Featured
-
-
Article
| Open Access Proteomic characterization of epithelial ovarian cancer delineates molecular signatures and therapeutic targets in distinct histological subtypes
Proteomic characterization of epithelial ovarian cancer delineates molecular signatures and therapeutic targets in distinct histological subtypes
The molecular phenotypic features of epithelial ovarian cancer (EOC) remain elusive. Here, the authors perform mass spectrometry-based proteomic profiling for 269 EOC patients and reveal molecularly distinct features and potential therapeutic targets among the histological subtypes of EOC.
- Ting-Ting Gong
- , Shuang Guo
- & Qi-Jun Wu
-
Article
| Open Access Plasma proteomic profiles predict individual future health risk
Plasma proteomic profiles predict individual future health risk
The predictive capability of future health risk using plasma proteomic profiles remains largely unexplored. Using 1461 proteins collected from 50k individuals, authors show proteins can derive much better or equivalent performance than established clinical indicators for more than 40 endpoints.
- Jia You
- , Yu Guo
- & Jin-Tai Yu
-
Article
| Open Access Time-resolved proteomic profiling reveals compositional and functional transitions across the stress granule life cycle
Time-resolved proteomic profiling reveals compositional and functional transitions across the stress granule life cycle
Stress granules (SGs) are dynamic compartments with a poorly characterized transition in composition and function during prolonged stress. In this study, the authors investigated the dynamic changes in SG constituents during acute to prolonged heat shock using time-resolved proteomic profiling.
- Shuyao Hu
- , Yufeng Zhang
- & Yun Bai
-
Article
| Open Access Evaluation of circulating plasma proteins in breast cancer using Mendelian randomisation
Evaluation of circulating plasma proteins in breast cancer using Mendelian randomisation
Proteomics of blood samples is a promising avenue for cancer diagnosis. Here, the authors conduct Mendelian randomisation analysis of protein levels across multiple cohorts, and identify 5 proteins that show promise as biomarkers for the long-term risk of breast cancer, and as potential drug targets.
- Anders Mälarstig
- , Felix Grassmann
- & Åsa K. Hedman
-
Article
| Open Access Improved in situ characterization of protein complex dynamics at scale with thermal proximity co-aggregation
Improved in situ characterization of protein complex dynamics at scale with thermal proximity co-aggregation
Vast majority of cellular activities are carried out by protein complexes that assembled dynamically in response to cellular needs and environmental cues. Here, the authors present Slim-TPCA, an effective and readily deployable strategy to unravel the functional roles of protein complexes en masse across various cellular processes.
- Siyuan Sun
- , Zhenxiang Zheng
- & Chris Soon Heng Tan
-
Article
| Open Access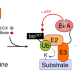 BioE3 identifies specific substrates of ubiquitin E3 ligases
BioE3 identifies specific substrates of ubiquitin E3 ligases
Here, the authors describe BioE3, a biotin-based method to discriminate direct substrates of ubiquitin E3 ligases of interest from mere interactors using proximity proteomics. BioE3 responds to chemical treatments, and works with RING- and HECT-type E3s, as well as ubiquitin-likes (e.g., SUMO).
- Orhi Barroso-Gomila
- , Laura Merino-Cacho
- & James D. Sutherland
-
Article
| Open Access Targetable lesions and proteomes predict therapy sensitivity through disease evolution in pediatric acute lymphoblastic leukemia
Targetable lesions and proteomes predict therapy sensitivity through disease evolution in pediatric acute lymphoblastic leukemia
The role of clonal evolution on the actionable proteome and response to therapy in childhood acute lymphoblastic leukemia (ALL) remains unknown. Here, targeted sequencing and proteomic analysis of paired ALL diagnosis and relapsed samples revealed PARP1 as a potential therapeutic target.
- Amanda C. Lorentzian
- , Jenna Rever
- & Philipp F. Lange
-
Article
| Open Access TOFIMS mass spectrometry-based immunopeptidomics refines tumor antigen identification
TOFIMS mass spectrometry-based immunopeptidomics refines tumor antigen identification
MS-based immunopeptidomics provides direct evidence for HLA peptide-antigen presentation, which is indispensable for therapeutic use. Here the authors present an ion mobility MS-based immunopeptidome workflow, largely expand benign reference databases and enables next generation tumor antigen discovery.
- Naomi Hoenisch Gravel
- , Annika Nelde
- & Juliane S. Walz
-
Article
| Open Access An adaptive stress response that confers cellular resilience to decreased ubiquitination
An adaptive stress response that confers cellular resilience to decreased ubiquitination
Hunt et al. identify the protein sets that are modulated by RNAi for each E2 ubiquitin-conjugating enzyme in human cells. By analyzing the UBA1/E2-sensitive proteome, they report an adaptive stress response that preserves peroxisomal protein import in cells with decreased ubiquitination capacity.
- Liam C. Hunt
- , Vishwajeeth Pagala
- & Fabio Demontis
-
Article
| Open Access Local membrane source gathering by p62 body drives autophagosome formation
Local membrane source gathering by p62 body drives autophagosome formation
Phase separated p62 body plays pivotal roles in autophagy. Here, the authors describe a spatial membrane gathering mode by which p62 body functions in autophagosome formation.
- Xuezhao Feng
- , Daxiao Sun
- & Na Mi
-
Article
| Open Access NULISA: a proteomic liquid biopsy platform with attomolar sensitivity and high multiplexing
NULISA: a proteomic liquid biopsy platform with attomolar sensitivity and high multiplexing
Unlocking the blood proteome requires exquisite sensitivity and multiplexing to detect low and high abundance proteins simultaneously. Here the authors describe a 200-plex immunoassay with attomolar sensitivity to detect important low abundance proteins in inflammatory diseases and COVID-19.
- Wei Feng
- , Joanne C. Beer
- & Xiao-Jun Ma
-
Article
| Open Access Spatially resolved mapping of proteome turnover dynamics with subcellular precision
Spatially resolved mapping of proteome turnover dynamics with subcellular precision
Mapping protein turnover dynamics with subcellular precision is crucial for understanding cell physiology and pathology. Here, the authors leveraged APEX2-mediated proximity labeling to develop prox-SILAC methods to profile protein turnover rates in the mitochondria and endoplasmic reticulum.
- Feng Yuan
- , Yi Li
- & Peng Zou
-
Article
| Open Access Multiple E3 ligases control tankyrase stability and function
Multiple E3 ligases control tankyrase stability and function
The poly(ADP-ribosyl)transferases, tankyrase 1 and 2, are regulated by RNF146-mediated K48-linked polyubiquitylation and degradation. Here the authors show that this is opposed by K11-linked diubiquitylation by RING-UIM E3 ligases RNF114 and 166 and further impacted by several PAR-binding E3 ligases.
- Jerome Perrard
- & Susan Smith
-
Article
| Open Access N-terminal acetylation shields proteins from degradation and promotes age-dependent motility and longevity
N-terminal acetylation shields proteins from degradation and promotes age-dependent motility and longevity
The most common protein modification in eukaryotes is N-terminal acetylation, but its functional impact has remained enigmatic. Here, the authors find that a key role for N-terminal acetylation is shielding proteins from ubiquitin ligase-mediated degradation, mediating motility and longevity.
- Sylvia Varland
- , Rui Duarte Silva
- & Thomas Arnesen
-
Article
| Open Access lesSDRF is more: maximizing the value of proteomics data through streamlined metadata annotation
lesSDRF is more: maximizing the value of proteomics data through streamlined metadata annotation
Public proteomics data often lack essential metadata, limiting their potential. To address this, the authors developed lesSDRF, a tool to simplify the process of metadata annotation, thereby ensuring that data leave a lasting, impactful legacy well beyond their initial publication.
- Tine Claeys
- , Tim Van Den Bossche
- & Lennart Martens
-
Article
| Open Access Profiling of the Helicobacter pylori redox switch HP1021 regulon using a multi-omics approach
Profiling of the Helicobacter pylori redox switch HP1021 regulon using a multi-omics approach
Helicobacter pylori adapted to the harsh conditions of the human stomach using a handful of regulatory proteins. Here, the authors identify H. pylori processes controlled by the HP1021 response regulator under optimal growth and oxidative stress.
- Mateusz Noszka
- , Agnieszka Strzałka
- & Anna Zawilak-Pawlik
-
Article
| Open Access Automated imaging and identification of proteoforms directly from ovarian cancer tissue
Automated imaging and identification of proteoforms directly from ovarian cancer tissue
Identification of tissue proteoforms by top-down mass spectrometry remains challenging. Here, the authors present AutoPiMS, a semi-automated multiplexed tandem mass spectrometry workflow for proteoform identification directly from tissue contexts.
- John P. McGee
- , Pei Su
- & Neil L. Kelleher
-
Article
| Open Access PPTC7 maintains mitochondrial protein content by suppressing receptor-mediated mitophagy
PPTC7 maintains mitochondrial protein content by suppressing receptor-mediated mitophagy
The mitochondrial phosphatase PPTC7 has previously been linked to the maintenance of mitochondrial content, but the mechanisms underlying this phenotype remain unclear. Here, the authors demonstrate that loss of Pptc7 results in metabolic defects and further suggest that PPTC7 is a regulator of receptor-mediated mitophagy.
- Natalie M. Niemi
- , Lia R. Serrano
- & David J. Pagliarini
-
Article
| Open Access MYH10 activation rescues contractile defects in arrhythmogenic cardiomyopathy (ACM)
MYH10 activation rescues contractile defects in arrhythmogenic cardiomyopathy (ACM)
Arrhythmogenic cardiomyopathy is an untreatable heart muscle disease and a common cause of sudden cardiac death in young athletes. The authors show a link between actomyosin dysregulation and cardiac dysfunction by studying nonsense PKP2 mutants classified as pathogenic to identify a potential treatment.
- Nieves García-Quintáns
- , Silvia Sacristán
- & Juan A. Bernal
-
Article
| Open Access Proteogenetic drug response profiling elucidates targetable vulnerabilities of myelofibrosis
Proteogenetic drug response profiling elucidates targetable vulnerabilities of myelofibrosis
Myelofibrosis is a form of myeloproliferative neoplasm with few treatment options available. Here, the authors profiled drug responses and proteomics ex vivo and identify molecularly-guided treatment strategies, including HDAC and BET inhibitors for CALR mutant myelofibrosis patients.
- Mattheus H. E. Wildschut
- , Julien Mena
- & Berend Snijder
-
Article
| Open Access A cyclin-dependent kinase-mediated phosphorylation switch of disordered protein condensation
A cyclin-dependent kinase-mediated phosphorylation switch of disordered protein condensation
The authors show that dynamics of protein phosphorylation in the vertebrate cell cycle is largely attributable to CDK-mediated regulation of intrinsically disordered proteins that are involved in biomolecular condensate formation.
- Juan Manuel Valverde
- , Geronimo Dubra
- & Maarten Altelaar
-
Article
| Open Access Spectroscape enables real-time query and visualization of a spectral archive in proteomics
Spectroscape enables real-time query and visualization of a spectral archive in proteomics
Proteomics data repositories are deluged with data that is scarcely reused. Here, the authors developed Spectroscape, an interactive web-based tool for efficient similarity search of a query spectrum against a repository-scale spectral archive, and real-time visualization of its neighborhood.
- Long Wu
- , Ayman Hoque
- & Henry Lam
-
Article
| Open Access Chemoproteomic capture of RNA binding activity in living cells
Chemoproteomic capture of RNA binding activity in living cells
Here the authors introduce a photo-activatable-competition and chemoproteomic enrichment (PACCE) method to localize protein-RNA interfaces using photoactivatable cellular RNA to protect RNA binding regions on proteins from electrophilic purine probe labeling.
- Andrew J. Heindel
- , Jeffrey W. Brulet
- & Ku-Lung Hsu
-
Article
| Open Access FGF18 alleviates hepatic ischemia-reperfusion injury via the USP16-mediated KEAP1/Nrf2 signaling pathway in male mice
FGF18 alleviates hepatic ischemia-reperfusion injury via the USP16-mediated KEAP1/Nrf2 signaling pathway in male mice
Hepatic ischemia-reperfusion injury (IRI) is a common complication that occurs during hepatic resection and transplantation. Here the authors show that Hepatic stellate cells secrete FGF18 which alleviates hepatocyte injury by USP16/KEAP1/Nrf2 signaling.
- Gaozan Tong
- , Yiming Chen
- & XiaoKun Li
-
Article
| Open Access Multi-omics profiling reveals rhythmic liver function shaped by meal timing
Multi-omics profiling reveals rhythmic liver function shaped by meal timing
Post-translational modifications (PTMs) couple feed-fast cycles to circadian clocks. Here, the authors systematically profile daily rhythms of the proteome, 4 PTMs and lipidome in mouse livers under TRF, providing a comprehensive resource detailing rhythmic liver functions shaped by meal timing.
- Rongfeng Huang
- , Jianghui Chen
- & Min-Dian Li
-
Article
| Open Access Metabolic crosstalk between skeletal muscle cells and liver through IRF4-FSTL1 in nonalcoholic steatohepatitis
Metabolic crosstalk between skeletal muscle cells and liver through IRF4-FSTL1 in nonalcoholic steatohepatitis
In this study Guo et al. show that skeletal muscle IRF4-FSTL1 exerts metabolic regulation on the liver via DIP2A/CD14 in NASH. FSTL1 from skeletal muscle that enters the circulation promotes the development of NASH.
- Shanshan Guo
- , Yonghao Feng
- & Xingxing Kong
-
Article
| Open Access AlphaFold-Multimer predicts cross-kingdom interactions at the plant-pathogen interface
AlphaFold-Multimer predicts cross-kingdom interactions at the plant-pathogen interface
AlphaFold-Multimer was used to screen of 1,879 small secreted proteins from plant pathogens to be inhibitors of six tomato defense enzymes. Four of these inhibit subtilase P69B, showing the use of AI to predict cross-kingdom protein interactions.
- Felix Homma
- , Jie Huang
- & Renier A. L. van der Hoorn
-
Article
| Open Access Exploration of cell state heterogeneity using single-cell proteomics through sensitivity-tailored data-independent acquisition
Exploration of cell state heterogeneity using single-cell proteomics through sensitivity-tailored data-independent acquisition
Single-cell proteomics by Mass Spectrometry (scpMS) provides unparalleled insights into cellular mechanisms from a proteome-centric standpoint. Here, the authors leverage sensitivity-tailored data acquisition methods to profile cell state heterogeneity in cultured model systems.
- Valdemaras Petrosius
- , Pedro Aragon-Fernandez
- & Erwin M. Schoof
-
Article
| Open Access SEPepQuant enhances the detection of possible isoform regulations in shotgun proteomics
SEPepQuant enhances the detection of possible isoform regulations in shotgun proteomics
Protein isoform quantification in shotgun proteomics is challenging due to the mapping of many peptides to multiple protein isoforms. Here, the authors present a computational method SEPepQuant and demonstrate its utility in revealing protein isoform level regulation in shotgun proteomics.
- Yongchao Dou
- , Yuejia Liu
- & Bing Zhang
-
Article
| Open Access Discovery and pharmacophoric characterization of chemokine network inhibitors using phage-display, saturation mutagenesis and computational modelling
Discovery and pharmacophoric characterization of chemokine network inhibitors using phage-display, saturation mutagenesis and computational modelling
Ticks inject evasins at the bite site to bind multiple redundant chemokines and inhibit inflammation allowing blood feeding. Here, the authors identify evasin derived short peptides with broad spectrum anti-chemokine activity that could be used to develop new treatments for inflammatory disease.
- Serena Vales
- , Jhanna Kryukova
- & Shoumo Bhattacharya
-
Article
| Open Access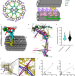 Integrated modeling of the Nexin-dynein regulatory complex reveals its regulatory mechanism
Integrated modeling of the Nexin-dynein regulatory complex reveals its regulatory mechanism
Motile cilia are hair-like structures that are found on the surface of eukaryotic cells providing cell motility. Here, authors reveal the twelve components of nexin-dynein regulatory complex and associated proteins in cilia from Tetrahymena thermophila.
- Avrin Ghanaeian
- , Sumita Majhi
- & Khanh Huy Bui
-
Article
| Open Access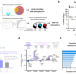 CSF proteome profiling reveals biomarkers to discriminate dementia with Lewy bodies from Alzheimer´s disease
CSF proteome profiling reveals biomarkers to discriminate dementia with Lewy bodies from Alzheimer´s disease
This study characterizes the CSF proteome changes underlying Dementia with Lewy Bodies (DLB) and identifies pathophysiological and diagnostic leads associated to this cause of dementia. Findings have been translated into a biomarker panel that could identify DLB patients with high accuracy across different cohorts.
- Marta del Campo
- , Lisa Vermunt
- & Charlotte E. Teunissen
-
Article
| Open Access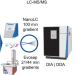 Deep and fast label-free Dynamic Organellar Mapping
Deep and fast label-free Dynamic Organellar Mapping
Regulated subcellular localization changes control protein function. Here, the authors provide a seamless spatial proteomics pipeline for mapping whole-cell protein localization dynamics, which includes a scalable workflow and a software suite for automated data analysis and visualization.
- Julia P. Schessner
- , Vincent Albrecht
- & Georg H. H. Borner
-
Article
| Open Access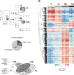 An integrated proteome and transcriptome of B cell maturation defines poised activation states of transitional and mature B cells
An integrated proteome and transcriptome of B cell maturation defines poised activation states of transitional and mature B cells
B cells pursue specific genetic programs to facilitate downstream cellular functions. Here the authors identify, using a combination of proteomic, transcriptomic and functional analyses, a group of mRNAs related to early activation and antibody production that are expressed in B cells without corresponding proteins, hinting a ‘poised’ state of B cells.
- Fiamma Salerno
- , Andrew J. M. Howden
- & Martin Turner
-
Article
| Open Access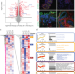 An integrated organoid omics map extends modeling potential of kidney disease
An integrated organoid omics map extends modeling potential of kidney disease
Lassé et al. show that genes involved in kidney organoid proteomic response to TNFα segregate a subset of individuals with poor outcomes in proteinuric kidney disease, demonstrating the relevance of kidney organoid modeling to human kidney disease.
- Moritz Lassé
- , Jamal El Saghir
- & Markus M. Rinschen
-
Article
| Open Access Proteomics and constraint-based modelling reveal enzyme kinetic properties of Chlamydomonas reinhardtii on a genome scale
Proteomics and constraint-based modelling reveal enzyme kinetic properties of Chlamydomonas reinhardtii on a genome scale
Closing a major gap in photosynthetic metabolic modelling, the authors provide over 500 estimates of in vivo enzyme catalytic rate in C. reinhardtii, which considerably improves predictions on how enzyme mass is allocated to different pathways.
- Marius Arend
- , David Zimmer
- & Zoran Nikoloski
-
Article
| Open Access Loss of N-terminal acetyltransferase A activity induces thermally unstable ribosomal proteins and increases their turnover in Saccharomyces cerevisiae
Loss of N-terminal acetyltransferase A activity induces thermally unstable ribosomal proteins and increases their turnover in Saccharomyces cerevisiae
N-terminal acetylation is a common modification with unclear function. Here, using multidimensional proteomics, the authors found that NatA-deficient yeast show increased ribosomal protein degradation and decreased ribosome thermostability, suggesting that N-terminal acetylation enhances proteome stability.
- Ulises H. Guzman
- , Henriette Aksnes
- & Jesper V. Olsen
-
Article
| Open Access MSBooster: improving peptide identification rates using deep learning-based features
MSBooster: improving peptide identification rates using deep learning-based features
There is a need for accessible ways to improve peptide spectrum match rescoring with deep learning predictions in bottom-up proteomics. Here, the authors demonstrate robust gains in peptide/protein identifications across various experiments, from single cell proteomics to immunopeptidomics.
- Kevin L. Yang
- , Fengchao Yu
- & Alexey I. Nesvizhskii
-
Article
| Open Access A ubiquitin-based effector-to-inhibitor switch coordinates early brain, craniofacial, and skin development
A ubiquitin-based effector-to-inhibitor switch coordinates early brain, craniofacial, and skin development
The molecular mechanisms ensuring early face, brain, and skin formation are unclear. Here, the authors uncover a posttranslational pathway that controls cytoskeletal signaling circuits to coordinate ectodermal patterning and neurulation.
- Anthony J. Asmar
- , Shaun R. Abrams
- & Achim Werner
-
Article
| Open Access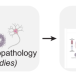 Compilation of reported protein changes in the brain in Alzheimer’s disease
Compilation of reported protein changes in the brain in Alzheimer’s disease
Proteomic studies in Alzheimer’s disease may be useful for understanding disease mechanisms and potential therapeutic targets. Here the authors describe a resource collating known protein changes throughout the progression of Alzheimer’s disease in human brain tissue.
- Manor Askenazi
- , Tomas Kavanagh
- & Eleanor Drummond
-
Article
| Open Access Mapping diversity in African trypanosomes using high resolution spatial proteomics
Mapping diversity in African trypanosomes using high resolution spatial proteomics
The molecular diversity between parasite species and life-stages correlate with diversity in the hosts they infect or the pathologies they cause. Here, Moloney et al., map the spatial proteomes of two African trypanosome species across two life stages and identify key routes of parasitic adaptation.
- Nicola M. Moloney
- , Konstantin Barylyuk
- & Paula MacGregor
-
Article
| Open Access Characterising the RNA-binding protein atlas of the mammalian brain uncovers RBM5 misregulation in mouse models of Huntington’s disease
Characterising the RNA-binding protein atlas of the mammalian brain uncovers RBM5 misregulation in mouse models of Huntington’s disease
RNA-Binding Proteins (RBPs) are critical regulators of RNA biology. Here, the authors describe the Brain-pCLAP methodology, uncover the RBP atlas of the mouse brain and demonstrate the differential binding of the splicing factor RBM5 to Huntington’s disease relevant transcripts in R6/2 mice.
- Meeli Mullari
- , Nicolas Fossat
- & Michael L. Nielsen
-
Article
| Open Access Proteogenomics of clear cell renal cell carcinoma response to tyrosine kinase inhibitor
Proteogenomics of clear cell renal cell carcinoma response to tyrosine kinase inhibitor
Many clear cell renal cell carcinoma (ccRCC) patients do not respond or develop resistance to tyrosine kinase inhibitors, such as Sunitinib. Here, the authors perform a proteogenomics analysis of Chinese ccRCC patients treated with Sunitinib and develop a multi-omics classifier to distinguish responders from non-responders.
- Hailiang Zhang
- , Lin Bai
- & Chen Ding
-
Article
| Open Access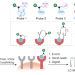 Deciphering intercellular signaling complexes by interaction-guided chemical proteomics
Deciphering intercellular signaling complexes by interaction-guided chemical proteomics
Systematic profiling of the indirect cell–cell interactions remains challenging. Here, the authors report a chemical proteomics method to identify ligand-receptor complexes formed between cell surface receptors and secreted proteins from neighboring cells.
- Jiangnan Zheng
- , Zhendong Zheng
- & Ruijun Tian
-
Article
| Open Access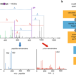 Detecting diagnostic features in MS/MS spectra of post-translationally modified peptides
Detecting diagnostic features in MS/MS spectra of post-translationally modified peptides
Protein modifications increase the complexity of data analysis in mass spectrometry-based proteomics, which may impair the comprehensive mapping of modification sites. Here, the authors develop an algorithm to extract diagnostic fragmentation patterns to improve modified peptide recovery and localization.
- Daniel J. Geiszler
- , Daniel A. Polasky
- & Alexey I. Nesvizhskii
-
Article
| Open Access Symbiont-host interactome mapping reveals effector-targeted modulation of hormone networks and activation of growth promotion
Symbiont-host interactome mapping reveals effector-targeted modulation of hormone networks and activation of growth promotion
Pathogens secrete effectors to promote disease, symbionts might use them to confer benefits. Here, the authors identify 106 candidate effectors from the symbiont Serendipita indica, characterise their interactions, and reveal their roles in regulating phytohormone signalling and promoting growth.
- Rory Osborne
- , Laura Rehneke
- & Patrick Schäfer
-
Article
| Open Access Glycopeptide database search and de novo sequencing with PEAKS GlycanFinder enable highly sensitive glycoproteomics
Glycopeptide database search and de novo sequencing with PEAKS GlycanFinder enable highly sensitive glycoproteomics
Accurate identification of intact glycopeptides from mass spectrometry data is essential for the characterization of glycosylation events in biological samples. Here, the authors propose GlycanFinder, a database search and de novo sequencing tool for the analysis of intact glycopeptides.
- Weiping Sun
- , Qianqiu Zhang
- & Baozhen Shan
-
Article
| Open Access Targeted protein degradation reveals BET bromodomains as the cellular target of Hedgehog pathway inhibitor-1
Targeted protein degradation reveals BET bromodomains as the cellular target of Hedgehog pathway inhibitor-1
Understanding the cellular target of hit compounds from phenotypic screens presents a major challenge yet is essential in the development of chemical probes. Here, the authors reveal the target of Hedgehog Pathway Inhibitor-1, by converting it to a bifunctional degrader, to be BET bromodomains.
- Meropi Bagka
- , Hyeonyi Choi
- & Sascha Hoogendoorn
-
Article
| Open Access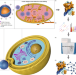 Targeted cross-linker delivery for the in situ mapping of protein conformations and interactions in mitochondria
Targeted cross-linker delivery for the in situ mapping of protein conformations and interactions in mitochondria
Current methods for analysing protein structures and interactions generally require the separation of specific organelles or changes to the intracellular environment. Here, authors developed nanocarrier-based cross-linking mass spectrometry techniques to assess mitochondrial proteins within living cells.
- Yuwan Chen
- , Wen Zhou
- & Yukui Zhang

 The protein interactome of the citrus Huanglongbing pathogen Candidatus Liberibacter asiaticus
The protein interactome of the citrus Huanglongbing pathogen Candidatus Liberibacter asiaticus