Featured
-
-
Article
| Open Access Single-molecule, full-length transcript isoform sequencing reveals disease-associated RNA isoforms in cardiomyocytes
Single-molecule, full-length transcript isoform sequencing reveals disease-associated RNA isoforms in cardiomyocytes
Alternative splicing generates RNA isoforms that contribute to phenotypic diversity. Here the authors perform single-molecule full-length RNA sequencing to identify disease-associated variant transcript isoforms.
- Chenchen Zhu
- , Jingyan Wu
- & Lars M. Steinmetz
-
Article
| Open Access TSSC4 is a component of U5 snRNP that promotes tri-snRNP formation
TSSC4 is a component of U5 snRNP that promotes tri-snRNP formation
The correct assembly and recycling of the multicomponent spliceosome remains largely elusive. Here, the authors show that a previously uncharacterized protein TSSC4 associates with de novo formed spliceosomal U5 snRNP as well as with a post-splicing U5-PRPF19 particle, and that TSSC4 is important for assembly of the splicing competent tri-snRNP.
- Klára Klimešová
- , Jitka Vojáčková
- & David Staněk
-
Article
| Open Access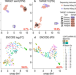 MOCCASIN: a method for correcting for known and unknown confounders in RNA splicing analysis
MOCCASIN: a method for correcting for known and unknown confounders in RNA splicing analysis
Confounding factors on gene expression analysis can be analyzed by several existing tools. Here the authors develop an algorithm called MOCCASIN to correct the effect of known and unknown confounders on RNA splicing quantification.
- Barry Slaff
- , Caleb M. Radens
- & Yoseph Barash
-
Article
| Open Access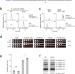 A single m6A modification in U6 snRNA diversifies exon sequence at the 5’ splice site
A single m6A modification in U6 snRNA diversifies exon sequence at the 5’ splice site
Spliceosomal U6 snRNA is modified by m6A. Here the authors show that m6A modification on U6 snRNA promotes efficient recognition of the splice site through cooperating with U5 snRNA, indicating that U6 snRNA m6A allows the 5’ exon sequence variation.
- Yuma Ishigami
- , Takayuki Ohira
- & Tsutomu Suzuki
-
Article
| Open Access The splicing factor XAB2 interacts with ERCC1-XPF and XPG for R-loop processing
The splicing factor XAB2 interacts with ERCC1-XPF and XPG for R-loop processing
XPA-binding protein (XAB)-2 is the human homologue of the yeast pre-mRNA splicing factor Syf1. Here the authors use an in vivo biotinylation tagging approach to show XAB2’s role in DNA repair, RNA splicing and transcription during mammalian development.
- Evi Goulielmaki
- , Maria Tsekrekou
- & George A. Garinis
-
Article
| Open Access Activation of Prp28 ATPase by phosphorylated Npl3 at a critical step of spliceosome remodeling
Activation of Prp28 ATPase by phosphorylated Npl3 at a critical step of spliceosome remodeling
Yeast helicase Prp28 promotes the first step of spliceosome remodeling. By placing a photoactivatable unnatural amino acid in Prp28, the authors capture Prp28 in action revealing its dynamic interactions and cofactor Npl3.
- Fu-Lung Yeh
- , Shang-Lin Chang
- & Tien-Hsien Chang
-
Article
| Open Access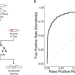 Hotspot exons are common targets of splicing perturbations
Hotspot exons are common targets of splicing perturbations
Splicing-disrupting mutations are linked to diseases. By employing a machine learning approach, the authors show that certain exons, termed hotspot exons, are enriched for splicing-disruption variants and susceptible to splicing perturbations.
- David T. Glidden
- , Jeramiah L. Buerer
- & William G. Fairbrother
-
Article
| Open Access The RNA binding protein FgRbp1 regulates specific pre-mRNA splicing via interacting with U2AF23 in Fusarium
The RNA binding protein FgRbp1 regulates specific pre-mRNA splicing via interacting with U2AF23 in Fusarium
Human RBM42 associates with the spliceosome complex. Here the authors show that the fungus counterpart of RBM42, FgRbp1 regulates splicing by interacting with FgU2AF23.
- Minhui Wang
- , Tianling Ma
- & Zhonghua Ma
-
Article
| Open Access Conserved long-range base pairings are associated with pre-mRNA processing of human genes
Conserved long-range base pairings are associated with pre-mRNA processing of human genes
Functional RNA secondary structure is important for the pre-mRNA processing including splicing, cleavage and polyadenylation, and RNA editing. Here the authors present a catalog of conserved long-range RNA structures in the human transcriptome by defining pairs of conserved complementary regions (PCCR) in pre-aligned evolutionarily conserved regions.
- Svetlana Kalmykova
- , Marina Kalinina
- & Dmitri Pervouchine
-
Article
| Open Access Role of Hakai in m6A modification pathway in Drosophila
Role of Hakai in m6A modification pathway in Drosophila
Drosophila m6A writer complex regulates alternative splicing of the Sex-lethal gene. Here the authors show that a potential E3 ligase Hakai interacts with the fly m6A writer complex and that m6A level is reduced in Hakai mutant flies.
- Yanhua Wang
- , Lifeng Zhang
- & Dong Yan
-
Article
| Open Access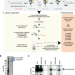 psiCLIP reveals dynamic RNA binding by DEAH-box helicases before and after exon ligation
psiCLIP reveals dynamic RNA binding by DEAH-box helicases before and after exon ligation
ATP-dependent helicases remodel the spliceosome and proofread splice site recognition. A new method – Purified Spliceosome iCLIP (psiCLIP) – probes protein-RNA interactions in defined spliceosome complexes to reveal how the helicases Prp16 and Prp22 promote correct mRNA synthesis through dynamic binding on their RNA substrates.
- Lisa M. Strittmatter
- , Charlotte Capitanchik
- & Kiyoshi Nagai
-
Article
| Open Access The methyltransferase SETD2 couples transcription and splicing by engaging mRNA processing factors through its SHI domain
The methyltransferase SETD2 couples transcription and splicing by engaging mRNA processing factors through its SHI domain
The methylation of Histone 3 at Lysine 36 (H3K36) has been implicated in the regulation of transcription and coupled processes such as mRNA splicing. Here the authors show that the histone methyltransferase SETD2 interacts with hnRNP L to mediate the crosstalk between the transcription and splicing machineries.
- Saikat Bhattacharya
- , Michaella J. Levy
- & Jerry L. Workman
-
Article
| Open Access Interaction of 7SK with the Smn complex modulates snRNP production
Interaction of 7SK with the Smn complex modulates snRNP production
The noncoding RNA 7SK controls the transcription of mRNAs. Here, the authors show that the 7SK complex interacts with the Smn complex, suggesting crosstalk between transcription and snRNP assembly.
- Changhe Ji
- , Jakob Bader
- & Michael Briese
-
Article
| Open Access Detection of aberrant splicing events in RNA-seq data using FRASER
Detection of aberrant splicing events in RNA-seq data using FRASER
Aberrant splicing is a major contributor to rare disease, but detection accuracy using current methods is limited. Here, the authors develop an algorithm that detects aberrant splicing and intron retention events from RNA-seq data and apply it to diagnosis in mitochondrial disease.
- Christian Mertes
- , Ines F. Scheller
- & Julien Gagneur
-
Article
| Open Access A spatially resolved brain region- and cell type-specific isoform atlas of the postnatal mouse brain
A spatially resolved brain region- and cell type-specific isoform atlas of the postnatal mouse brain
Alternative RNA splicing varies across the brain. Its mapping at single cell resolution is unclear. Here, the authors provide a spatial and single-cell splicing atlas reporting brain region- and cell type-specific expression of different isoforms in the postnatal mouse brain.
- Anoushka Joglekar
- , Andrey Prjibelski
- & Hagen U. Tilgner
-
Article
| Open Access Structure of SRSF1 RRM1 bound to RNA reveals an unexpected bimodal mode of interaction and explains its involvement in SMN1 exon7 splicing
Structure of SRSF1 RRM1 bound to RNA reveals an unexpected bimodal mode of interaction and explains its involvement in SMN1 exon7 splicing
SRSF1 is an oncoprotein that plays important roles in RNA metabolism. We reveal the structure of the human SRSF1 RRM1 bound to RNA, and propose a bimodal mode of interaction of the protein with RNA. A single mutation in RRM1 changed SRSF1 specificity for RNA and made it active on SMN2 exon7 splicing.
- Antoine Cléry
- , Miroslav Krepl
- & Frédéric H.-T. Allain
-
Article
| Open Access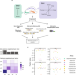 Differential contribution of transcriptomic regulatory layers in the definition of neuronal identity
Differential contribution of transcriptomic regulatory layers in the definition of neuronal identity
Post-transcriptional gene regulation is an important contributor to cell type-specific differences at the transcriptomic level. Here, the authors use a multiomics approach to characterize neuronal diversity in the mouse nervous system, analyzing the relative contributions of multiple layers of transcriptomic regulation in the specification of cell type identity.
- Kevin C. H. Ha
- , Timothy Sterne-Weiler
- & Benjamin J. Blencowe
-
Article
| Open Access STRAP regulates alternative splicing fidelity during lineage commitment of mouse embryonic stem cells
STRAP regulates alternative splicing fidelity during lineage commitment of mouse embryonic stem cells
STRAP (serine threonine kinase receptor-associated protein) promotes tumorigenicity. Here the authors report that STRAP associates with spliceosome and regulates alternative splicing during embryonic stem cell lineage commitment and early mouse embryo organogenesis.
- Lin Jin
- , Yunjia Chen
- & Pran K. Datta
-
Article
| Open Access Defective minor spliceosomes induce SMA-associated phenotypes through sensitive intron-containing neural genes in Drosophila
Defective minor spliceosomes induce SMA-associated phenotypes through sensitive intron-containing neural genes in Drosophila
Spinal muscular atrophy (SMA) is associated with minor splicing-related defects. Here the authors develop Drosophila models with minor spliceosomal-snRNA deletions, and demonstrate SMA-like phenotypes.
- Liang Li
- , Zhan Ding
- & Yong-Zhen Xu
-
Article
| Open Access Elucidation of the aberrant 3′ splice site selection by cancer-associated mutations on the U2AF1
Elucidation of the aberrant 3′ splice site selection by cancer-associated mutations on the U2AF1
U2AF1 binds to the 3’ splice site of introns and its mutation lead to abnormal splicing. Here the authors solve the crystal structures of wild type and pathogenic mutant U2AF1 bound to target RNA, showing that different target sequence is preferred by pathogenic mutant.
- Hisashi Yoshida
- , Sam-Yong Park
- & Eiji Obayashi
-
Article
| Open Access Antisense oligonucleotide modulation of non-productive alternative splicing upregulates gene expression
Antisense oligonucleotide modulation of non-productive alternative splicing upregulates gene expression
Restoration of normal gene expression is one way to treat monogenic disorders. Here the authors target naturally occurring non-productive alternative splicing using antisense oligonucleotides to promote the production of functional proteins.
- Kian Huat Lim
- , Zhou Han
- & Isabel Aznarez
-
Article
| Open Access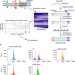 Comprehensive identification of mRNA isoforms reveals the diversity of neural cell-surface molecules with roles in retinal development and disease
Comprehensive identification of mRNA isoforms reveals the diversity of neural cell-surface molecules with roles in retinal development and disease
Here the authors present an approach that can reveal the full complement of mRNA isoforms encoded by individual genes, and they identify a major isoform of the retinal degeneration gene CRB1 which functions at the cell-cell junctions of the outer limiting membrane to promote photoreceptor survival.
- Thomas A. Ray
- , Kelly Cochran
- & Jeremy N. Kay
-
Article
| Open Access Intragenic recruitment of NF-κB drives splicing modifications upon activation by the oncogene Tax of HTLV-1
Intragenic recruitment of NF-κB drives splicing modifications upon activation by the oncogene Tax of HTLV-1
The nuclear factors κB (NF-κB) is a transcription factor involved in immune functions, inflammation, and cancer. Here, the authors show that the NF-κB factor RELA regulates splicing of target genes by recruiting DDX17 on chromatin upon expression of the viral oncogene Tax.
- Lamya Ben Ameur
- , Paul Marie
- & Franck Mortreux
-
Article
| Open Access A kinase-deficient NTRK2 splice variant predominates in glioma and amplifies several oncogenic signaling pathways
A kinase-deficient NTRK2 splice variant predominates in glioma and amplifies several oncogenic signaling pathways
Tropomyosin receptor kinase B (TrkB), encoded by the neurotrophic tyrosine receptor kinase 2 (NTRK2) gene, exhibits intricate splicing patterns and post-translational modifications. Here, the authors perform whole gene and transcript-level analyses and report the TrkB.T1 splice variant enhances PDGF-driven gliomas in vivo and augments PI3K signaling cascades in vitro.
- Siobhan S. Pattwell
- , Sonali Arora
- & Eric C. Holland
-
Article
| Open Access CRISPR artificial splicing factors
CRISPR artificial splicing factors
Control over splicing could be used for both therapeutic and engineering applications. Here the authors create artificial splicing factors using RNA-targeting CRISPR systems under small molecule control.
- Menghan Du
- , Nathaniel Jillette
- & Albert Wu Cheng
-
Article
| Open Access Mutational bias and the protein code shape the evolution of splicing enhancers
Mutational bias and the protein code shape the evolution of splicing enhancers
Splicing is regulated by cis-acting elements in pre-mRNAs such as exonic or intronic splicing enhancers and silencers. Here the authors show that exonic splicing enhancers are enriched in exons compared to introns due to mutational bias coupled with purifying selection on the protein code.
- Stephen Rong
- , Luke Buerer
- & William G. Fairbrother
-
Article
| Open Access High-resolution annotation of the mouse preimplantation embryo transcriptome using long-read sequencing
High-resolution annotation of the mouse preimplantation embryo transcriptome using long-read sequencing
Until now, the transcriptome of preimplantation mouse embryos has only been analysed by short-read sequencing. Here, the authors perform long-read sequencing to provide a more detailed transcriptome of the preimplantation mouse embryo, identifying various novel transcripts, for example Kdm4dl.
- Yunbo Qiao
- , Chao Ren
- & Wenjie Shu
-
Article
| Open Access ORF Capture-Seq as a versatile method for targeted identification of full-length isoforms
ORF Capture-Seq as a versatile method for targeted identification of full-length isoforms
Most human protein-coding genes are expressed as multiple isoforms. Here the authors present ORF Capture-seq that uses cloned ORFs as probes to capture and sequence full length transcript sequences. This enables highly sensitive characterization of eukaryotic transcriptomes.
- Gloria M. Sheynkman
- , Katharine S. Tuttle
- & Marc Vidal
-
Article
| Open Access Loss of MBNL1 induces RNA misprocessing in the thymus and peripheral blood
Loss of MBNL1 induces RNA misprocessing in the thymus and peripheral blood
The activity of the RNA splicing factor MBNL1 is altered in myotonic dystrophy (DM) patients. Here the authors characterize the thymic phenotype of Mbnl1 knockout mice, including developmental defects, transcriptome changes, and RNA mis-splicing of transcripts encoding thymic transcription factors.
- Łukasz J. Sznajder
- , Marina M. Scotti
- & Maurice S. Swanson
-
Article
| Open Access Full-length transcript characterization of SF3B1 mutation in chronic lymphocytic leukemia reveals downregulation of retained introns
Full-length transcript characterization of SF3B1 mutation in chronic lymphocytic leukemia reveals downregulation of retained introns
Long-read sequencing is useful in determining exon-connectivity of full-length mRNA isoforms. Here, by long-read nanopore sequencing, the authors report that intron retention is downregulated in SF3B1 mutant chronic lymphocytic leukemia cells than normal B cells.
- Alison D. Tang
- , Cameron M. Soulette
- & Angela N. Brooks
-
Article
| Open Access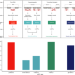 Regulatory sites for splicing in human basal ganglia are enriched for disease-relevant information
Regulatory sites for splicing in human basal ganglia are enriched for disease-relevant information
Regulation of gene expression and splicing are thought to be tissue-specific. Here, the authors obtain genomic and transcriptomic data from putamen and substantia nigra of 117 neurologically healthy human brains and find that splicing eQTLs are enriched for neuron-specific regulatory information.
- Sebastian Guelfi
- , Karishma D’Sa
- & Mina Ryten
-
Article
| Open Access The dynamic proteome of influenza A virus infection identifies M segment splicing as a host range determinant
The dynamic proteome of influenza A virus infection identifies M segment splicing as a host range determinant
Avian influenza A virus (IAV) strains replicate poorly in mammalian hosts, but mechanisms underlying species restriction are incompletely understood. Here, Bogdanow et al. show that avian and mammalian adapted IAV strains have evolved different RNA structure features for regulation of M segment RNA splicing.
- Boris Bogdanow
- , Xi Wang
- & Matthias Selbach
-
Article
| Open Access CRISPR-Cas9-based mutagenesis frequently provokes on-target mRNA misregulation
CRISPR-Cas9-based mutagenesis frequently provokes on-target mRNA misregulation
CRISPR-Cas9 genome editing is presumed to knock out gene function by generating a frameshift during NHEJ repair. Here, the authors investigate mRNA and protein expression in edited lines and find genome editing can generate internal ribosome entry sites or alternatively spliced variants.
- Rubina Tuladhar
- , Yunku Yeu
- & Lawrence Lum
-
Article
| Open Access Smu1 and RED are required for activation of spliceosomal B complexes assembled on short introns
Smu1 and RED are required for activation of spliceosomal B complexes assembled on short introns
Human spliceosome components Smu1 and RED regulate alternative splicing. Here the authors show that Smu1 and RED are also required for constitutive splicing of short introns.
- Sandra Keiper
- , Panagiotis Papasaikas
- & Reinhard Lührmann
-
Article
| Open Access Alternative splicing regulates stochastic NLRP3 activity
Alternative splicing regulates stochastic NLRP3 activity
Leucine-rich repeat (LRR) domains are commonly present in immune regulatory proteins. Here the authors show that LRR exonic modularity and alternative splicing of an LRR-containing protein, NLRP3, modulate the ratio of functional/afunctional NLRP3 isoforms to instill a stochastic regulation of NLRP3-mediated inflammation and innate immunity.
- Florian Hoss
- , James L. Mueller
- & Eicke Latz
-
Article
| Open Access Defining the genetic and evolutionary architecture of alternative splicing in response to infection
Defining the genetic and evolutionary architecture of alternative splicing in response to infection
Genetic ancestry might influence immunological response to infection at different regulatory levels. Here, the authors use RNA-Seq to investigate the variability of alternative splicing patterns in resting and stimulated monocytes of African- and European-descent.
- Maxime Rotival
- , Hélène Quach
- & Lluis Quintana-Murci
-
Article
| Open Access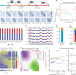 Dynamics of genome reorganization during human cardiogenesis reveal an RBM20-dependent splicing factory
Dynamics of genome reorganization during human cardiogenesis reveal an RBM20-dependent splicing factory
The spatial organization of the genome plays an important but unclearly defined role in gene regulation. Here, the authors integrate Hi-C, RNA-seq and ATAC-seq data to map cardiogenesis from pluripotent stem cells and describe an RBM20-dependent splicing factory assembling the TTN locus with other RBM20 targets.
- Alessandro Bertero
- , Paul A. Fields
- & Charles E. Murry
-
Article
| Open Access The splicing factor RBM25 controls MYC activity in acute myeloid leukemia
The splicing factor RBM25 controls MYC activity in acute myeloid leukemia
Splicing factors are often mutated in hematological malignancies. Here, the authors perform an in vivo shRNA screen in a CEBPA mutant AML mouse model and identify that RBM25 controls the splicing of pre-mRNAs encoding BCL-X and BIN1 to exert its tumour suppressor activities in AML.
- Ying Ge
- , Mikkel Bruhn Schuster
- & Bo Torben Porse
-
Article
| Open Access Non-invasive monitoring of alternative splicing outcomes to identify candidate therapies for myotonic dystrophy type 1
Non-invasive monitoring of alternative splicing outcomes to identify candidate therapies for myotonic dystrophy type 1
Myotonic dystrophy type 1 (DM1) is associated with aberrant transcript splicing. Here, the authors develop a transgenic mouse model expressing a bi-chromatic reporter system that allows non-invasive monitoring of splicing of a transcript altered in DM1 in vivo, and show that it allows for evaluation of the therapeutic response to treatment with antisense oligonucleotides.
- Ningyan Hu
- , Layal Antoury
- & Thurman M. Wheeler
-
Article
| Open Access SHQ1 regulation of RNA splicing is required for T-lymphoblastic leukemia cell survival
SHQ1 regulation of RNA splicing is required for T-lymphoblastic leukemia cell survival
T-acute lymphoblastic leukemia is an aggressive cancer. Here the authors provide insights into the functional role of SHQ1, an H/ACA snoRNP assembly factor involved in snRNA pseudouridylation, in T-lymphoblastic leukemia cell survival through regulating the maturation of MYC mRNA.
- Hexiu Su
- , Juncheng Hu
- & Hudan Liu
-
Article
| Open Access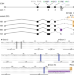 Genetic and mechanistic basis for APOBEC3H alternative splicing, retrovirus restriction, and counteraction by HIV-1 protease
Genetic and mechanistic basis for APOBEC3H alternative splicing, retrovirus restriction, and counteraction by HIV-1 protease
Human APOBEC3H has several haplotypes and splice variants with distinct anti-HIV-1 activities, but the genetics underlying the expression of these variants are unclear. Here, the authors identify an intronic deletion in A3H haplotype II resulting in production of the most active splice variant, which is counteracted by HIV-1 protease.
- Diako Ebrahimi
- , Christopher M. Richards
- & Reuben S. Harris
-
Article
| Open Access Regulatory mechanisms of incomplete huntingtin mRNA splicing
Regulatory mechanisms of incomplete huntingtin mRNA splicing
Incomplete splicing of HTT results in the production of the highly pathogenic exon 1 HTT protein. Here the authors identify the necessary intronic regions and the underlying mechanisms that contribute to this process.
- Andreas Neueder
- , Anaelle A. Dumas
- & Gillian P. Bates
-
Article
| Open Access Aberrant splicing and defective mRNA production induced by somatic spliceosome mutations in myelodysplasia
Aberrant splicing and defective mRNA production induced by somatic spliceosome mutations in myelodysplasia
Mutations to the splicing machinery may have an important role in myelodysplasia. Here, the authors describe splicing factor gene mutations in myelodysplasia and report tumor suppressor, epigenetic, iron metabolism and heme biosynthesis genes as their targets.
- Yusuke Shiozawa
- , Luca Malcovati
- & Mario Cazzola
-
Article
| Open Access Mutually exclusive acetylation and ubiquitylation of the splicing factor SRSF5 control tumor growth
Mutually exclusive acetylation and ubiquitylation of the splicing factor SRSF5 control tumor growth
Changes in glucose metabolism can lead to tumor development, but the involvement of splicing factors is unclear. Here, the authors screened for SR proteins and identified SRSF5 stability is enhanced in response to glucose elevation to promote alternative splicing of CCAR1 which facilitates tumor growth.
- Yuhan Chen
- , Qingyang Huang
- & Lingqiang Zhang
-
Article
| Open Access Co-regulatory activity of hnRNP K and NS1-BP in influenza and human mRNA splicing
Co-regulatory activity of hnRNP K and NS1-BP in influenza and human mRNA splicing
Alternative splicing of influenza A virus (IAV) M transcript is regulated by hnRNP K and NS1-BP, but mechanistic details are unknown. Here, Thompson et al. show how hnRNP K and NS1-BP bind M mRNA and that these proteins regulate splicing of host transcripts in both the absence and presence of IAV infection.
- Matthew G. Thompson
- , Raquel Muñoz-Moreno
- & Kristen W. Lynch
-
Article
| Open Access Molecular basis of differential 3′ splice site sensitivity to anti-tumor drugs targeting U2 snRNP
Molecular basis of differential 3′ splice site sensitivity to anti-tumor drugs targeting U2 snRNP
Several families of natural compounds target core components of the pre-mRNA splicing machinery and display anti-tumor activity. Here the authors show that particular sequence features can be linked to drug response, and that drugs with very similar chemical structures display substantially different effects on splicing regulation.
- Luisa Vigevani
- , André Gohr
- & Juan Valcárcel
-
Article
| Open Access CryoEM structure of Saccharomyces cerevisiae U1 snRNP offers insight into alternative splicing
CryoEM structure of Saccharomyces cerevisiae U1 snRNP offers insight into alternative splicing
U1 snRNP is critical for 5′ splicing site recognition in pre-mRNA splicing. Here the authors describe the cryo-EM structure of the yeast U1 snRNP and suggest that PrpF39 is an alternative splicing factor essential for the successful recruitment of U1 snRNP by other alternative splicing factors.
- Xueni Li
- , Shiheng Liu
- & Rui Zhao
-
Article
| Open Access Differential alternative splicing coupled to nonsense-mediated decay of mRNA ensures dietary restriction-induced longevity
Differential alternative splicing coupled to nonsense-mediated decay of mRNA ensures dietary restriction-induced longevity
Alternative splicing coupled to nonsense-mediated decay (AS-NMD) is a conserved mechanism for post-transcriptional gene regulation. Here, the authors provide evidence that AS-NMD is enhanced during dietary restriction (DR) and is required for DR-mediated longevity assurance in C. elegans.
- Syed Shamsh Tabrez
- , Ravi Datta Sharma
- & Arnab Mukhopadhyay
-
Article
| Open Access Functional and dynamic polymerization of the ALS-linked protein TDP-43 antagonizes its pathologic aggregation
Functional and dynamic polymerization of the ALS-linked protein TDP-43 antagonizes its pathologic aggregation
TDP-43 aggregation is observed in amyotrophic lateral sclerosis. Here the authors combine X-ray crystallography, nuclear magnetic resonance and electron microscopy studies and show that physiological oligomerization of TDP-43 is mediated through its N-terminal domain, which forms functional and dynamic oligomers antagonizing pathologic aggregation.
- Tariq Afroz
- , Eva-Maria Hock
- & Magdalini Polymenidou

 Therapeutic manipulation of IKBKAP mis-splicing with a small molecule to cure familial dysautonomia
Therapeutic manipulation of IKBKAP mis-splicing with a small molecule to cure familial dysautonomia