Article
|
Open Access
Featured
-
-
Article
| Open Access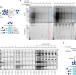 Signal-noise metrics for RNA binding protein identification reveal broad spectrum protein-RNA interaction frequencies and dynamics
Signal-noise metrics for RNA binding protein identification reveal broad spectrum protein-RNA interaction frequencies and dynamics
The identification of RNA-bound proteomes is hampered by a lack of quantitative metrics for evaluating RNA binding function. Here, the authors report LEAP-RBP as a method for purification of RNA-bound proteins and introduce signal-based metrics for robust profiling of RNA-bound proteomes.
- JohnCarlo Kristofich
- & Christopher V. Nicchitta
-
Article
| Open Access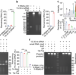 Discovery of the major 15–30 nt mammalian small RNAs, their biogenesis and function
Discovery of the major 15–30 nt mammalian small RNAs, their biogenesis and function
The authors uncover the major 15-30 nt mouse and human small RNAs (sRNAs), which mainly end with 2’,3’-cyclic phosphate, and show that many of these sRNAs can be generated by Angiogenin or RNase 4 and function in Ago2 complex as miRNAs.
- Hejin Lai
- , Ning Feng
- & Qiwei Zhai
-
Article
| Open Access The master energy homeostasis regulator PGC-1α exhibits an mRNA nuclear export function
The master energy homeostasis regulator PGC-1α exhibits an mRNA nuclear export function
PGC-1α is a master regulator activating the transcription of key genes controlling the cell’s energy production. Here the authors show that PGC-1α has a function in the NXF1-dependent nuclear export of mRNAs.
- Simeon R. Mihaylov
- , Lydia M. Castelli
- & Guillaume M. Hautbergue
-
Article
| Open Access Dynamically regulated two-site interaction of viral RNA to capture host translation initiation factor
Dynamically regulated two-site interaction of viral RNA to capture host translation initiation factor
RNA viruses use elements of their genomic RNA to commandeer the host translational machinery. Here, the authors use NMR and cryo-EM to reveal the sophisticated strategy by which a viral RNA engages host translational factors in a dynamically regulated two-site interaction.
- Shunsuke Imai
- , Hiroshi Suzuki
- & Ichio Shimada
-
Article
| Open Access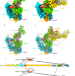 Structural mechanism of R2D2 and Loqs-PD synergistic modulation on DmDcr-2 oligomers
Structural mechanism of R2D2 and Loqs-PD synergistic modulation on DmDcr-2 oligomers
R2D2 and Loqs-PD are cofactors of Drosophila Dicer-2 (DmDcr-2), which generates siRNAs. Here the authors report the cryo-EM structures of DmDcr-2/R2D2/Loqs-PD with dsRNAs showing that these complexes can form oligomers and assemble into fibers.
- Ting Deng
- , Shichen Su
- & Jia Wang
-
Article
| Open Access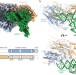 Structural basis for a degenerate tRNA identity code and the evolution of bimodal specificity in human mitochondrial tRNA recognition
Structural basis for a degenerate tRNA identity code and the evolution of bimodal specificity in human mitochondrial tRNA recognition
Aminoacyl-tRNA synthetases catalyze the ligation of amino acids to their cognate tRNAs. Here the authors report the cryo-EM structure of a human mitochondrial seryl-tRNA synthetase•mtRNASer complex showing how strong mutation pressure on mtRNA genes drove a rewiring of intermolecular recognition rules.
- Bernhard Kuhle
- , Marscha Hirschi
- & Paul Schimmel
-
Article
| Open Access Modulation of translational decoding by m6A modification of mRNA
Modulation of translational decoding by m6A modification of mRNA
m6A is an mRNA modification that slows down translation elongation. Here, Jain et al. show that m6A delays decoding and increases tRNA drop-off from the ribosome by favoring alternative codon conformations that are rejected by the ribosome.
- Sakshi Jain
- , Lukasz Koziej
- & Marina V. Rodnina
-
Article
| Open Access Molecular basis of the pleiotropic effects by the antibiotic amikacin on the ribosome
Molecular basis of the pleiotropic effects by the antibiotic amikacin on the ribosome
Here the authors use fast kinetics, X-ray crystallography, and cryo-EM to uncover the mechanism of ribosome inhibition by amikacin and kanamycin. They find that amikacin binds near the P-site tRNA, offering new strategies to fight antibiotic resistance.
- Savannah M. Seely
- , Narayan P. Parajuli
- & Matthieu G. Gagnon
-
Article
| Open Access Structural basis for specific RNA recognition by the alternative splicing factor RBM5
Structural basis for specific RNA recognition by the alternative splicing factor RBM5
The RNA binding protein RBM5 regulates alternative splicing of genes implicated in cancer. Here the authors show structural mechanisms how multiple RNA binding domains of RBM5 cooperate to recognize specific target RNA sequences.
- Komal Soni
- , Pravin Kumar Ankush Jagtap
- & Michael Sattler
-
Article
| Open Access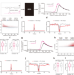 Exon-intron boundary inhibits m6A deposition, enabling m6A distribution hallmark, longer mRNA half-life and flexible protein coding
Exon-intron boundary inhibits m6A deposition, enabling m6A distribution hallmark, longer mRNA half-life and flexible protein coding
m6A mRNA modification is not typically found near splice junctions in mRNAs. Here the authors show exon-intron boundary inhibits m6A deposition at ~100 nt region nearby splice site, enabling m6A distribution hallmark, more stable mRNA and flexible protein coding.
- Zhiyuan Luo
- , Qilian Ma
- & Shengdong Ke
-
Article
| Open Access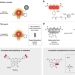 Fast-exchanging spirocyclic rhodamine probes for aptamer-based super-resolution RNA imaging
Fast-exchanging spirocyclic rhodamine probes for aptamer-based super-resolution RNA imaging
Live-cell RNA imaging with high spatial and temporal resolution remains a major challenge. Here the authors design spirocyclic rhodamine probes that enable a fluorescent light-up aptamer system suitable for visualizing RNAs in live or fixed cells with two different super-resolution microscopy modalities SMLM and STED.
- Daniel Englert
- , Eva-Maria Burger
- & Murat Sunbul
-
Article
| Open Access The mRNA methyltransferase Mettl3 modulates cytokine mRNA stability and limits functional responses in mast cells
The mRNA methyltransferase Mettl3 modulates cytokine mRNA stability and limits functional responses in mast cells
The m6A mRNA modification is essential for immune cell function. Here, the Monticelli lab optimized methods of gene editing by CRISPR-Cas9 in mast cells and revealed how the m6A machinery is required to sustain proliferation and to limit the production of inflammatory cytokines by these cells.
- Cristina Leoni
- , Marian Bataclan
- & Silvia Monticelli
-
Article
| Open Access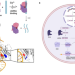 An unnatural enzyme with endonuclease activity towards small non-coding RNAs
An unnatural enzyme with endonuclease activity towards small non-coding RNAs
Endonucleases play crucial roles in various biological processes but endonucleases that target small non-coding RNAs have not been reported. Here, the authors combined the metal binding non-canonical amino acid BpyAla and a high affinity binder to engineer a catalyst that degrades small non-coding RNAs.
- Noreen Ahmed
- , Nadine Ahmed
- & John Paul Pezacki
-
Article
| Open Access Co-translational binding of importins to nascent proteins
Co-translational binding of importins to nascent proteins
Importins are known to facilitate nucleocytoplasmic transport and cytoplasmic chaperoning of some proteins. Here, the authors uncover that these proteins also act as co-translational chaperones for specific sets of proteins, for example ribonucleic acid binding factors.
- Maximilian Seidel
- , Natalie Romanov
- & Martin Beck
-
Article
| Open Access Systematic detection of tertiary structural modules in large RNAs and RNP interfaces by Tb-seq
Systematic detection of tertiary structural modules in large RNAs and RNP interfaces by Tb-seq
Compact RNA structural motifs control many aspects of gene expression, but methods for their identification are lacking. Here the authors present a sequencing-based terbium probing approach to detect complex 3D structural elements, which can be used to pinpoint potential riboregulatory elements.
- Shivali Patel
- , Alec N. Sexton
- & Anna Marie Pyle
-
Article
| Open Access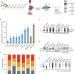 In vivo PAR-CLIP (viP-CLIP) of liver TIAL1 unveils targets regulating cholesterol synthesis and secretion
In vivo PAR-CLIP (viP-CLIP) of liver TIAL1 unveils targets regulating cholesterol synthesis and secretion
Here the authors develop viP-CLIP, a technique allowing the identification of RNA binding proteins (RBP) targets in mouse tissues, and identify the RBP Tial1 as a regulator of cholesterol metabolism in the liver.
- Hasan Vatandaslar
- , Aitor Garzia
- & Markus Stoffel
-
Article
| Open Access The preference signature of the SARS-CoV-2 Nucleocapsid NTD for its 5’-genomic RNA elements
The preference signature of the SARS-CoV-2 Nucleocapsid NTD for its 5’-genomic RNA elements
SARS-CoV-2 genome turnover is mediated by its N protein, but precise parameters driving the necessary RNA specificity have remained enigmatic. Here, Korn et al. reveal N’s N-terminal domain to distinguish regulatory viral RNA motifs with a preference for transiently folded elements of functional impact.
- Sophie Marianne Korn
- , Karthikeyan Dhamotharan
- & Andreas Schlundt
-
Article
| Open Access Intracellular RNA and DNA tracking by uridine-rich internal loop tagging with fluorogenic bPNA
Intracellular RNA and DNA tracking by uridine-rich internal loop tagging with fluorogenic bPNA
Commonly used protein-based tools to monitor intracellular RNA and DNA can impact steric accessibility and native nucleic acid biology. Here, the authors show that fluorogenic uridine-rich internal loop tagging bPNA probes can be used to label nucleic acids in fixed and live cells.
- Yufeng Liang
- , Sydney Willey
- & Dennis Bong
-
Article
| Open Access Co-crystal structures of the fluorogenic aptamer Beetroot show that close homology may not predict similar RNA architecture
Co-crystal structures of the fluorogenic aptamer Beetroot show that close homology may not predict similar RNA architecture
The recently discovered aptamer Beetroot is a homodimeric RNA that binds and activates DFAME, a conditional, red-shifted fluorophore derived from GFP. Here the authors determine the Beetroot-DFAME co-crystal structure, which is distinctively different from that of similar RNA aptamer Corn.
- Luiz F. M. Passalacqua
- , Mary R. Starich
- & Adrian R. Ferré-D’Amaré
-
Article
| Open Access Cap analogs with a hydrophobic photocleavable tag enable facile purification of fully capped mRNA with various cap structures
Cap analogs with a hydrophobic photocleavable tag enable facile purification of fully capped mRNA with various cap structures
Removing immunogenic uncapped mRNA from transcribed mRNA can be challenging, but is critical in mRNA research and clinical applications such as vaccines. Here, authors develop hydrophobic photocaged tag-modified cap analogs, which can be used to separate capped mRNA from uncapped mRNA, with subsequent tag removal using photo-irradiation.
- Masahito Inagaki
- , Naoko Abe
- & Hiroshi Abe
-
Article
| Open Access Observation of structural switch in nascent SAM-VI riboswitch during transcription at single-nucleotide and single-molecule resolution
Observation of structural switch in nascent SAM-VI riboswitch during transcription at single-nucleotide and single-molecule resolution
Observation of single nascent RNA at single-nucleotide resolution enables building a conformation landscape of how RNAs change during transcription. Here the authors develop a method to monitor the conformational change of individual SAM-VI riboswitch RNA (riboSAM) during transcription at single-nucleotide resolution as it is synthesized by transcription.
- Yanyan Xue
- , Jun Li
- & Yu Liu
-
Article
| Open Access RNA binding protein SYNCRIP maintains proteostasis and self-renewal of hematopoietic stem and progenitor cells
RNA binding protein SYNCRIP maintains proteostasis and self-renewal of hematopoietic stem and progenitor cells
Hematopoietic stem cells (HSCs) are essential for long-term repopulation after secondary transplantation. Here they show that SYNCRIP safeguards HSC self-renewal during regenerative stress by maintaining both proteostasis and CDC42-regulated cell polarity.
- Florisela Herrejon Chavez
- , Hanzhi Luo
- & Michael G. Kharas
-
Article
| Open Access Copy number variation in tRNA isodecoder genes impairs mammalian development and balanced translation
Copy number variation in tRNA isodecoder genes impairs mammalian development and balanced translation
Enigmatically tRNA genes exist in several hundred copies in mammalian genomes. Here the authors find a precipitous failure of development and increased amino acid misincorporation upon systematic elimination of tRNA-Phe genes in mice using CRISPR.
- Laetitia A. Hughes
- , Danielle L. Rudler
- & Aleksandra Filipovska
-
Article
| Open Access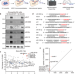 Screening of Drosophila microRNA-degradation sequences reveals Argonaute1 mRNA’s role in regulating miR-999
Screening of Drosophila microRNA-degradation sequences reveals Argonaute1 mRNA’s role in regulating miR-999
miRNAs typically bind to target mRNAs to induce their degradation but when base-pairing between miRNAs and the target RNAs are extended, miRNAs themselves can be degraded. Here, by employing AGO1-CLASH in Drosophila cells, the authors show that RNA sequence in AGO1 mRNA 3′UTR induces decay of miR-999.
- Peike Sheng
- , Lu Li
- & Mingyi Xie
-
Article
| Open Access Crystal structure of a highly conserved enteroviral 5′ cloverleaf RNA replication element
Crystal structure of a highly conserved enteroviral 5′ cloverleaf RNA replication element
A cloverleaf-like RNA domain within the enterovirus genome is essential for replication. Here, the authors determine the 1.9 Å resolution crystal structure of such RNA from coxsackievirus B3 – a model enterovirus to study many other human viruses.
- Naba K. Das
- , Nele M. Hollmann
- & Deepak Koirala
-
Article
| Open Access Bombyx Vasa sequesters transposon mRNAs in nuage via phase separation requiring RNA binding and self-association
Bombyx Vasa sequesters transposon mRNAs in nuage via phase separation requiring RNA binding and self-association
Bombyx Vasa assembles Vasa bodies, the site of transposon silencing by Siwi and Ago3-piRISC formation. Here, the authors show Vasa sequesters transposon mRNAs in Vasa bodies via phase separation requiring RNA binding and self-association of Vasa.
- Hiroya Yamazaki
- , Yurika Namba
- & Mikiko C. Siomi
-
Article
| Open Access Multi-modal quantification of pathway activity with MAYA
Multi-modal quantification of pathway activity with MAYA
Pathways can be activated through various signaling cascades depending on cell type. Here, the authors introduce MAYA, a computational method that can detect and score multiple modes of activation for each pathway, improving the granularity of pathway analysis for single-cell datasets.
- Yuna Landais
- & Céline Vallot
-
Article
| Open Access Snapshots of the second-step self-splicing of Tetrahymena ribozyme revealed by cryo-EM
Snapshots of the second-step self-splicing of Tetrahymena ribozyme revealed by cryo-EM
The Tetrahymena ribozyme changes conformations to perform its self-splicing function. Here, the authors capture six structural snapshots of ribozyme during the second step of self-splicing from a single specimen, revealing the structural basis of how it promotes and coordinates self-splicing reactions.
- Shanshan Li
- , Michael Z. Palo
- & Kaiming Zhang
-
Article
| Open Access Well-TEMP-seq as a microwell-based strategy for massively parallel profiling of single-cell temporal RNA dynamics
Well-TEMP-seq as a microwell-based strategy for massively parallel profiling of single-cell temporal RNA dynamics
Gene expression of cells is a heterogeneous and dynamic program involved in various biological processes. Here, authors develop Well-TEMPseq, a high-throughput, cost-effective, and accurate method for massively parallel profiling of the temporal dynamics of single-cell gene expression.
- Shichao Lin
- , Kun Yin
- & Chaoyong Yang
-
Article
| Open Access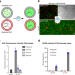 Periodic temperature changes drive the proliferation of self-replicating RNAs in vesicle populations
Periodic temperature changes drive the proliferation of self-replicating RNAs in vesicle populations
How primordial cells could achieve inheritance of encapsulated components is still an open question. Here, the authors show that ribozymes can assemble in active forms and replicate in populations of membrane vesicles thanks to freeze-thaw cycles.
- Elia Salibi
- , Benedikt Peter
- & Hannes Mutschler
-
Article
| Open Access Evaluating native-like structures of RNA-protein complexes through the deep learning method
Evaluating native-like structures of RNA-protein complexes through the deep learning method
RNA-protein docking is a very challenging area. Here, the authors develop a deep-learning based method, DRPScore, to evaluate RNA-protein complexes. DRPScore is robust and consistently performs better than existing methods on representative testing sets.
- Chengwei Zeng
- , Yiren Jian
- & Yunjie Zhao
-
Article
| Open Access Mechanisms of the RNA helicases DDX42 and DDX46 in human U2 snRNP assembly
Mechanisms of the RNA helicases DDX42 and DDX46 in human U2 snRNP assembly
The U2 snRNP recognizes the intron during spliceosome assembly. Here the authors report the high-resolution structures of two assembly precursors of human U2 snRNP and reveal that the RNA helicases DDX42 and DDX46 are mutually exclusive in terms of SF3B1-binding.
- Fenghua Yang
- , Tong Bian
- & Yigong Shi
-
Article
| Open Access Cryo-EM captures early ribosome assembly in action
Cryo-EM captures early ribosome assembly in action
The production of ribosomes is a precisely orchestrated energy consuming cellular process of highest priority. Here, the authors use cryo-EM to show that bacterial ribosomal subunits, self-assembled from their purified RNA and protein components, mature along parallel pathways.
- Bo Qin
- , Simon M. Lauer
- & Rainer Nikolay
-
Article
| Open Access Visualizing orthogonal RNAs simultaneously in live mammalian cells by fluorescence lifetime imaging microscopy (FLIM)
Visualizing orthogonal RNAs simultaneously in live mammalian cells by fluorescence lifetime imaging microscopy (FLIM)
No multi-color RNA fluorescent tags are currently available for use in live cells. Here, the authors show that fluorescence lifetime imaging microscopy is advantageous for multiplexed RNA visualization while achieving robust cellular contrast.
- Nadia Sarfraz
- , Emilia Moscoso
- & Esther Braselmann
-
Article
| Open Access Chemoproteomic discovery of a human RNA ligase
Chemoproteomic discovery of a human RNA ligase
RNA ligases are present across all forms of life. Here, the hitherto uncharacterised human protein C12orf29 was identified as a human enzyme promoting RNA ligation between 5′-PO4 and 3′-OH termini. This data provides the groundwork for establishing a human RNA repair pathway.
- Yizhi Yuan
- , Florian M. Stumpf
- & Andreas Marx
-
Article
| Open Access Visualizing RNA conformational and architectural heterogeneity in solution
Visualizing RNA conformational and architectural heterogeneity in solution
RNA conformational heterogeneity is important to diverse functions. Here, the authors use AFM to directly visualize individual RNA molecules that are in various conformational states under near physiological solution conditions for the first time.
- Jienyu Ding
- , Yun-Tzai Lee
- & Yun-Xing Wang
-
Article
| Open Access Two distinct binding modes provide the RNA-binding protein RbFox with extraordinary sequence specificity
Two distinct binding modes provide the RNA-binding protein RbFox with extraordinary sequence specificity
Here the authors show that the RRM of RbFox accomplishes extraordinary sequence specificity by employing functionally and structurally distinct binding modes - one for its cognate RNA and one for all non-cognate RNAs.
- Xuan Ye
- , Wen Yang
- & Fan Yang
-
Article
| Open Access Reanalysis of ribosome profiling datasets reveals a function of rocaglamide A in perturbing the dynamics of translation elongation via eIF4A
Reanalysis of ribosome profiling datasets reveals a function of rocaglamide A in perturbing the dynamics of translation elongation via eIF4A
The compound Rocaglamide A (RocA) is known for repressing translation initiation. Here the authors identify a dual mode of action for RocA in blocking translation initiation and elongation via eIF4A using previous datasets and new analyses.
- Fajin Li
- , Jianhuo Fang
- & Xuerui Yang
-
Article
| Open Access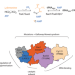 A paralog of Pcc1 is the fifth core subunit of the KEOPS tRNA-modifying complex in Archaea
A paralog of Pcc1 is the fifth core subunit of the KEOPS tRNA-modifying complex in Archaea
Many eukaryotic and archaeal tRNAs carry a modified adenosine (t6A) that is synthesized by the KEOPS complex, which is composed of four subunits. A fifth subunit (Gon7) is found only in fungi and metazoa. Here the authors show that archaea also possess a fifth subunit, which is structurally and functionally similar to eukaryotic Gon7.
- Marie-Claire Daugeron
- , Sophia Missoury
- & Tamara Basta
-
Article
| Open Access Preclinical development of kinetin as a safe error-prone SARS-CoV-2 antiviral able to attenuate virus-induced inflammation
Preclinical development of kinetin as a safe error-prone SARS-CoV-2 antiviral able to attenuate virus-induced inflammation
The search for antivirals against SARS-CoV-2 continue due to the emergence of variants of concerns, able to escape the vaccinal humoral response. In this work, authors pre-clinically explore the potential of kinetin against SARS-CoV-2, which could be used alone or in combination with other antivirals.
- Thiago Moreno L. Souza
- , Vagner D. Pinho
- & Jaime A. Rabi
-
Article
| Open Access Decoding of the ubiquitin code for clearance of colliding ribosomes by the RQT complex
Decoding of the ubiquitin code for clearance of colliding ribosomes by the RQT complex
The colliding ribosomes are ubiquitinated by the sensor protein Hel2, leading to noncanonical subunit dissociation by the ribosome associated quality control trigger (RQT) complex. Here the authors reveal the decoding mechanism of the ribosome ubiquitin code by the RQT complex.
- Yoshitaka Matsuo
- , Takayuki Uchihashi
- & Toshifumi Inada
-
Article
| Open Access Targeted systematic evolution of an RNA platform neutralizing DNMT1 function and controlling DNA methylation
Targeted systematic evolution of an RNA platform neutralizing DNMT1 function and controlling DNA methylation
Here the authors generate an RNA-based platform to neutralize the major epigenetic player DNMT1. Using this targeted approach, aberrant DNA methylation in cancer can be corrected.
- Carla L. Esposito
- , Ida Autiero
- & Annalisa Di Ruscio
-
Article
| Open Access Translation factor eIF5a is essential for IFNγ production and cell cycle regulation in primary CD8+ T lymphocytes
Translation factor eIF5a is essential for IFNγ production and cell cycle regulation in primary CD8+ T lymphocytes
The translational elongation factor eIF5a may have a function in T cell protein translation. Here the authors show that hypusination of eIF5a or deletion of eIF5a alters the profile of mRNA translation in T cells and that eIF5 regulates the proliferation and effector function of these T cells.
- Thomas C. J. Tan
- , Van Kelly
- & Rose Zamoyska
-
Article
| Open Access Broad misappropriation of developmental splicing profile by cancer in multiple organs
Broad misappropriation of developmental splicing profile by cancer in multiple organs
The molecular mechanisms underlying the overlap between oncogenic and embryonic development remain to be explored. Here, the authors use temporal transcriptomic data during development in multiple human organs and suggest the involvement of alternative splicing events, splicing factors, and transcription factors.
- Arashdeep Singh
- , Arati Rajeevan
- & Sridhar Hannenhalli
-
Article
| Open Access DAP5 enables main ORF translation on mRNAs with structured and uORF-containing 5′ leaders
DAP5 enables main ORF translation on mRNAs with structured and uORF-containing 5′ leaders
RNA structure and upstream ORFs can modulate the translation of human transcripts. Here, the authors show that structured mRNAs with pervasive uORF translation require DAP5, an eIF4G-like protein, for translation of the main ORF.
- Ramona Weber
- , Leon Kleemann
- & Cátia Igreja
-
Article
| Open Access Altered tRNA processing is linked to a distinct and unusual La protein in Tetrahymena thermophila
Altered tRNA processing is linked to a distinct and unusual La protein in Tetrahymena thermophila
La proteins are conserved factors critical for the maturation of RNA polymerase III transcripts. In the ciliate T. thermophila and related alveolates, La proteins have a novel domain arrangement and are linked to a distinct pre-tRNA processing pathway.
- Kyra Kerkhofs
- , Jyoti Garg
- & Mark A. Bayfield
-
Article
| Open Access Structural basis for recognition of transcriptional terminator structures by ProQ/FinO domain RNA chaperones
Structural basis for recognition of transcriptional terminator structures by ProQ/FinO domain RNA chaperones
In this work, the authors determine the crystal structure of a ProQ/FinO RNA chaperone bound to its RNA target. This provides insight into how this family of bacterial proteins recognize transcriptional terminator structures.
- Hyeong Jin Kim
- , Mazzen Black
- & J. N. Mark Glover
-
Article
| Open Access Structure of the IscB–ωRNA ribonucleoprotein complex, the likely ancestor of CRISPR-Cas9
Structure of the IscB–ωRNA ribonucleoprotein complex, the likely ancestor of CRISPR-Cas9
The cryo-EM structure of the IscB–ωRNA–target DNA complex provide insights into the ωRNA-guided DNA cleavage mechanism by IscB and the evolution of the type II CRISPR-Cas9 effector complexes
- Kazuki Kato
- , Sae Okazaki
- & Hiroshi Nishimasu
-
Article
| Open Access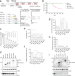 RNA-binding proteins of KHDRBS and IGF2BP families control the oncogenic activity of MLL-AF4
RNA-binding proteins of KHDRBS and IGF2BP families control the oncogenic activity of MLL-AF4
The establishment of a mouse model of MLL-AF4-induced leukaemia is a challenge as the introduction of fusion gene of human MLL-AF4 does not cause leukaemia in mice. Here the authors reveal that MLL-AF4 is post-transcriptionally regulated by RNA-binding proteins that inhibits MLL-AF4 translation, thus, hampering MLL-AF4-mediated leukemic transformation.
- Hiroshi Okuda
- , Ryo Miyamoto
- & Akihiko Yokoyama

 Epitranscriptomic subtyping, visualization, and denoising by global motif visualization
Epitranscriptomic subtyping, visualization, and denoising by global motif visualization