Featured
-
-
Article
| Open Access Late-stage guanine C8–H alkylation of nucleosides, nucleotides, and oligonucleotides via photo-mediated Minisci reaction
Late-stage guanine C8–H alkylation of nucleosides, nucleotides, and oligonucleotides via photo-mediated Minisci reaction
Chemically modified nucleobases and oligonucleotides are essential in several fields but introducing functional groups into nucleobases requires laborious chemical synthesis. Here, the authors report site-selective alkylation at the C8-position of guanines in guanosine, GMP, GDP, and GTP, as well as late-stage alkylation of RNA/DNA oligonucleotides through photomediated Minisci reaction.
- Ruoqian Xie
- , Wanlu Li
- & Gang Chen
-
Article
| Open Access Trabectedin derails transcription-coupled nucleotide excision repair to induce DNA breaks in highly transcribed genes
Trabectedin derails transcription-coupled nucleotide excision repair to induce DNA breaks in highly transcribed genes
The antitumor drug trabectedin is more toxic to DNA-repair-proficient cells. Here the authors show that this is caused by persistent DNA breaks induced from an abortive repair reaction and develop “TRABI-Seq” to map the breaks to transcribed regions of the genome. Trabectedin may thus be used as a diagnostic and therapeutic in precision oncology.
- Kook Son
- , Vakil Takhaveev
- & Orlando D. Schärer
-
Article
| Open Access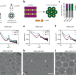 Liquid crystalline inverted lipid phases encapsulating siRNA enhance lipid nanoparticle mediated transfection
Liquid crystalline inverted lipid phases encapsulating siRNA enhance lipid nanoparticle mediated transfection
The authors display the bottom-up design, assembly, and in-depth characterization of defined lipid-RNA structures in the core of lipid nanoparticles. The inverted structures are thermostable and provide better transfection over lamellar structures.
- Roy Pattipeiluhu
- , Ye Zeng
- & Thomas H. Sharp
-
Article
| Open Access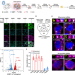 Profiling stress-triggered RNA condensation with photocatalytic proximity labeling
Profiling stress-triggered RNA condensation with photocatalytic proximity labeling
Stress granules (SGs) are highly dynamic cytoplasmic membraneless organelles that assemble when cells are challenged by stress. Herein, the authors apply a proximity-dependent RNA labeling method, CAP-seq, to comprehensively investigate the content of SG-proximal transcriptome and the dynamic change in SG-proximal transcriptome along the time course of granule assembly and disassembly processes in live mammalian cells.
- Ziqi Ren
- , Wei Tang
- & Peng Zou
-
Article
| Open Access Enzymatic synthesis and nanopore sequencing of 12-letter supernumerary DNA
Enzymatic synthesis and nanopore sequencing of 12-letter supernumerary DNA
Unnatural base pairing xenonucleic acids (XNAs) can be used to expand life’s alphabet beyond ATGC. Here, authors show strategies for enzymatic synthesis and next-generation nanopore sequencing of XNA base pairs for reading and writing 12-letter DNA (ATGCBSPZXKJV).
- Hinako Kawabe
- , Christopher A. Thomas
- & Jorge A. Marchand
-
Article
| Open Access Extensive breaking of genetic code degeneracy with non-canonical amino acids
Extensive breaking of genetic code degeneracy with non-canonical amino acids
Genetic code expansion is limited by the degeneracy of the 61 sense codons which encode for only 20 amino acids. Here, the authors show that by combining hyperaccurate ribosomes and in vitro transcribed tRNAs, dramatic and extensive breaking of sense codon degeneracy can be achieved.
- Clinton A. L. McFeely
- , Bipasana Shakya
- & Matthew C. T. Hartman
-
Article
| Open Access Hidden modes of DNA binding by human nuclear receptors
Hidden modes of DNA binding by human nuclear receptors
Nuclear receptors (NR) are drug-responsive master regulators. Here, authors map DNA binding profiles of all human NRs. Their MinSeq Find algorithm identifies masked NR binding sites in genomes and maps ~10% of orphan SNPs linked to numerous diseases.
- Devesh Bhimsaria
- , José A. Rodríguez-Martínez
- & Aseem Z. Ansari
-
Article
| Open Access Chemical evolution of an autonomous DNAzyme with allele-specific gene silencing activity
Chemical evolution of an autonomous DNAzyme with allele-specific gene silencing activity
Low activity currently prevents the wider use of DNA enzymes (DNAzymes). Here the authors report the chemical evolution of a DNAzyme with high catalytic activity under near physiological conditions: the enzyme achieves ~65 turnovers in 30 minutes.
- Kim Nguyen
- , Turnee N. Malik
- & John C. Chaput
-
Article
| Open Access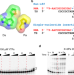 Structural basis of transcription recognition of a hydrophobic unnatural base pair by T7 RNA polymerase
Structural basis of transcription recognition of a hydrophobic unnatural base pair by T7 RNA polymerase
T7 RNA polymerase (RNAP) is widely used for synthesizing RNA molecules with synthetic modifications and unnatural base pairs (UBPs). Here, authors show the structural basis of how UBPs are recognized as template and substrate, providing mechanistic insights into UBP transcription by T7 RNAP.
- Juntaek Oh
- , Michiko Kimoto
- & Dong Wang
-
Article
| Open Access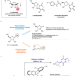 Additive-controlled asymmetric iodocyclization enables enantioselective access to both α- and β-nucleosides
Additive-controlled asymmetric iodocyclization enables enantioselective access to both α- and β-nucleosides
Nucleosides and their analogs are pharmacologically important molecules. Here, the authors report an additive-controlled stereodivergent iodocyclization method for the selective synthesis of α- or β-nucleosides.
- Qi Wang
- , Jiayi Mu
- & Fener Chen
-
Article
| Open Access Shedding light on the base-pair opening dynamics of nucleic acids in living human cells
Shedding light on the base-pair opening dynamics of nucleic acids in living human cells
Base-pair opening is important for nucleic acids to exert biological functions, but studying its dynamics inside living cells is challenging. Here, the authors determine the base-pair opening kinetics of hairpin and G-quadruplex structures inside living human cells by the in-cell NMR technique, and demonstrate a difference in dynamics of nucleic acids between in-cell and in vitro conditions.
- Yudai Yamaoki
- , Takashi Nagata
- & Masato Katahira
-
Article
| Open Access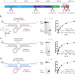 XNAzymes targeting the SARS-CoV-2 genome inhibit viral infection
XNAzymes targeting the SARS-CoV-2 genome inhibit viral infection
RNA viruses have been responsible for large-scale epidemics and pandemics throughout the last few centuries. Here, the authors show the design, synthesis and screening of artificial RNA endonuclease XNAzymes capable of cleaving genomic SARS-CoV-2 RNA and self-assembling into enzymatic nanostructures inhibiting cellular viral replication.
- Pehuén Pereyra Gerber
- , Maria J. Donde
- & Alexander I. Taylor
-
Article
| Open Access HIF1α-AS1 is a DNA:DNA:RNA triplex-forming lncRNA interacting with the HUSH complex
HIF1α-AS1 is a DNA:DNA:RNA triplex-forming lncRNA interacting with the HUSH complex
Using a composite bioinformatics approach, the DNA:DNA:RNA triplex-forming lncRNAs HIF1α-AS1 was identified in human endothelial cells which recruits an epigenetic silencing complex to limit expression of triplex target genes.
- Matthias S. Leisegang
- , Jasleen Kaur Bains
- & Ralf P. Brandes
-
Article
| Open Access Microfluidic space coding for multiplexed nucleic acid detection via CRISPR-Cas12a and recombinase polymerase amplification
Microfluidic space coding for multiplexed nucleic acid detection via CRISPR-Cas12a and recombinase polymerase amplification
Fast, low-cost and multiplexed nucleic acid detection is challenging. Here the authors report a strategy that couples microfluidic space coding, CRISPRCas12a, and multiplex RPA for the rapid detection of up to 30 targets with only one fluorescent probe.
- Zhichen Xu
- , Dongjuan Chen
- & Maili Liu
-
Article
| Open Access Controllable DNA hybridization by host–guest complexation-mediated ligand invasion
Controllable DNA hybridization by host–guest complexation-mediated ligand invasion
Direct dissociation of nucleic acid duplex structures without heating or specific binding proteins is challenging. Here the authors use the cucurbit[7]uril-based host–guest system to construct a ligand-invasion pathway for controllable DNA hybridisation.
- Lin Xiao
- , Liang-Liang Wang
- & Liang Xu
-
Article
| Open Access Efficient DNA fluorescence labeling via base excision trapping
Efficient DNA fluorescence labeling via base excision trapping
Methods for fluorescently labelling DNAs are expensive and labour-intensive. Here the authors report an in situ DNA labelling strategy for oligonucleotides as well as dsDNA that makes use of aldehyde-reactive rotor dyes to trap AP sites resulting from excision of deaminated DNA bases.
- Yong Woong Jun
- , Emily M. Harcourt
- & Eric T. Kool
-
Article
| Open Access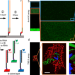 Simple synthesis of massively parallel RNA microarrays via enzymatic conversion from DNA microarrays
Simple synthesis of massively parallel RNA microarrays via enzymatic conversion from DNA microarrays
RNA microarrays have many potential applications, but are difficult to produce. Here, the AUs present a method for converting commercial, customizable DNA microarrays into RNA microarrays using an accessible three-step process involving primer photocrosslinking, extension, and template degradation.
- Erika Schaudy
- , Kathrin Hölz
- & Mark M. Somoza
-
Article
| Open Access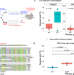 Targeting double-strand break indel byproducts with secondary guide RNAs improves Cas9 HDR-mediated genome editing efficiencies
Targeting double-strand break indel byproducts with secondary guide RNAs improves Cas9 HDR-mediated genome editing efficiencies
Programmable double-strand DNA breaks (DSBs) can be harnessed for precision genome editing through manipulation of the homology-directed repair (HDR) pathway. Here the authors report the development of the double tap - double tap implements secondary gRNAs which target Cas9 to common indel sequences and provides a second chance at HDR.
- Zsolt Bodai
- , Alena L. Bishop
- & Alexis C. Komor
-
Article
| Open Access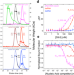 Structural basis of R-loop recognition by the S9.6 monoclonal antibody
Structural basis of R-loop recognition by the S9.6 monoclonal antibody
The S9.6 monoclonal antibody is widely used to map R-loops genome wide. Here, Bou-Nader et al., define the nucleic acid-binding specificity of S9.6 and report its crystal structures free and bound to a hybrid, which reveal the asymmetric recognition of the RNA and DNA strands and its A-form conformation.
- Charles Bou-Nader
- , Ankur Bothra
- & Jinwei Zhang
-
Article
| Open Access Gene editing with CRISPR-Cas12a guides possessing ribose-modified pseudoknot handles
Gene editing with CRISPR-Cas12a guides possessing ribose-modified pseudoknot handles
Development of Cas12a for human therapeutics and diagnostics may significantly benefit from, or even require, chemical modification of its guide RNA. Here the authors show that the noncanonical 5′ pseudoknot structure of the AsCas12a crRNA guide can be heavily modified and still retain very high editing activity in cells and trans cleavage activity in vitro.
- Eman A. Ageely
- , Ramadevi Chilamkurthy
- & Keith T. Gagnon
-
Article
| Open Access A chemical probe based on the PreQ1 metabolite enables transcriptome-wide mapping of binding sites
A chemical probe based on the PreQ1 metabolite enables transcriptome-wide mapping of binding sites
The small modified nucleotide PreQ1 binds to the PreQ1 riboswitch and regulates gene expression by inducing RNA conformation change. Here the authors design and characterize a specific preQ1-derived probe by x-ray crystallography, mass spectral analysis and transcriptome-wide using Chem-CLIP.
- Sumirtha Balaratnam
- , Curran Rhodes
- & John S. Schneekloth Jr
-
Article
| Open Access A non-enzymatic, isothermal strand displacement and amplification assay for rapid detection of SARS-CoV-2 RNA
A non-enzymatic, isothermal strand displacement and amplification assay for rapid detection of SARS-CoV-2 RNA
The reliance on enzymes in SARS-CoV-2 RNA detection imposes limits on transport and storage conditions. Here the authors use non-enzymatic isothermal amplification to detect RNA with no need for reverse transcription.
- Mohsen Mohammadniaei
- , Ming Zhang
- & Yi Sun
-
Article
| Open Access A kinetically controlled platform for ligand-oligonucleotide transduction
A kinetically controlled platform for ligand-oligonucleotide transduction
Ligand-oligonucleotide interactions can integrate both small molecules and proteins into nucleic acid-based circuits. Here the authors design ligand-aptamer complexes to control strand-displacement reactions for versatile ligand transduction.
- Qiu-Long Zhang
- , Liang-Liang Wang
- & Liang Xu
-
Article
| Open Access Fully automated fast-flow synthesis of antisense phosphorodiamidate morpholino oligomers
Fully automated fast-flow synthesis of antisense phosphorodiamidate morpholino oligomers
PMOs (phosphorodiamidate morpholino oligomers) have huge potential for antisense therapy but complex and slow synthesis limits application. Here, the authors report the development of automated flow synthesis methods which reduce nucleobase coupling times from hours to minutes removing human errors and allow for high-throughput production.
- Chengxi Li
- , Alex J. Callahan
- & Bradley L. Pentelute
-
Article
| Open Access Nonenzymatic polymerase-like template-directed synthesis of acyclic l-threoninol nucleic acid
Nonenzymatic polymerase-like template-directed synthesis of acyclic l-threoninol nucleic acid
A world preceding the prebiotic RNA-world may have been based on xeno nucleic acids (XNAs), but their replication likely did not require enzymes. Here, the authors demonstrate template-directed non-enzymatic synthesis of an XNA, acyclic l-threoninol nucleic acid, via chemical ligation mediated by N-cyanoimidazole, and achieve a pseudo-primer extension of this XNA with all four nucleobases.
- Keiji Murayama
- , Hikari Okita
- & Hiroyuki Asanuma
-
Article
| Open Access Optoribogenetic control of regulatory RNA molecules
Optoribogenetic control of regulatory RNA molecules
Short hairpin RNAs can be used to modulate and regulate gene expression. Here the authors generate chimeric RNAs that interact with the photoreceptor PAL, allowing for optoribogenetic control of cell physiology.
- Sebastian Pilsl
- , Charles Morgan
- & Günter Mayer
-
Article
| Open Access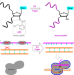 Conditional control of RNA-guided nucleic acid cleavage and gene editing
Conditional control of RNA-guided nucleic acid cleavage and gene editing
Constituitively active CRISPR systems have the risk of adverse off-target effects. Here the authors use chemical masking and activation of gRNA to control activity.
- Shao-Ru Wang
- , Ling-Yu Wu
- & Xiang Zhou
-
Article
| Open Access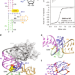 SAM-VI riboswitch structure and signature for ligand discrimination
SAM-VI riboswitch structure and signature for ligand discrimination
Riboswitches are conserved RNA domains located in the non-coding region of mRNA that recognize cellular metabolites and, in turn, regulate gene expression. Here the authors report the structure of the recently identified SAM-VI riboswitch and provide insight into its mechanism of ligand discrimination.
- Aiai Sun
- , Catherina Gasser
- & Aiming Ren
-
Article
| Open Access Time-resolved NMR monitoring of tRNA maturation
Time-resolved NMR monitoring of tRNA maturation
Transfer RNA (tRNA) is regulated by RNA modifications. Here the authors employ time-resolved NMR to monitor modifications of yeast tRNAPhe in cellular extracts, revealing a sequential order and cross-talk between modifications.
- Pierre Barraud
- , Alexandre Gato
- & Carine Tisné
-
Article
| Open Access An artificial triazole backbone linkage provides a split-and-click strategy to bioactive chemically modified CRISPR sgRNA
An artificial triazole backbone linkage provides a split-and-click strategy to bioactive chemically modified CRISPR sgRNA
For CRISPR-Cas9 genome editing, Cas9 protein is guided to its target by single guide (sg) RNA. Here, the authors synthesised sgRNAs via convergent ‘click’ ligation of variable 20-mer RNAs that target the genome and a Cas9-binding 79-mer chimeric RNA/2´-OMe RNA of fixed sequence in a single tube.
- Lapatrada Taemaitree
- , Arun Shivalingam
- & Tom Brown
-
Article
| Open Access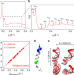 Helical antimicrobial peptides assemble into protofibril scaffolds that present ordered dsDNA to TLR9
Helical antimicrobial peptides assemble into protofibril scaffolds that present ordered dsDNA to TLR9
Amphihelical antimicrobial peptides (AMPs) are bactericidal host defense factors, but their function as immunomodulators is emerging. Here the authors show that several AMPs organize DNA into periodic nanocrystals by self-assembling into superhelical protofibril scaffolds, which potentiates DNA sensing by TLR9.
- Ernest Y. Lee
- , Changsheng Zhang
- & Gerard C. L. Wong
-
Article
| Open Access Template-directed RNA polymerization and enhanced ribozyme catalysis inside membraneless compartments formed by coacervates
Template-directed RNA polymerization and enhanced ribozyme catalysis inside membraneless compartments formed by coacervates
Membraneless compartments have been theorized to be prebiotic micro-compartments as they spontaneously encapsulate RNA and proteins. Here, the authors report membraneless compartments can enhance RNA chemistries, affecting template directed RNA polymerization and stimulating nucleic acid enzymes.
- Raghav R. Poudyal
- , Rebecca M. Guth-Metzler
- & Philip C. Bevilacqua
-
Comment
| Open Access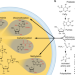 Non-canonical nucleosides and chemistry of the emergence of life
Non-canonical nucleosides and chemistry of the emergence of life
- Sidney Becker
- , Christina Schneider
- & Thomas Carell
-
Comment
| Open AccessPrebiotic plausibility and networks of paradox-resolving independent models
- Steven A. Benner
-
Comment
| Open Access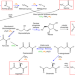 Life as a guide to prebiotic nucleotide synthesis
Life as a guide to prebiotic nucleotide synthesis
- Stuart A. Harrison
- & Nick Lane
-
-
Comment
| Open Access Searching for lost nucleotides of the pre-RNA World with a self-refining model of early Earth
Searching for lost nucleotides of the pre-RNA World with a self-refining model of early Earth
- Nicholas V. Hud
-
Comment
| Open Access Prebiotic nucleic acids need space to grow
Prebiotic nucleic acids need space to grow
- Daniel Whitaker
- & Matthew W. Powner
-
Comment
| Open Access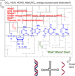 Experimentally investigating the origin of DNA/RNA on early Earth
Experimentally investigating the origin of DNA/RNA on early Earth
- Ramanarayanan Krishnamurthy
-
Article
| Open Access Translation of non-standard codon nucleotides reveals minimal requirements for codon-anticodon interactions
Translation of non-standard codon nucleotides reveals minimal requirements for codon-anticodon interactions
The recognition of the mRNA codon by the tRNA anticodon is crucial for protein synthesis. Here the authors introduce non-standard nucleotides in bacterial and eukaryotic mRNA to reveal the minimal hydrogen bond requirement of codon-anticodon interaction for efficient and accurate translation.
- Thomas Philipp Hoernes
- , Klaus Faserl
- & Matthias David Erlacher
-
Article
| Open Access Chemical and structural studies provide a mechanistic basis for recognition of the MYC G-quadruplex
Chemical and structural studies provide a mechanistic basis for recognition of the MYC G-quadruplex
Targeting noncoding nucleic acids with small molecules represents an important and significant challenge in chemical biology and drug discovery. Here the authors characterize DC-34, a small molecule that exhibits selective binding to specific G4 structures, and provide a structural basis for its selectivity
- David R. Calabrese
- , Xiang Chen
- & John S. Schneekloth Jr.
-
Article
| Open Access An anionic phthalocyanine decreases NRAS expression by breaking down its RNA G-quadruplex
An anionic phthalocyanine decreases NRAS expression by breaking down its RNA G-quadruplex
Hyperactivity of the gene NRAS contributes to the proliferation and metastatic nature of many types of cancer cells. Here, the authors show that NRAS can be controlled by an anionic phthalocyanine coordinating Zn2+ in combination with photo-irradiation.
- Keiko Kawauchi
- , Wataru Sugimoto
- & Daisuke Miyoshi
-
Article
| Open Access Modular cell-internalizing aptamer nanostructure enables targeted delivery of large functional RNAs in cancer cell lines
Modular cell-internalizing aptamer nanostructure enables targeted delivery of large functional RNAs in cancer cell lines
Large RNAs and ribonucleoprotein complexes have shown potential as novel therapeutic agents, but their targeted delivery to cells is still challenging. Here the authors present a modular aptamer nanostructure for intracellular delivery of RNAs up to 250 nucleotides to cancer cells.
- David Porciani
- , Leah N. Cardwell
- & Donald H. Burke
-
Article
| Open Access Comprehensive profiling of the ligand binding landscapes of duplexed aptamer families reveals widespread induced fit
Comprehensive profiling of the ligand binding landscapes of duplexed aptamer families reveals widespread induced fit
Duplexed aptamers are a common biosensor format; however, how complementary strand sequence, length, and position modulate ligand binding is not well understood. Here, the authors introduce ACE-Scan to comprehensively map binding landscapes, uncovering hotspots of enhanced binding by induced fit.
- Jeffrey D. Munzar
- , Andy Ng
- & David Juncker
-
Article
| Open Access Structural basis for TNA synthesis by an engineered TNA polymerase
Structural basis for TNA synthesis by an engineered TNA polymerase
The laboratory-evolved polymerase Kod-RI catalyzes α-L-threose nucleic acid (TNA) synthesis. Here, the authors present Kod-RI crystal structures that give insights into how TNA triphosphates are selected and extended in a template-dependent manner, which will help to engineer improved TNA polymerases for synthetic genetics applications.
- Nicholas Chim
- , Changhua Shi
- & John C. Chaput
-
Article
| Open Access Target guided synthesis using DNA nano-templates for selectively assembling a G-quadruplex binding c-MYC inhibitor
Target guided synthesis using DNA nano-templates for selectively assembling a G-quadruplex binding c-MYC inhibitor
Identification of inhibitors can be accelerated by using the target as a template for ligand formation. Here the authors show that DNA-functionalised magnetic nanoparticles guide templating of G-quadruplex bindingc-MYCinhibitors from an array of building blocks, and can be isolated by magnetic decanting.
- Deepanjan Panda
- , Puja Saha
- & Jyotirmayee Dash
-
Article
| Open Access Activating frataxin expression by repeat-targeted nucleic acids
Activating frataxin expression by repeat-targeted nucleic acids
Expansion of the trinucleotide GAA within an intronic FXN RNA can cause Friedreich's Ataxia (FRDA), an incurable genetic disorder. Here, the authors show that anti-GAA duplex RNAs or single-stranded locked nucleic acids increases FXN protein expression in patient-derived cells to levels similar to wild-type cells.
- Liande Li
- , Masayuki Matsui
- & David R. Corey
-
Article
| Open Access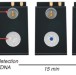 Ultrasensitive visual read-out of nucleic acids using electrocatalytic fluid displacement
Ultrasensitive visual read-out of nucleic acids using electrocatalytic fluid displacement
Point-of-care analytical devices are of interest for diagnostic applications where larger scale laboratory instruments are not feasible or available. Here, the authors present a direct read-out colorimetric sensor which uses catalytic gas production to visualize picomolar concentrations of DNA.
- Justin D. Besant
- , Jagotamoy Das
- & Shana O. Kelley
-
Article
| Open Access Freely orbiting magnetic tweezers to directly monitor changes in the twist of nucleic acids
Freely orbiting magnetic tweezers to directly monitor changes in the twist of nucleic acids
Rotational motion and torsional strain affects DNA replication, transcription and repair. Lipfertet al. have developed a new technique that uses freely orbiting magnetic tweezers to measure equilibrium fluctuations and determine the twist of tethered nucleic acid molecules.
- Jan Lipfert
- , Matthew Wiggin
- & Nynke H. Dekker

 Expedient production of site specifically nucleobase-labelled or hypermodified RNA with engineered thermophilic DNA polymerases
Expedient production of site specifically nucleobase-labelled or hypermodified RNA with engineered thermophilic DNA polymerases