Article
|
Open Access
Featured
-
-
Article
| Open Access Prophages and satellite prophages are widespread in Streptococcus and may play a role in pneumococcal pathogenesis
Prophages and satellite prophages are widespread in Streptococcus and may play a role in pneumococcal pathogenesis
Prophages are viral genomes integrated within bacterial genomes. Here, Rezaei Javan et al. identify nearly 800 prophages and satellite prophages in > 1300 Streptococcus genomes, and show that a satellite prophage is associated with virulence in a mouse model of pneumococcal infection.
- Reza Rezaei Javan
- , Elisa Ramos-Sevillano
- & Angela B. Brueggemann
-
Article
| Open Access The structure of a polygamous repressor reveals how phage-inducible chromosomal islands spread in nature
The structure of a polygamous repressor reveals how phage-inducible chromosomal islands spread in nature
Staphylococcus aureus pathogenicity islands (SaPIs) encode the master repressor Stl and after bacteriophage infection Stl interacts with specific phage proteins leading to a derepression of SaPIs. Here the authors provide structural insights into this family of repressors by determining the crystal structures of SaPIbov1 Stl alone and in complex with two structurally unrelated phage dUTPases.
- J. Rafael Ciges-Tomas
- , Christian Alite
- & Alberto Marina
-
Article
| Open Access Structural basis for the adsorption of a single-stranded RNA bacteriophage
Structural basis for the adsorption of a single-stranded RNA bacteriophage
Single-stranded RNA bacteriophages use a single maturation protein (Mat) to attach to a retractile pilus of the bacterial host. Here, the authors report the structures of the MS2 phage bound to the host receptor F-pili and define the orientations of Mat relative to the cell and emanating F-pili, providing new insights into the F-like type IV secretion systems.
- Ran Meng
- , Mengqiu Jiang
- & Junjie Zhang
-
Article
| Open Access Identification and characterization of a direct activator of a gene transfer agent
Identification and characterization of a direct activator of a gene transfer agent
Gene transfer agents (GTAs) are ‘domesticated’ bacteriophages that can transfer any genes between bacteria. Here, Paul Fogg identifies a protein that directly regulates transcription of GTA genes and whose expression is in turn controlled by a global cell-cycle regulator and a quorum-sensing regulator.
- Paul C. M. Fogg
-
Article
| Open Access CRISPR analysis suggests that small circular single-stranded DNA smacoviruses infect Archaea instead of humans
CRISPR analysis suggests that small circular single-stranded DNA smacoviruses infect Archaea instead of humans
Smacoviruses are found in the intestinal tract of humans and animals but their precise host remains elusive. Here, the authors identify smacovirus-matching CRISPR spacer sequences in the faecal archaeon Candidatus Methanomassiliicoccus intestinalis, implicating Archaea as a potential host.
- César Díez-Villaseñor
- & Francisco Rodriguez-Valera
-
Article
| Open Access Widespread anti-CRISPR proteins in virulent bacteriophages inhibit a range of Cas9 proteins
Widespread anti-CRISPR proteins in virulent bacteriophages inhibit a range of Cas9 proteins
Some phages carry genes coding for anti-CRISPR (Acr) proteins that interfere with the activity of bacterial CRISPR-Cas systems. Here, Hynes et al. characterize a new Acr family from streptococcal phages and investigate its potential in genome-editing applications.
- Alexander P. Hynes
- , Geneviève M. Rousseau
- & Sylvain Moineau
-
Article
| Open Access Anti-phage islands force their target phage to directly mediate island excision and spread
Anti-phage islands force their target phage to directly mediate island excision and spread
Mobile genetic elements called PLEs protect Vibrio cholerae from infection with phage ICP1 by unclear mechanisms. Here, McKitterick and Seed show that a PLE-encoded large serine recombinase exploits an ICP1 protein as a recombination directionality factor to excise this PLE in response to phage infection.
- Amelia C. McKitterick
- & Kimberley D. Seed
-
Article
| Open Access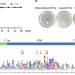 Regulatory protein SrpA controls phage infection and core cellular processes in Pseudomonas aeruginosa
Regulatory protein SrpA controls phage infection and core cellular processes in Pseudomonas aeruginosa
You et al. show that SrpA, a small protein widely conserved among bacteria, controls core cellular processes in response to phage infection and environmental signals in Pseudomonas aeruginosa, including cell motility, chemotaxis, biofilm formation, and virulence.
- Jiajia You
- , Li Sun
- & Hongjiang Yang
-
Article
| Open Access Bacteriophage T5 tail tube structure suggests a trigger mechanism for Siphoviridae DNA ejection
Bacteriophage T5 tail tube structure suggests a trigger mechanism for Siphoviridae DNA ejection
Host cell recognition is mediated by the phage tail tip proteins, which then triggers viral genome delivery via the phage tail. Here, the authors combine crystallography and cryoEM to structurally characterise the bacteriophage T5 tail tube structure before and after interaction with its host receptor.
- Charles-Adrien Arnaud
- , Grégory Effantin
- & Cécile Breyton
-
Article
| Open Access Internalization of a polysialic acid-binding Escherichia coli bacteriophage into eukaryotic neuroblastoma cells
Internalization of a polysialic acid-binding Escherichia coli bacteriophage into eukaryotic neuroblastoma cells
Eukaryotic organisms are continuously exposed to bacteriophages, but these are not thought to enter non-phagocytic cells. Here, Lehti et al. show that a bacteriophage can bind to a specific receptor on the surface of human neuroblastoma cells in vitro, and be internalized via the endolysosomal route.
- Timo A. Lehti
- , Maria I. Pajunen
- & Jukka Finne
-
Article
| Open Access Ecogenomics of virophages and their giant virus hosts assessed through time series metagenomics
Ecogenomics of virophages and their giant virus hosts assessed through time series metagenomics
Virophages are recently-identified small viruses that infect larger viruses, yet their diversity and ecological roles are poorly understood. Here, Roux and colleagues present time series metagenomics data revealing new virophage genera and their putative ecological interactions in two freshwater lakes.
- Simon Roux
- , Leong-Keat Chan
- & Matthew B. Sullivan
-
Article
| Open Access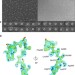 Portal protein functions akin to a DNA-sensor that couples genome-packaging to icosahedral capsid maturation
Portal protein functions akin to a DNA-sensor that couples genome-packaging to icosahedral capsid maturation
Tailed bacteriophages assemble empty precursor capsids known as procapsids that are subsequently filled with viral DNA by a genome-packaging motor. Here the authors present a structure-based analysis that suggests the signal for termination of genome packaging is achieved through a DNA-dependent symmetrization of portal protein.
- Ravi K. Lokareddy
- , Rajeshwer S. Sankhala
- & Gino Cingolani
-
Article
| Open Access Asymmetric cryo-EM reconstruction of phage MS2 reveals genome structure in situ
Asymmetric cryo-EM reconstruction of phage MS2 reveals genome structure in situ
MS2 is a single-stranded RNA bacteriophage that infects its host via adsorption to bacterial pili. Here the authors visualize the MS2 virion with asymmetric cryo-EM reconstruction, revealing that the genome of MS2 adopts a specific structure of asymmetrically distributed stem-loops connected to the capsid.
- Roman I Koning
- , Josue Gomez-Blanco
- & Abraham J. Koster
-
Article |
 Adaptation in bacterial CRISPR-Cas immunity can be driven by defective phages
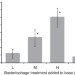
Adaptation in bacterial CRISPR-Cas immunity can be driven by defective phages
The bacterial ‘adaptive’ immune system known as CRISPR-Cas destroys foreign DNA molecules, such as viral genomes, to which the cells have previously been exposed. Here, Hynes et al.show that this gain of immunity is favoured by exposure to defective viruses, a result reminiscent of vaccination.
- Alexander P. Hynes
- , Manuela Villion
- & Sylvain Moineau

 Coordination of cohabiting phage elements supports bacteria–phage cooperation
Coordination of cohabiting phage elements supports bacteria–phage cooperation