Featured
-
-
Article
| Open Access Proteogenomics refines the molecular classification of chronic lymphocytic leukemia
Proteogenomics refines the molecular classification of chronic lymphocytic leukemia
Proteomics can be used to refine cancer classification. Here, the authors characterise chronic lymphocytic leukaemia patients by proteogenomics, and identified a subtype of patients with poor prognosis associated with aberrant B cell receptor signalling.
- Sophie A. Herbst
- , Mattias Vesterlund
- & Sascha Dietrich
-
Article
| Open Access PIM1 promotes hepatic conversion by suppressing reprogramming-induced ferroptosis and cell cycle arrest
PIM1 promotes hepatic conversion by suppressing reprogramming-induced ferroptosis and cell cycle arrest
Protein kinase-mediated phosphorylation plays a critical role in many biological processes. Here the authors develop a trans-omics-based algorithm called Central Kinase Inference to integrate quantitative transcriptomic and phosphoproteomic data, finding that PIM1 promotes hepatic conversion by suppressing reprogramming-induced ferroptosis and cell cycle arrest.
- Yangyang Yuan
- , Chenwei Wang
- & Pengyu Huang
-
Article
| Open Access A method for Boolean analysis of protein interactions at a molecular level
A method for Boolean analysis of protein interactions at a molecular level
Determination of interactions between native proteins in cells is important for understanding function. Here the authors report MolBoolean as a method to detect interactions between endogenous proteins in subcellular compartments, using antibody-DNA conjugates for identification and signal amplification.
- Doroteya Raykova
- , Despoina Kermpatsou
- & Ola Söderberg
-
Article
| Open Access dia-PASEF data analysis using FragPipe and DIA-NN for deep proteomics of low sample amounts
dia-PASEF data analysis using FragPipe and DIA-NN for deep proteomics of low sample amounts
The dia-PASEF technology uses ion mobility separation to reduce signal interferences and increase sensitivity of mass spectrometry-based proteomics. The authors present algorithms and a software solution, which boost proteomic depth in dia-PASEF experiments by up to 83% compared to previous work, and are specifically beneficial for fast proteomic experiments and those with low sample amounts.
- Vadim Demichev
- , Lukasz Szyrwiel
- & Markus Ralser
-
Article
| Open Access HarmonizR enables data harmonization across independent proteomic datasets with appropriate handling of missing values
HarmonizR enables data harmonization across independent proteomic datasets with appropriate handling of missing values
Dataset integration is common practice to overcome limitations in statistically underpowered omics datasets. Here the authors present “HarmonizR”, a tool for missing data tolerant experimental variance reduction in large, integrated but independently generated datasets without data imputation, adjustable for individual dataset modalities, correction algorithm, and user preferences.
- Hannah Voß
- , Simon Schlumbohm
- & Christoph Krisp
-
Article
| Open Access A synaptomic analysis reveals dopamine hub synapses in the mouse striatum
A synaptomic analysis reveals dopamine hub synapses in the mouse striatum
The neurotransmitter dopamine is an important regulator of brain function. Here the authors describe “dopamine hub synapses”, where dopamine transmission may act in synergy with other neurotransmitters.
- Vincent Paget-Blanc
- , Marlene E. Pfeffer
- & Etienne Herzog
-
Article
| Open Access Cell type-specific biotin labeling in vivo resolves regional neuronal and astrocyte proteomic differences in mouse brain
Cell type-specific biotin labeling in vivo resolves regional neuronal and astrocyte proteomic differences in mouse brain
Current isolation-based approaches for cell type-specific proteomics pose several challenges. Here, the authors present an approach for in vivo cell type-specific protein labeling to characterize proteomic differences between neurons and astrocytes in their native state in adult mouse brain.
- Sruti Rayaprolu
- , Sara Bitarafan
- & Srikant Rangaraju
-
Article
| Open Access Mitochondrial calcium uniporter stabilization preserves energetic homeostasis during Complex I impairment
Mitochondrial calcium uniporter stabilization preserves energetic homeostasis during Complex I impairment
Mitochondrial complex I deficiency is frequent in congenital, neurologic and cardiovascular disease. Here the authors demonstrate that Complex I stimulates the turnover of a mitochondrial calcium channel, which becomes stabilized during Complex I deficiency, preserving energetic homeostasis.
- Enrique Balderas
- , David R. Eberhardt
- & Dipayan Chaudhuri
-
Article
| Open Access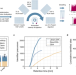 SPIN enables high throughput species identification of archaeological bone by proteomics
SPIN enables high throughput species identification of archaeological bone by proteomics
Available methods to identify species from fragmented archaeological bone and remains suffer a trade-off between cost and resolution. Here, the authors present a workflow that uses automated sample preparation, 10 to 20 times faster data acquisition, and computerized data interpretation to make the technology applicable to large-scale studies.
- Patrick Leopold Rüther
- , Immanuel Mirnes Husic
- & Jesper Velgaard Olsen
-
Article
| Open Access Hidden information on protein function in censuses of proteome foldedness
Hidden information on protein function in censuses of proteome foldedness
Proteomics can define features of proteome foldedness by assessing the reactivity of surface exposed amino acids. Here, the authors show that such exposure patterns yield insight to structural changes in chaperones as they bind to unfolded proteins in urea-denatured mammalian cell lysate.
- Dezerae Cox
- , Ching-Seng Ang
- & Danny M. Hatters
-
Article
| Open Access Tissue extracellular matrix hydrogels as alternatives to Matrigel for culturing gastrointestinal organoids
Tissue extracellular matrix hydrogels as alternatives to Matrigel for culturing gastrointestinal organoids
The culture of gastrointestinal organoids relies on Matrigel that has several drawbacks for clinical application. Here, the authors report the feasibility of gastrointestinal tissue-mimetic matrices as effective alternatives to Matrigel for organoid culture and transplantation.
- Suran Kim
- , Sungjin Min
- & Seung-Woo Cho
-
Article
| Open Access A microphysiological model of human trophoblast invasion during implantation
A microphysiological model of human trophoblast invasion during implantation
Normal and abnormal pregnancy is challenging to study and involves complex interactions between maternal and fetal cells. Here the authors present an implantation-on-a-chip device capable of modeling trophoblast invasion, a process critical to the establishment of pregnancy.
- Ju Young Park
- , Sneha Mani
- & Dan Dongeun Huh
-
Article
| Open Access A proximity biotinylation-based approach to identify protein-E3 ligase interactions induced by PROTACs and molecular glues
A proximity biotinylation-based approach to identify protein-E3 ligase interactions induced by PROTACs and molecular glues
PROTACs and molecular glues target E3 ubiquitin ligases to substrate proteins. Here, the authors develop a proximity biotinylation-based method to identify drug-induced E3 ligase-substrate interactions, enabling the assessment of the target spectrum of PROTACs and molecular glues in cells.
- Satoshi Yamanaka
- , Yuto Horiuchi
- & Tatsuya Sawasaki
-
Article
| Open Access An atlas of protein turnover rates in mouse tissues
An atlas of protein turnover rates in mouse tissues
Protein turnover underpins biology but is challenging to measure in vivo across the entire proteome. Here, the authors provide a comprehensive resource of protein turnover in mouse tissues and develop a visualization platform to analyze these data.
- Zach Rolfs
- , Brian L. Frey
- & Nathan V. Welham
-
Article
| Open Access Synergistic insights into human health from aptamer- and antibody-based proteomic profiling
Synergistic insights into human health from aptamer- and antibody-based proteomic profiling
Broad-capture affinity-based proteomic technologies inform how the readout of our genes affects human health. Here, the authors integrate aptamer- and antibody-based profiling to understand the mechanisms underlying gene-protein-disease associations.
- Maik Pietzner
- , Eleanor Wheeler
- & Claudia Langenberg
-
Article
| Open Access Smooth muscle-specific MMP17 (MT4-MMP) regulates the intestinal stem cell niche and regeneration after damage
Smooth muscle-specific MMP17 (MT4-MMP) regulates the intestinal stem cell niche and regeneration after damage
While the role of smooth muscle in peristalsis has been studied extensively, little is known about its other functions in the intestine. Here the authors identify MMP17, expressed by smooth muscle cells, as a modulator of intestinal epithelial regeneration and the intestinal stem cell niche.
- Mara Martín-Alonso
- , Sharif Iqbal
- & Menno J. Oudhoff
-
Article
| Open Access Photoactivatable ribonucleosides mark base-specific RNA-binding sites
Photoactivatable ribonucleosides mark base-specific RNA-binding sites
RNA-protein interactions play critical roles in post-transcriptional gene regulation. Here the authors demonstrate pRBS-ID, an updated MS/MS-based method that combines the benefits of photoactivatable ribonucleosides and the chemical cleavage of RNA.
- Jong Woo Bae
- , Sangtae Kim
- & Jong-Seo Kim
-
Article
| Open Access The regulatory landscape of the human HPF1- and ARH3-dependent ADP-ribosylome
The regulatory landscape of the human HPF1- and ARH3-dependent ADP-ribosylome
ADP-ribosylation is regulated by HPF1 and ARH3, but the cellular target spectrum of these enzymes is not fully understood. Here, the authors use quantitative proteomics to define the HPF1- and ARH3-dependent ADP-ribosylome, providing evidence that mono-ADP-ribosylation of serine predominates in cells.
- Ivo A. Hendriks
- , Sara C. Buch-Larsen
- & Michael L. Nielsen
-
Article
| Open Access Protein identification by nanopore peptide profiling
Protein identification by nanopore peptide profiling
Peptide mass fingerprinting is a traditional approach for protein identification by mass spectrometry. Here, the authors provide evidence that peptide mass fingerprinting is also feasible using FraC nanopores, demonstrating protein identification based on nanopore measurements of digested peptides.
- Florian Leonardus Rudolfus Lucas
- , Roderick Corstiaan Abraham Versloot
- & Giovanni Maglia
-
Article
| Open Access Spatiotemporal proteomic profiling of the pro-inflammatory response to lipopolysaccharide in the THP-1 human leukaemia cell line
Spatiotemporal proteomic profiling of the pro-inflammatory response to lipopolysaccharide in the THP-1 human leukaemia cell line
“Protein relocalisation plays a major role in the innate immune response but remains incompletely characterised. Here, the authors combine temporal proteomics with LOPIT, a spatial proteomic workflow, in a fully Bayesian framework to elucidate spatiotemporal proteomic changes during the LPS-induced immune response in THP-1 cells.
- Claire M. Mulvey
- , Lisa M. Breckels
- & Kathryn S. Lilley
-
Article
| Open Access Off-the-shelf proximity biotinylation for interaction proteomics
Off-the-shelf proximity biotinylation for interaction proteomics
Proximity biotinylation is a powerful tool to profile interactomes, but it requires genetic engineering of the target protein. Here, the authors develop a proximity biotinylation enzyme that can be directed to the target using antibodies, enabling interactome profiling of endogenous proteins or PTMs.
- Irene Santos-Barriopedro
- , Guido van Mierlo
- & Michiel Vermeulen
-
Article
| Open Access Cell-type and subcellular compartment-specific APEX2 proximity labeling reveals activity-dependent nuclear proteome dynamics in the striatum
Cell-type and subcellular compartment-specific APEX2 proximity labeling reveals activity-dependent nuclear proteome dynamics in the striatum
Mapping neuronal proteomes with genetic, subcellular, and temporal specificity is a challenging task. This study uncovers proteome dynamics in two classes of striatal spiny projection neurons in the mouse brain using a genetically targeted APEX2-based proximity labeling approach.
- V. Dumrongprechachan
- , R. B. Salisbury
- & Y. Kozorovitskiy
-
Article
| Open Access Insulin signaling regulates longevity through protein phosphorylation in Caenorhabditis elegans
Insulin signaling regulates longevity through protein phosphorylation in Caenorhabditis elegans
How phosphorylation mediated by Insulin/IGF-1 Signaling kinases regulates lifespan remains unclear. Here the authors perform a large-scale quantitative phosphoproteomic analysis of wildtype and IIS mutant C. elegans strains to reveal detailed functional insights into longevity.
- Wen-Jun Li
- , Chen-Wei Wang
- & Meng-Qiu Dong
-
Article
| Open Access Composition and stage dynamics of mitochondrial complexes in Plasmodium falciparum
Composition and stage dynamics of mitochondrial complexes in Plasmodium falciparum
Applying complexome profiling, Evers et al. unravel the composition of mitochondrial oxidative phosphorylation complexes in P. falciparum asexual and sexual blood stages. Abundance of these complexes differs between both stages, supporting the hypothesis that a mitochondrial metabolic switch is central to gametocyte development and functioning.
- Felix Evers
- , Alfredo Cabrera-Orefice
- & Taco W. A. Kooij
-
Article
| Open Access Multianalyte serology in home-sampled blood enables an unbiased assessment of the immune response against SARS-CoV-2
Multianalyte serology in home-sampled blood enables an unbiased assessment of the immune response against SARS-CoV-2
Here, Roxhed et al. develop a multiplexed approach to screen IgG and IgM levels against several SARS-CoV-2 proteins in home-sampled dried blood spots and estimate seroprevalence of 12.5% in Stockholm in spring of 2020.
- Niclas Roxhed
- , Annika Bendes
- & Jochen M. Schwenk
-
Article
| Open Access Next generation plasma proteome profiling to monitor health and disease
Next generation plasma proteome profiling to monitor health and disease
The proximity extension assay (PEA) is a popular tool to measure plasma protein levels. Here, the authors extend the proteome coverage of PEA by combining it with next-generation sequencing, enabling the analysis of nearly 1500 proteins from minute amounts of plasma.
- Wen Zhong
- , Fredrik Edfors
- & Mathias Uhlén
-
Article
| Open Access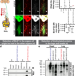 Proteomics of protein trafficking by in vivo tissue-specific labeling
Proteomics of protein trafficking by in vivo tissue-specific labeling
The network of proteins secreted for interorgan communication is poorly understood. Here, the authors develop a method, based on protein labeling, to study cell-specific secretomes and interorgan protein trafficking, and demonstrate their approach in Drosophila and mouse models.
- Ilia A. Droujinine
- , Amanda S. Meyer
- & Norbert Perrimon
-
Article
| Open Access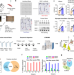 Deep muscle-proteomic analysis of freeze-dried human muscle biopsies reveals fiber type-specific adaptations to exercise training
Deep muscle-proteomic analysis of freeze-dried human muscle biopsies reveals fiber type-specific adaptations to exercise training
Skeletal muscle conveys the beneficial effects of physical exercise but due to its heterogeneity, studying the effects of exercise on muscle fibres is challenging. Here, the authors carry out proteomic analysis of myofibres from freeze-dried muscle biopsies, show fibre-type specific changes in response to exercise, and show that the oxidative and glycolytic muscle fibers adapt differentially to exercise training.
- A. S. Deshmukh
- , D. E. Steenberg
- & J. F. P. Wojtaszewski
-
Article
| Open Access Data-independent acquisition method for ubiquitinome analysis reveals regulation of circadian biology
Data-independent acquisition method for ubiquitinome analysis reveals regulation of circadian biology
Protein ubiquitylation is often studied by proteomics but how data independent acquisition (DIA) may advance these studies remains to be explored. Here, the authors show that DIA improves ubiquitylation site identification and quantification, enabling them to characterize the circadian ubiquitinome in human cells.
- Fynn M. Hansen
- , Maria C. Tanzer
- & Matthias Mann
-
Article
| Open Access Establishing a mass spectrometry-based system for rapid detection of SARS-CoV-2 in large clinical sample cohorts
Establishing a mass spectrometry-based system for rapid detection of SARS-CoV-2 in large clinical sample cohorts
Large population testing is a key step to controlling the COVID-19 pandemic. Here, the authors develop a targeted mass spectrometry system for peptide-based SARS-CoV-2 detection, allowing analysis of over 500 swab samples per day and enabling virus detection even after prolonged sample storage at room temperature.
- Karina Helena Morais Cardozo
- , Adriana Lebkuchen
- & Valdemir Melechco Carvalho
-
Article
| Open Access Immune suppression in the early stage of COVID-19 disease
Immune suppression in the early stage of COVID-19 disease
How COVID-19 pathology differs from other drivers of pneumonia is unclear. Here the authors analyze urine from patients with COVID-19 and identify an immunosuppressive protein expression pattern that is distinct from the pattern in healthy individuals or patients with non-COVID-19 pneumonia.
- Wenmin Tian
- , Nan Zhang
- & Catherine C. L. Wong
-
Perspective
| Open Access A high-stringency blueprint of the human proteome
A high-stringency blueprint of the human proteome
The Human Proteome Project (HPP) was launched in 2010 to enhance accurate annotation of the genome-encoded proteome. Ten years later, the HPP releases its first blueprint of the human proteome, annotating 90% of all known proteins at high-stringency and discussing the implications of proteomics for precision medicine.
- Subash Adhikari
- , Edouard C. Nice
- & Mark S. Baker
-
Article
| Open Access Identification of a herpes simplex virus 1 gene encoding neurovirulence factor by chemical proteomics
Identification of a herpes simplex virus 1 gene encoding neurovirulence factor by chemical proteomics
Here the authors use chemical proteomics to identify the herpes simplex virus 1 encoded proteome in infected cells. Functional characterization of one of the nine identified proteins, designated piUL49, shows that it acts as neurovirulence factor in mice by regulating a virally encoded dUTPase.
- Akihisa Kato
- , Shungo Adachi
- & Yasushi Kawaguchi
-
Article
| Open Access Widespread protein lysine acetylation in gut microbiome and its alterations in patients with Crohn’s disease
Widespread protein lysine acetylation in gut microbiome and its alterations in patients with Crohn’s disease
Intestinal microbiota is increasingly reported to influence human health, but little is known on how its functions are regulated. Here the authors characterize microbiome protein acetylation and demonstrate its potential roles in shaping gut microbial functions and the onset of Crohn’s disease.
- Xu Zhang
- , Zhibin Ning
- & Daniel Figeys
-
Article
| Open Access Multiscale causal networks identify VGF as a key regulator of Alzheimer’s disease
Multiscale causal networks identify VGF as a key regulator of Alzheimer’s disease
To investigate the molecular foundation of sporadic Alzheimer’s disease (AD), Beckmann et al. constructed multiscale causal networks on a large human AD multi-omics dataset, detecting AD-associated networks and their top predicted regulator, VGF, with extensive validation in the 5xFAD mouse model.
- Noam D. Beckmann
- , Wei-Jye Lin
- & Eric E. Schadt
-
Article
| Open Access Integrated pharmaco-proteogenomics defines two subgroups in isocitrate dehydrogenase wild-type glioblastoma with prognostic and therapeutic opportunities
Integrated pharmaco-proteogenomics defines two subgroups in isocitrate dehydrogenase wild-type glioblastoma with prognostic and therapeutic opportunities
The heterogeneity of IDH1/2 wild-type glioblastoma limits its prognosis and therapy. Here, the authors show a binary stratification, based on quantitative proteomic analysis of samples from patients with glioblastoma, with different prognosis and therapeutic vulnerabilities.
- Sejin Oh
- , Jeonghun Yeom
- & Hyun Seok Kim
-
Article
| Open Access Blood circulation of soft nanomaterials is governed by dynamic remodeling of protein opsonins at nano-biointerface
Blood circulation of soft nanomaterials is governed by dynamic remodeling of protein opsonins at nano-biointerface
The blood circulation time is important to the biomedical application of nanomaterials. Here, the authors explore the effect of protein corona formation on the blood residency of nanomaterials and show circulation times are governed by the dynamic remodelling of protein opsonins in vivo.
- Srinivas Abbina
- , Lily E. Takeuchi
- & Jayachandran N. Kizhakkedathu
-
Article
| Open Access Plasma-derived extracellular vesicles from Plasmodium vivax patients signal spleen fibroblasts via NF-kB facilitating parasite cytoadherence
Plasma-derived extracellular vesicles from Plasmodium vivax patients signal spleen fibroblasts via NF-kB facilitating parasite cytoadherence
Extracellular vesicles (EVs) in plasma can affect pathogenesis of parasites, but details remain unclear. Here, Toda et al. characterize plasma-derived EVs from Plasmodium vivax patients and show that PvEVs are preferentially taken up by human spleen fibroblasts, facilitating parasite cytoadherence.
- Haruka Toda
- , Miriam Diaz-Varela
- & Hernando A. del Portillo
-
Article
| Open Access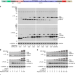 Split Intein-Mediated Protein Ligation for detecting protein-protein interactions and their inhibition
Split Intein-Mediated Protein Ligation for detecting protein-protein interactions and their inhibition
Protein-protein interactions are fundamental to the regulation of protein activity and cellular phyisology. Here the authors present Split Intein-Mediated Protein Ligation, which uses bait and prey proteins fused to intein fragments to generate single intact proteins upon interaction.
- Zhong Yao
- , Farzaneh Aboualizadeh
- & Igor Stagljar
-
Article
| Open Access A shift in glutamine nitrogen metabolism contributes to the malignant progression of cancer
A shift in glutamine nitrogen metabolism contributes to the malignant progression of cancer
Glucose metabolism is known to be dysregulated in cancer. Here, the authors show that glutamine nitrogen is also affected in cancer and demonstrate that glutaminase 1 and phosphoribosyl pyrophosphate amidotransferase are the key enzymes that control this metabolic switch.
- Manabu Kodama
- , Kiyotaka Oshikawa
- & Keiichi I. Nakayama
-
Article
| Open Access Temporal dynamics of protein complex formation and dissociation during human cytomegalovirus infection
Temporal dynamics of protein complex formation and dissociation during human cytomegalovirus infection
Here, Hashimoto et al. apply mass spectrometry-based thermal proximity coaggregation to characterize the temporal dynamics of virus-host protein-protein interactions during human cytomegalovirus (HCMV) infection, uncovering proviral functions including the internalization of the HCMV receptor integrin beta 1 with CD63.
- Yutaka Hashimoto
- , Xinlei Sheng
- & Ileana M. Cristea
-
Article
| Open Access Robust, reproducible and quantitative analysis of thousands of proteomes by micro-flow LC–MS/MS
Robust, reproducible and quantitative analysis of thousands of proteomes by micro-flow LC–MS/MS
Mass spectrometry-based proteomics typically relies on highly sensitive nano-flow liquid chromatography (LC) but this can reduce robustness and reproducibility. Here, the authors show that micro-flow LC enables robust and reproducible high-throughput proteomics experiments at a very moderate loss of sensitivity.
- Yangyang Bian
- , Runsheng Zheng
- & Bernhard Kuster
-
Article
| Open Access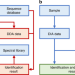 In silico spectral libraries by deep learning facilitate data-independent acquisition proteomics
In silico spectral libraries by deep learning facilitate data-independent acquisition proteomics
Data-independent acquisition (DIA) is an emerging technology in proteomics but it typically relies on spectral libraries built by data-dependent acquisition (DDA). Here, the authors use deep learning to generate in silico spectral libraries directly from protein sequences that enable more comprehensive DIA experiments than DDA-based libraries.
- Yi Yang
- , Xiaohui Liu
- & Liang Qiao
-
Article
| Open Access Classification of mouse B cell types using surfaceome proteotype maps
Classification of mouse B cell types using surfaceome proteotype maps
Analysis of the cell surface proteome (surfaceome) is essential for cell classification but is technically challenging. Here the authors miniaturize and automate the Cell Surface Capture method to increase sensitivity, reproducibility and throughput, and use it to create population-specific surfaceome maps of developing mouse B cells.
- Marc van Oostrum
- , Maik Müller
- & Bernd Wollscheid
-
Comment
| Open AccessThe ever expanding scope of electrospray mass spectrometry—a 30 year journey
John Fenn’s electrospray mass spectrometry (ESMS) was awarded the chemistry Nobel Prize in 2002 and is now the basis of the entire field of MS-based proteomics. Technological progress continues unabated, enabling single cell sensitivity and clinical applications.
- Matthias Mann
-
Article
| Open Access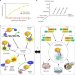 Maximizing binary interactome mapping with a minimal number of assays
Maximizing binary interactome mapping with a minimal number of assays
Comprehensive mapping of binary protein-protein interactions requires to combine several complementary assays. Here, the authors show that complete coverage could be reached with a minimal number of assays as long as they explore various experimental conditions.
- Soon Gang Choi
- , Julien Olivet
- & Yves Jacob
-
Article
| Open Access Profiling surface proteins on individual exosomes using a proximity barcoding assay
Profiling surface proteins on individual exosomes using a proximity barcoding assay
The use of antibodies to capture and profile exosomes limits the number of target proteins that can be detected. Here the authors develop a proximity-dependent barcoding assay that allows profiling of 38 surface proteins on individual exosomes from heterogeneous samples such as serum and seminal fluid.
- Di Wu
- , Junhong Yan
- & Masood Kamali-Moghaddam
-
Article
| Open Access Deep multiomics profiling of brain tumors identifies signaling networks downstream of cancer driver genes
Deep multiomics profiling of brain tumors identifies signaling networks downstream of cancer driver genes
Multi-omic profiling is a powerful approach to dissecting molecular mechanisms in disease. Here the authors generate whole proteome, phosphoproteome and transcriptome profiles from two mouse models of high-grade glioma driven by different oncogenes, and validate identified master regulators with a CRISPR screen.
- Hong Wang
- , Alexander K. Diaz
- & Junmin Peng
-
Article
| Open Access Proteogenomic landscape of squamous cell lung cancer
Proteogenomic landscape of squamous cell lung cancer
Squamous cell lung cancer has dismal prognosis due to the dearth of effective treatments. Here, the authors perform an integrated proteogenomic analysis of the disease, revealing three proteomics-based subtypes and suggesting potential therapeutic opportunities.
- Paul A. Stewart
- , Eric A. Welsh
- & Eric B. Haura

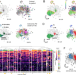 Single-cell profiling reveals a memory B cell-like subtype of follicular lymphoma with increased transformation risk
Single-cell profiling reveals a memory B cell-like subtype of follicular lymphoma with increased transformation risk