Featured
-
-
Article
| Open Access The Burkholderia pseudomallei intracellular ‘TRANSITome’
The Burkholderia pseudomallei intracellular ‘TRANSITome’
Prokaryotic cell transcriptomics has been limited to mixed or sub-population dynamics and individual cells within heterogeneous populations. Here the authors develop a ‘TRANSITomic’ approach to profile transcriptomes of single Burkholderia pseudomallei cells as they transit through host cell infection.
- Yun Heacock-Kang
- , Ian A. McMillan
- & Tung T. Hoang
-
Article
| Open Access Machine learning uncovers independently regulated modules in the Bacillus subtilis transcriptome
Machine learning uncovers independently regulated modules in the Bacillus subtilis transcriptome
The systems-level regulatory structure underlying gene expression in bacteria can be inferred using machine learning algorithms. Here we show this structure for Bacillus subtilis, present five hypotheses gleaned from it, and analyse the process of sporulation from its perspective.
- Kevin Rychel
- , Anand V. Sastry
- & Bernhard O. Palsson
-
Article
| Open Access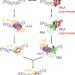 Single-molecule live-cell imaging visualizes parallel pathways of prokaryotic nucleotide excision repair
Single-molecule live-cell imaging visualizes parallel pathways of prokaryotic nucleotide excision repair
In Escherichia coli, the UvrAB damage sensor recognizes helix-distorting lesions by itself or via Mfd bound to stalled RNA polymerase. Here authors use single-molecule fluorescence imaging to quantify the kinetic signatures of interactions of UvrA with Mfd and UvrB in live cells.
- Harshad Ghodke
- , Han Ngoc Ho
- & Antoine M. van Oijen

 Ultrastructure of macromolecular assemblies contributing to bacterial spore resistance revealed by in situ cryo-electron tomography
Ultrastructure of macromolecular assemblies contributing to bacterial spore resistance revealed by in situ cryo-electron tomography