven development of generalizable machine learning algorithms
Constructing the human reference atlas requires integration and analysis of massive amounts of data. Here the authors report the setup and results of the Hacking the Human Body machine learning algorithm development competition hosted by the Human Biomolecular Atlas and the Human Protein Atlas teams.
- Yashvardhan Jain
- , Leah L. Godwin
- & Katy Börner
-
Article
| Open Access Guided construction of single cell reference for human and mouse lung
Guided construction of single cell reference for human and mouse lung
Accurate cell-type identification is vital for single-cell analysis. Here, the authors develop a computational pipeline called “LungMAP CellRef” for efficient, automated cell-type annotation of normal and disease human and mouse lung single-cell datasets.
- Minzhe Guo
- , Michael P. Morley
- & Yan Xu
-
Article
| Open Access MSBooster: improving peptide identification rates using deep learning-based features
MSBooster: improving peptide identification rates using deep learning-based features
There is a need for accessible ways to improve peptide spectrum match rescoring with deep learning predictions in bottom-up proteomics. Here, the authors demonstrate robust gains in peptide/protein identifications across various experiments, from single cell proteomics to immunopeptidomics.
- Kevin L. Yang
- , Fengchao Yu
- & Alexey I. Nesvizhskii
-
Article
| Open Access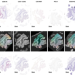 Virtual alignment of pathology image series for multi-gigapixel whole slide images
Virtual alignment of pathology image series for multi-gigapixel whole slide images
The spatial organization of a tumor affects how it grows and responds to treatment. Here, the authors present VALIS, a software to align sets of whole slide images (WSI) with state-of-the-art accuracy, enabling spatial studies of the tumor ecology.
- Chandler D. Gatenbee
- , Ann-Marie Baker
- & Alexander R. A. Anderson
-
Article
| Open Access Identification of transcriptional programs using dense vector representations defined by mutual information with GeneVector
Identification of transcriptional programs using dense vector representations defined by mutual information with GeneVector
In single-cell RNA-seq analyses, it would be critical to measure the relationships between genes. Here, the authors develop a framework for single-cell dimensionality reduction that incorporates gene-specific relationships - GeneVector -, and use it for tasks such as annotating cell types and analysing pathway variation after treatment.
- Nicholas Ceglia
- , Zachary Sethna
- & Andrew McPherson
-
Article
| Open Access Atlas-scale single-cell multi-sample multi-condition data integration using scMerge2
Atlas-scale single-cell multi-sample multi-condition data integration using scMerge2
Recent advances in multi-condition single-cell multi-cohort studies enable exploration of diverse cell states. Here, authors present scMerge2, an algorithm that allows integration of a large COVID-19 data collection with over five million cells to uncover distinct signatures of disease progression.
- Yingxin Lin
- , Yue Cao
- & Jean Y. H. Yang
-
Article
| Open Access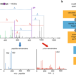 Detecting diagnostic features in MS/MS spectra of post-translationally modified peptides
Detecting diagnostic features in MS/MS spectra of post-translationally modified peptides
Protein modifications increase the complexity of data analysis in mass spectrometry-based proteomics, which may impair the comprehensive mapping of modification sites. Here, the authors develop an algorithm to extract diagnostic fragmentation patterns to improve modified peptide recovery and localization.
- Daniel J. Geiszler
- , Daniel A. Polasky
- & Alexey I. Nesvizhskii
-
Article
| Open Access Multi-batch single-cell comparative atlas construction by deep learning disentanglement
Multi-batch single-cell comparative atlas construction by deep learning disentanglement
Comparing single-cell RNA-seq and ATAC-seq data from multiple batches is challenging due to technical artifacts. Here, the authors propose a method that disentangles technical and biological effects, facilitating batch-confounded chromatin and gene expression state discovery and enhancing the analysis of perturbation effects on cell populations.
- Allen W. Lynch
- , Myles Brown
- & Clifford A. Meyer
-
Article
| Open Access Analysis of DIA proteomics data using MSFragger-DIA and FragPipe computational platform
Analysis of DIA proteomics data using MSFragger-DIA and FragPipe computational platform
DIA-MS has emerged as a widely used technological platform for quantitative protein profiling. Here, the authors develop MSFragger-DIA, a robust and fast tool to directly identify peptides from DIA spectra. It demonstrates excellent performance across applications from large-scale tumor studies to single-cell proteomics.
- Fengchao Yu
- , Guo Ci Teo
- & Alexey I. Nesvizhskii
-
Article
| Open Access Trackable and scalable LC-MS metabolomics data processing using asari
Trackable and scalable LC-MS metabolomics data processing using asari
Reproducible and scalable data processing is key to the progress of metabolomics. Here, the authors present a software tool that offers predictable metabolomics feature detection and improved computational performance in large datasets.
- Shuzhao Li
- , Amnah Siddiqa
- & Shujian Zheng
-
Article
| Open Access High throughput single cell long-read sequencing analyses of same-cell genotypes and phenotypes in human tumors
High throughput single cell long-read sequencing analyses of same-cell genotypes and phenotypes in human tumors
There is a need for methods that allow the analysis of single-cell long-read sequencing data without depending on known barcode lists or short-read sequencing. Here, the authors develop scNanoGPS, a tool that can independently deconvolute long reads into single cells and single molecules, and apply it on tumour and cell line data.
- Cheng-Kai Shiau
- , Lina Lu
- & Ruli Gao
-
Article
| Open Access nnSVG for the scalable identification of spatially variable genes using nearest-neighbor Gaussian processes
nnSVG for the scalable identification of spatially variable genes using nearest-neighbor Gaussian processes
The identification of top spatially variable genes is a key step in the analysis of spatially-resolved transcriptomics data. Here, the authors develop a scalable method based on nearest-neighbor Gaussian processes and evaluate performance compared to existing and baseline methods.
- Lukas M. Weber
- , Arkajyoti Saha
- & Stephanie C. Hicks
-
Article
| Open Access Glycopeptide database search and de novo sequencing with PEAKS GlycanFinder enable highly sensitive glycoproteomics
Glycopeptide database search and de novo sequencing with PEAKS GlycanFinder enable highly sensitive glycoproteomics
Accurate identification of intact glycopeptides from mass spectrometry data is essential for the characterization of glycosylation events in biological samples. Here, the authors propose GlycanFinder, a database search and de novo sequencing tool for the analysis of intact glycopeptides.
- Weiping Sun
- , Qianqiu Zhang
- & Baozhen Shan
-
Article
| Open Access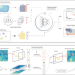 Leveraging spatial transcriptomics data to recover cell locations in single-cell RNA-seq with CeLEry
Leveraging spatial transcriptomics data to recover cell locations in single-cell RNA-seq with CeLEry
Cell location information is important for understanding how tissue is spatially organized. Here, the authors develop CeLEry, a machine learning method that aims to recover cell locations for single-cell RNA-seq data by leveraging information learned from spatial transcriptomics.
- Qihuang Zhang
- , Shunzhou Jiang
- & Mingyao Li
-
Article
| Open Access Detecting temporal and spatial malaria patterns from first antenatal care visits
Detecting temporal and spatial malaria patterns from first antenatal care visits
Pregnant people visiting antenatal clinics may represent a useful sentinel surveillance population for monitoring infections such as malaria. Here, the authors investigate the potential of this approach by comparing malaria prevalence in pregnant people and children living in the same area of southern Mozambique.
- Arnau Pujol
- , Nanna Brokhattingen
- & Alfredo Mayor
-
Article
| Open Access CAJAL enables analysis and integration of single-cell morphological data using metric geometry
CAJAL enables analysis and integration of single-cell morphological data using metric geometry
Cell morphology is one of the most described phenotypes in biology, yet systematic quantification and classification of morphology remains limited. Here, the authors present a computational approach for cell morphometry and multi-modal analysis based on concepts from metric geometry.
- Kiya W. Govek
- , Patrick Nicodemus
- & Pablo G. Camara
-
Article
| Open Access grandR: a comprehensive package for nucleotide conversion RNA-seq data analysis
grandR: a comprehensive package for nucleotide conversion RNA-seq data analysis
Nucleotide conversion approaches facilitate metabolic RNA labelling experiments but complicate computational analysis. Here, the authors develop a methodology and software package to enable specific analysis methods for nucleotide conversion RNA-seq data.
- Teresa Rummel
- , Lygeri Sakellaridi
- & Florian Erhard
-
Article
| Open Access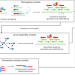 INSurVeyor: improving insertion calling from short read sequencing data
INSurVeyor: improving insertion calling from short read sequencing data
Current methods for detecting insertions from short read sequencing data generally have low sensitivity. Here, the authors develop a new tool that runs quickly and detects significantly more true positive insertions compared to any combination of existing methods.
- Ramesh Rajaby
- , Dong-Xu Liu
- & Wing-Kin Sung
-
Article
| Open Access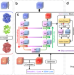 Improvement of cryo-EM maps by simultaneous local and non-local deep learning
Improvement of cryo-EM maps by simultaneous local and non-local deep learning
Map post-processing is crucial for cryo-EM modeling building. Here, the authors present a deep learning approach to improve both the quality and interpretability of cryo-EM maps by simultaneously considering local and non-local effects.
- Jiahua He
- , Tao Li
- & Sheng-You Huang
-
Article
| Open Access Inference of cell type-specific gene regulatory networks on cell lineages from single cell omic datasets
Inference of cell type-specific gene regulatory networks on cell lineages from single cell omic datasets
Cell type-specific gene expression patterns are outputs of transcriptional gene regulatory networks (GRNs) that connect transcription factors and signaling proteins to target genes. Here, the authors present single-cell Multi-Task Network Inference (scMTNI), a multi-task learning framework to infer cell type-specific GRN dynamics from scRNA-seq and scATAC-seq datasets collected for diverse cell fate specification trajectories.
- Shilu Zhang
- , Saptarshi Pyne
- & Sushmita Roy
-
Article
| Open Access ExpressAnalyst: A unified platform for RNA-sequencing analysis in non-model species
ExpressAnalyst: A unified platform for RNA-sequencing analysis in non-model species
RNA-sequencing data analysis is difficult for non-model species that have no reference genome. ExpressAnalyst enables RNA-sequencing analysis for any eukaryotic species in less than 24 h, on a laptop, and without any programming.
- Peng Liu
- , Jessica Ewald
- & Jianguo Xia

 EASTR: Identifying and eliminating systematic alignment errors in multi-exon genes
EASTR: Identifying and eliminating systematic alignment errors in multi-exon genes