Featured
-
-
Article |
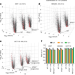 Systematic quantitative analysis of ribosome inventory during nutrient stress
Systematic quantitative analysis of ribosome inventory during nutrient stress
During nutrient stress, ribosomal protein abundance is regulated primarily by translational and non-autophagic degradative mechanisms, but ribosome density per cell is largely maintained by reductions in cell volume and rates of cell division.
- Heeseon An
- , Alban Ordureau
- & J. Wade Harper
-
Article |
 Lipid signalling drives proteolytic rewiring of mitochondria by YME1L
Lipid signalling drives proteolytic rewiring of mitochondria by YME1L
Under conditions such as hypoxia or starvation, an mTORC1-lipid signalling pathway initiates mitochondrial proteolysis by YME1L.
- Thomas MacVicar
- , Yohsuke Ohba
- & Thomas Langer
-
Article |
 Mitochondrial protein translocation-associated degradation
Mitochondrial protein translocation-associated degradation
In Saccharomyces cerevisiae, Ubx2 promotes clearing trapped precursor proteins from the channel of the translocase of the outer membrane, in a translocation-associated degradation pathway that maintains the protein import capacity of mitochondria.
- Christoph U. Mårtensson
- , Chantal Priesnitz
- & Thomas Becker
-
Letter |
 Distinct proteostasis circuits cooperate in nuclear and cytoplasmic protein quality control
Distinct proteostasis circuits cooperate in nuclear and cytoplasmic protein quality control
Ubiquitin chains linked to cytoplasmic misfolded proteins are different from those linked to nuclear misfolded proteins, each requiring a distinct combination of molecular chaperones and ubiquitination circuitries.
- Rahul S. Samant
- , Christine M. Livingston
- & Judith Frydman
-
Letter |
 Cotranslational assembly of protein complexes in eukaryotes revealed by ribosome profiling
Cotranslational assembly of protein complexes in eukaryotes revealed by ribosome profiling
Cotranslational assembly is a prevalent mechanism for the formation of oligomeric complexes in Saccharomyces cerevisiae, with one subunit serving as scaffold for the translation of partner subunits.
- Ayala Shiber
- , Kristina Döring
- & Bernd Bukau
-
Letter |
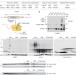 Vms1 and ANKZF1 peptidyl-tRNA hydrolases release nascent chains from stalled ribosomes
Vms1 and ANKZF1 peptidyl-tRNA hydrolases release nascent chains from stalled ribosomes
The Cdc48 adaptor Vms1 is a peptidyl-tRNA hydrolase that cooperates with the ribosome quality control complex to catalyse the removal of nascent polypeptides from stalled ribosomes.
- Rati Verma
- , Kurt M. Reichermeier
- & Raymond J. Deshaies
-
Article |
 An evolutionarily conserved pathway controls proteasome homeostasis
An evolutionarily conserved pathway controls proteasome homeostasis
Proteasome abundance is crucial for cell survival, but how cells maintain adequate amounts of proteasome is unclear; an analysis in yeast identifies TORC1 and Mpk1 as central components of a pathway regulating proteasome homeostasis through the coordinated regulation of regulatory particle assembly chaperones and proteasome subunits—this pathway is evolutionarily conserved with mTOR and ERK5 regulating proteasome abundance in mammals.
- Adrien Rousseau
- & Anne Bertolotti
-
Letter |
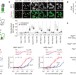 Protein quality control at the inner nuclear membrane
Protein quality control at the inner nuclear membrane
A protein degradation pathway is found at the inner nuclear membrane that is distinct from, but complementary to, endoplasmic-reticulum-associated protein degradation, and which is mediated by the Asi protein complex; a genome-wide library screening of yeast identifies more than 20 substrates of this pathway, which is shown to target mislocalized integral membrane proteins for degradation.
- Anton Khmelinskii
- , Ewa Blaszczak
- & Michael Knop
-
Letter |
 Role of a ribosome-associated E3 ubiquitin ligase in protein quality control
Role of a ribosome-associated E3 ubiquitin ligase in protein quality control
The translation of messenger RNA that lacks stop codons results in the production of aberrant proteins, which may have harmful effects on the cell. It is unclear how eukaryotic cells eliminate these 'non-stop' proteins. Here it is shown that, in Saccharomyces cerevisiae, an E3 ubiquitin ligase called Ltn1 acts in the quality-control pathway. It associates with ribosomes and marks non-stop proteins with ubiquitin, which targets the proteins for degradation.
- Mario H. Bengtson
- & Claudio A. P. Joazeiro

 Structural basis for dimerization quality control
Structural basis for dimerization quality control