Article
|
Open Access
Featured
-
-
Article
| Open Access Tyrosine phosphorylation of CARM1 promotes its enzymatic activity and alters its target specificity
Tyrosine phosphorylation of CARM1 promotes its enzymatic activity and alters its target specificity
Coactivator-associated arginine methyltransferase 1 (CARM1) is an important target in hematologic malignancies. In this work, the authors show that the hyperactivation of Janus kinase 2 (JAK2) by the V617F mutation phosphorylates CARM1 which regulates its methyltransferase activity and alters its target specificity.
- Hidehiro Itonaga
- , Adnan K. Mookhtiar
- & Stephen D. Nimer
-
Article
| Open Access Loss-of-function mutation in PRMT9 causes abnormal synapse development by dysregulation of RNA alternative splicing
Loss-of-function mutation in PRMT9 causes abnormal synapse development by dysregulation of RNA alternative splicing
Mutations in protein arginine methyltransferase 9 (PRMT9) are linked to intellectual disability. Here, the authors show that mutant PRMT9 fails to methylate its primary substrate SF3B2, causing aberrant RNA splicing and abnormal synapse development.
- Lei Shen
- , Xiaokuang Ma
- & Yanzhong Yang
-
Article
| Open Access The Mycobacterium tuberculosis methyltransferase Rv2067c manipulates host epigenetic programming to promote its own survival
The Mycobacterium tuberculosis methyltransferase Rv2067c manipulates host epigenetic programming to promote its own survival
Singh et al. show how the M. tuberculosis methyltransferase Rv2067c outsmarts host epigenetic machinery by methylating histone H3 prior to its assembly into nucleosomes, thereby ensuring the pathogen’s intracellular survival/success.
- Prakruti R. Singh
- , Venkatareddy Dadireddy
- & Valakunja Nagaraja
-
Article
| Open Access PRMT2 promotes HIV-1 latency by preventing nucleolar exit and phase separation of Tat into the Super Elongation Complex
PRMT2 promotes HIV-1 latency by preventing nucleolar exit and phase separation of Tat into the Super Elongation Complex
The mechanisms regulating Tat function for HIV-1 replication remain poorly understood. Here, the authors reveal that the transcriptional activity of Tat is modulated by a PRMT2-licensed switch between its nucleolar sequestration and phase separation into the nucleoplasmic Super Elongation Complex (SEC) droplets.
- Jiaxing Jin
- , Hui Bai
- & Deqing Hu
-
Article
| Open Access MePMe-seq: antibody-free simultaneous m6A and m5C mapping in mRNA by metabolic propargyl labeling and sequencing
MePMe-seq: antibody-free simultaneous m6A and m5C mapping in mRNA by metabolic propargyl labeling and sequencing
Methylation is the dominant modification in mRNA and occurs at a variety of sites. Here, Hartstock et al. show that a clickable analogue of the key cosubstrate S-adenosyl-L-methionine (SAM) can be produced in cells, allowing for identification and mapping of different methylated nucleosides in mRNA.
- Katja Hartstock
- , Nadine A. Kueck
- & Andrea Rentmeister
-
Article
| Open Access An autoinhibited state of 53BP1 revealed by small molecule antagonists and protein engineering
An autoinhibited state of 53BP1 revealed by small molecule antagonists and protein engineering
Here, using small molecule antagonists and protein engineering, the authors identify an autoinhibited state of 53BP1 leading to its chromatin binding surface being obstructed. Such small molecule ligands present a potential avenue for the development of cancer therapy drugs.
- Gaofeng Cui
- , Maria Victoria Botuyan
- & Georges Mer
-
Article
| Open Access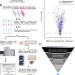 A seven-transmembrane methyltransferase catalysing N-terminal histidine methylation of lytic polysaccharide monooxygenases
A seven-transmembrane methyltransferase catalysing N-terminal histidine methylation of lytic polysaccharide monooxygenases
N-terminal histidine methylation modification has only been observed on certain fungal proteins. Here, the authors identify and validate the methyltransferase responsible for this modification through the combination of mass spectrometry-based proteomics and CRISPR/Cas9.
- Tanveer S. Batth
- , Jonas L. Simonsen
- & Jesper V. Olsen
-
Comment
| Open Access Limited choice of natural amino acids as mimetics restricts design of protein lysine methylation studies
Limited choice of natural amino acids as mimetics restricts design of protein lysine methylation studies
Protein lysine methylation plays important biological roles but its experimental characterization is limited by the lack of suitable mimetics of methylated and unmethylated lysine among the natural amino acids. Here, we summarize the consequent challenges and discuss alternative approaches for biochemical and cellular lysine methylation studies.
- Sara Weirich
- & Albert Jeltsch
-
Article
| Open Access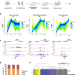 Histone H1 facilitates restoration of H3K27me3 during DNA replication by chromatin compaction
Histone H1 facilitates restoration of H3K27me3 during DNA replication by chromatin compaction
Liu et al. investigated the dynamic re-establishment of H3K27me3 on nascent DNA during DNA replication. They found H1-mediated chromatin compaction facilitates the propagation and restoration of H3K27me3 after DNA replication.
- Cuifang Liu
- , Juan Yu
- & Guohong Li
-
Article
| Open Access ASH1L-MRG15 methyltransferase deposits H3K4me3 and FACT for damage verification in nucleotide excision repair
ASH1L-MRG15 methyltransferase deposits H3K4me3 and FACT for damage verification in nucleotide excision repair
Due to the naturally dense packing of the genome, DNA repair factors encounter a topologically very intricate cellular context. We describe how the human ASH1L protein navigates excision repair factors to accelerate their search for DNA lesions.
- Corina Maritz
- , Reihaneh Khaleghi
- & Hanspeter Naegeli
-
Article
| Open Access Decoding protein methylation function with thermal stability analysis
Decoding protein methylation function with thermal stability analysis
Methylation is a common modification that affects protein function but, compared to other modifications, our knowledge is limited. Here, the authors use a method based on thermal stability to study how protein methylation regulates processes such as mRNA binding proteins and chromosome compaction.
- Cristina Sayago
- , Jana Sánchez-Wandelmer
- & Javier Munoz
-
Article
| Open Access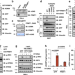 PRMT1 mediated methylation of cGAS suppresses anti-tumor immunity
PRMT1 mediated methylation of cGAS suppresses anti-tumor immunity
cGAS/STING mediated immunity is linked to the anti-tumor response, but how tumor-intrinsic cGAS signals are countered during tumorigenesis and immune evasion is poorly understood. Here the authors show PRMT1 suppresses the anti-tumor immune response via arginine methylation of cGAS.
- Jing Liu
- , Xia Bu
- & Wenyi Wei
-
Article
| Open Access Arabidopsis TRB proteins function in H3K4me3 demethylation by recruiting JMJ14
Arabidopsis TRB proteins function in H3K4me3 demethylation by recruiting JMJ14
TRB proteins have been shown to recruit PRC2 for H3K27me3 deposition. This work shows that TRBs also recruit JMJ14 to remove H3K4me3, demonstrating that TRBs silence the target genes via a combinatorial histone modification mechanism.
- Ming Wang
- , Zhenhui Zhong
- & Steven E. Jacobsen
-
Article
| Open Access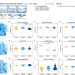 Paternal methotrexate exposure affects sperm small RNA content and causes craniofacial defects in the offspring
Paternal methotrexate exposure affects sperm small RNA content and causes craniofacial defects in the offspring
Anti-folate drugs, such as methotrexate, have been largely prohibited for pregnant women because of the teratogenic effect on their descendant. Here, the authors report a intergenerational mechanism by why paternal methotrexate exposure causes craniofacial defects on their offspring.
- Nagif Alata Jimenez
- , Mauricio Castellano
- & Pablo H. Strobl-Mazzulla
-
Article
| Open Access Systematic characterization of chromodomain proteins reveals an H3K9me1/2 reader regulating aging in C. elegans
Systematic characterization of chromodomain proteins reveals an H3K9me1/2 reader regulating aging in C. elegans
Chromodomains mainly function as histone methyl-lysine readers to regulate gene expression. Here the authors delineate a functional map of chromodomain proteins and identify an H3K9me1/2 writer, MET-2, and reader, CEC-5, that are required for the normal lifespan of C. elegans.
- Xinhao Hou
- , Mingjing Xu
- & Xuezhu Feng
-
Article
| Open Access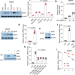 N6-methyladenosine of Spi2a attenuates inflammation and sepsis-associated myocardial dysfunction in mice
N6-methyladenosine of Spi2a attenuates inflammation and sepsis-associated myocardial dysfunction in mice
N6-methyladenosine (m6A) methylation of RNA is a common type of RNA modification that regulates gene expression. Here, the authors report m6A methylation of serpin 2 A negatively regulates inflammation and reduces heart dysfunction in the setting of sepsis.
- Xiangyu Wang
- , Yan Ding
- & Jie Du
-
Article
| Open Access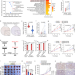 PHGDH arginine methylation by PRMT1 promotes serine synthesis and represents a therapeutic vulnerability in hepatocellular carcinoma
PHGDH arginine methylation by PRMT1 promotes serine synthesis and represents a therapeutic vulnerability in hepatocellular carcinoma
The role of protein arginine methylation in serine metabolism of cancer cells in hepatocellular carcinoma (HCC) remains to be explored. Here, the authors show that phosphoglycerate dehydrogenase (PHGDH) is activated by PRMT1-mediated R236 methylation, promoting serine synthesis, redox homeostasis and HCC growth.
- Kui Wang
- , Li Luo
- & Canhua Huang
-
Article
| Open Access Pivotal role for S-nitrosylation of DNA methyltransferase 3B in epigenetic regulation of tumorigenesis
Pivotal role for S-nitrosylation of DNA methyltransferase 3B in epigenetic regulation of tumorigenesis
Here the authors demonstrate that de novo DNA methylation mediated by DNMT3B is regulated by nitric oxide (NO). They also isolate a unique modulator (DBIC) that inhibits S-nitrosylation of DNMT3B, which mitigates cell proliferation and tumorigenic conversion in vivo.
- Kosaku Okuda
- , Kengo Nakahara
- & Takashi Uehara
-
Article
| Open Access The NFIB/CARM1 partnership is a driver in preclinical models of small cell lung cancer
The NFIB/CARM1 partnership is a driver in preclinical models of small cell lung cancer
Protein arginine methylation can contribute to tumor progression. Here the authors show that protein arginine methyltransferase, CARM1, methylates transcription factor NFIB to promote the growth of small cell lung cancers.
- Guozhen Gao
- , Simone Hausmann
- & Mark T. Bedford
-
Article
| Open Access A specific G9a inhibitor unveils BGLT3 lncRNA as a universal mediator of chemically induced fetal globin gene expression
A specific G9a inhibitor unveils BGLT3 lncRNA as a universal mediator of chemically induced fetal globin gene expression
This study describes RK-701, an inhibitor of histone methyltransferases G9a/GLP, as a promising therapeutic candidate for sickle cell disease and a universal role of BGLT3 lncRNA in fetal hemoglobin reactivation by chemical inducers including RK-701.
- Shohei Takase
- , Takashi Hiroyama
- & Akihiro Ito
-
Article
| Open Access A small molecule antagonist of SMN disrupts the interaction between SMN and RNAP II
A small molecule antagonist of SMN disrupts the interaction between SMN and RNAP II
The SMN protein recognizes symmetric dimethylarginine by its Tudor domain, and SMN deficiency leads to spinal muscular atrophy. Here, Liu et al. discover a small molecule that binds to the SMN Tudor domain and disrupts the interaction between SMN and RNA Polymerase II.
- Yanli Liu
- , Aman Iqbal
- & Jinrong Min
-
Article
| Open Access CHIKV infection reprograms codon optimality to favor viral RNA translation by altering the tRNA epitranscriptome
CHIKV infection reprograms codon optimality to favor viral RNA translation by altering the tRNA epitranscriptome
Viruses completely depend on the host translational machinery, but their genomes are often enriched in rare codons and therefore should be translated with poor efficiency. Here, Jungfleisch et al. apply Ribo-Seq and RNASeq to provide a global view on the translational changes occurring during Chikungunya virus (CHIKV) infection. CHIKV infection induces a codon-specific reprogramming of the host translation machinery to favor the translation of viral RNA genomes over host mRNAs via tRNA modification.
- Jennifer Jungfleisch
- , René Böttcher
- & Juana Díez
-
Article
| Open Access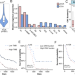 The genomic landscape of cholangiocarcinoma reveals the disruption of post-transcriptional modifiers
The genomic landscape of cholangiocarcinoma reveals the disruption of post-transcriptional modifiers
Cholangiocarcinoma is a heterogenous group of cancers, with large genetic variation seen within subtypes. Here, the authors find 12 significantly mutated genes and 5 focal CNA regions were found in perihilar cholangiocarcinoma, and identified METTL14 to have a potential tumour suppressive role.
- Yaodong Zhang
- , Zijian Ma
- & Xiangcheng Li
-
Article
| Open Access KMT5A-methylated SNIP1 promotes triple-negative breast cancer metastasis by activating YAP signaling
KMT5A-methylated SNIP1 promotes triple-negative breast cancer metastasis by activating YAP signaling
SNIP1 methylation initiates its oncogenic functions. Here, the authors show that SNIP1 is methylated by KMT5A and this leads to downstream signalling that activates the YAP pathway, resulting in tumorigenesis and metastasis.
- Bo Yu
- , Jun Su
- & Jianming Tang
-
Article
| Open Access Inducible and reversible RNA N6-methyladenosine editing
Inducible and reversible RNA N6-methyladenosine editing
RNA modifications, including N6-methyladenosine (m6A), have been reported to regulate fundamental RNA processes and properties, and directly linked to various human diseases. Here, the authors develop a chemically inducible and reversible RNA m6A modification editing platform integrating chemically induced proximity (CIP) and CRISPR methods.
- Huaxia Shi
- , Ying Xu
- & Fu-Sen Liang
-
Article
| Open Access Oxygen-sensitive methylation of ULK1 is required for hypoxia-induced autophagy
Oxygen-sensitive methylation of ULK1 is required for hypoxia-induced autophagy
Hypoxia induces mitochondrial clearance and autophagy, although the upstream mechanisms are not well defined. Here, the authors identify that oxygen-sensitive methylation of the key autophagy regulator ULK1 promotes ULK1 activation and subsequent autophagosome formation and mitochondrial clearance.
- Jingyi Li
- , Tao Zhang
- & Rui Liu
-
Article
| Open Access Inhibition of cytoplasmic EZH2 induces antitumor activity through stabilization of the DLC1 tumor suppressor protein
Inhibition of cytoplasmic EZH2 induces antitumor activity through stabilization of the DLC1 tumor suppressor protein
The tumor suppressor gene DLC1 can be downregulated in cancers through genetic and non-genetic mechanisms. Here, the authors show cytoplasmic EZH2 can directly methylate and destabilize the DLC1 protein, and EZH2 inhibition can increase the DLC1 protein half-life at least 20-fold.
- Brajendra K. Tripathi
- , Meghan F. Anderman
- & Douglas R. Lowy
-
Article
| Open Access Arginine methylation and ubiquitylation crosstalk controls DNA end-resection and homologous recombination repair
Arginine methylation and ubiquitylation crosstalk controls DNA end-resection and homologous recombination repair
Post-translational modifications are critical for regulating the DNA damage response. Here, the authors identify a methylation-deubiquitination crosstalk between methyltransferase PRMT1 and deubiquitinase USP11, showing that the enzymes regulate each other’s functions in DNA repair.
- Maria Pilar Sanchez-Bailon
- , Soo-Youn Choi
- & Clare C. Davies
-
Article
| Open Access m6Am-seq reveals the dynamic m6Am methylation in the human transcriptome
m6Am-seq reveals the dynamic m6Am methylation in the human transcriptome
m6Am is a dynamic and reversible RNA modification found on the mRNA cap. Here the authors report m6Am-seq to directly distinguish m6Am from m6A and identify functional methylation sites.
- Hanxiao Sun
- , Kai Li
- & Chengqi Yi
-
Article
| Open Access Arginine methylation promotes siRNA-binding specificity for a spermatogenesis-specific isoform of the Argonaute protein CSR-1
Arginine methylation promotes siRNA-binding specificity for a spermatogenesis-specific isoform of the Argonaute protein CSR-1
The Argonaute protein CSR-1 is essential for fertility and viability in C. elegans. Here the authors show that CSR-1A isoform associates preferentially with small RNAs mapping to spermatogenesis-specific genes while CSR-1B isoform binds small RNAs mapping to oogenesis-specific genes. Arginine methylation of CSR-1A promotes small RNA-binding specificity.
- Dieu An H. Nguyen
- & Carolyn M. Phillips
-
Article
| Open Access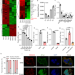 The long non-coding RNA βFaar regulates islet β-cell function and survival during obesity in mice
The long non-coding RNA βFaar regulates islet β-cell function and survival during obesity in mice
Beta-cell function is often impaired in obesity through incompletely understood mechanisms. Here the authors show that the long noncoding RNA βFaar is reduced by diet-induced obesity in mice, which leads to impaired beta-cell function via miR-138-5p and survival via TRAF3 Interacting Protein 2.
- Fangfang Zhang
- , Yue Yang
- & Liang Jin
-
Article
| Open Access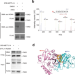 Arginine methylation of METTL14 promotes RNA N6-methyladenosine modification and endoderm differentiation of mouse embryonic stem cells
Arginine methylation of METTL14 promotes RNA N6-methyladenosine modification and endoderm differentiation of mouse embryonic stem cells
The methyltransferase complex of METTL3-METTL14-WTAP is responsible for m6A modification on RNA. Here the authors report that METTL14 arginine 255 (R255) is methylated by PRMT1 and this modification increases interaction of METTL3/METTL14 interaction with WTAP and substrate RNA, promoting m6A methylation activity of the complex.
- Xiaona Liu
- , Hailong Wang
- & Shan Xiao
-
Article
| Open Access PRMT5-mediated arginine methylation activates AKT kinase to govern tumorigenesis
PRMT5-mediated arginine methylation activates AKT kinase to govern tumorigenesis
Arginine methylation has been reported to regulate AKT-associated signalling. Here, the authors show that PRMT5-mediated arginine methylation promotes AKT activity and induces tumourigenesis.
- Shasha Yin
- , Liu Liu
- & Wenjian Gan
-
Article
| Open Access Profiling PRMT methylome reveals roles of hnRNPA1 arginine methylation in RNA splicing and cell growth
Profiling PRMT methylome reveals roles of hnRNPA1 arginine methylation in RNA splicing and cell growth
Arginine methyltransferases (PRMTs) are involved in the regulation of various physiological and pathological conditions. Using proteomics, the authors here profile the methylation substrates of PRMTs 4, 5 and 7 and characterize the roles of these enzymes in cancer-associated splicing regulation.
- Wen-juan Li
- , Yao-hui He
- & Wen Liu
-
Article
| Open Access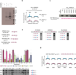 The methyltransferase METTL9 mediates pervasive 1-methylhistidine modification in mammalian proteomes
The methyltransferase METTL9 mediates pervasive 1-methylhistidine modification in mammalian proteomes
Only very few enzymes are known to catalyze protein histidine methylation. Here, the authors show that METTL9 is responsible for most 1-methylhistidine modifications in mouse and human proteomes, and characterize METTL9′s substrate specificity and potential cellular functions.
- Erna Davydova
- , Tadahiro Shimazu
- & Pål Ø. Falnes
-
Article
| Open Access Alpha-ketoglutarate ameliorates age-related osteoporosis via regulating histone methylations
Alpha-ketoglutarate ameliorates age-related osteoporosis via regulating histone methylations
α-ketoglutarate is an intermediate of the Krebs Cycle that was recently reported to extend lifespan in C.Elegans. Here, the authors show that administration of α-ketoglutarate to mice reduces age-related bone loss, by ameliorating senescence of bone-marrow derived mesenchymal stem cells.
- Yuan Wang
- , Peng Deng
- & Quan Yuan
-
Article
| Open Access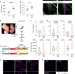 Ybx1 fine-tunes PRC2 activities to control embryonic brain development
Ybx1 fine-tunes PRC2 activities to control embryonic brain development
Polycomb repressive complex 2 (PRC2) methylates H3K27 and suppresses RNA polymerase II transcription by promoting a closed chromatin. Here the authors identify the transcription factor Ybx1 as an interactor that regulates the binding of PRC2 to chromatin and H3K27 methylation to promote the genetic programs underlying neural lineages and neural progenitor self-renewal–differentiation choices.
- Myron K. Evans
- , Yurika Matsui
- & Jamy C. Peng
-
Article
| Open Access Pharmacological inhibition of PRMT7 links arginine monomethylation to the cellular stress response
Pharmacological inhibition of PRMT7 links arginine monomethylation to the cellular stress response
Protein arginine methyltransferases (PRMTs) are increasingly recognized as potential therapeutic targets but PRMT7 remains an understudied member of this enzyme family. Here, the authors develop a chemical probe for PRMT7 and apply it to elucidate the role of PRMT7 in the cellular stress response.
- Magdalena M. Szewczyk
- , Yoshinori Ishikawa
- & Dalia Barsyte-Lovejoy
-
Article
| Open Access Protein lysine 43 methylation by EZH1 promotes AML1-ETO transcriptional repression in leukemia
Protein lysine 43 methylation by EZH1 promotes AML1-ETO transcriptional repression in leukemia
The oncogenic fusion protein AML1-ETO has the ability of AML1 to interact with DNA but blocks AML1-dependent transcription. Here the authors report that histone lysine methyltransferase EZH1 interacts with AML1-ETO and methylates AML1-ETO at lysine 43, promoting AML1-ETO transcriptional repression in leukemia.
- Liping Dou
- , Fei Yan
- & Li Yu
-
Article
| Open Access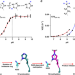 Structural basis for the target specificity of actin histidine methyltransferase SETD3
Structural basis for the target specificity of actin histidine methyltransferase SETD3
SETD3 is the first known metazoan protein histidine methyltransferase but the molecular basis for its target specificity is unclear. Here, the authors elucidate the structural and molecular determinants for the histidine specificity of SETD3.
- Shaobo Dai
- , John R. Horton
- & Xiaodong Cheng
-
Article
| Open Access The Polycomb protein Ezl1 mediates H3K9 and H3K27 methylation to repress transposable elements in Paramecium
The Polycomb protein Ezl1 mediates H3K9 and H3K27 methylation to repress transposable elements in Paramecium
H3K9me3 and H3K27me3 chromatin silencing marks are usually deposited by different SET-domain proteins. Here the authors show that the Enhancer-of-zeste-like protein Ezl1, from the unicellular eukaryote Paramecium tetraurelia, catalyzes methylation of histone H3 in vitro and in vivo with an apparent specificity toward K9 and K27, and controls the repression of transposable elements.
- Andrea Frapporti
- , Caridad Miró Pina
- & Sandra Duharcourt
-
Article
| Open Access SMARCAD1 ATPase activity is required to silence endogenous retroviruses in embryonic stem cells
SMARCAD1 ATPase activity is required to silence endogenous retroviruses in embryonic stem cells
Tight regulation of retrotransposons such as endogenous retroviruses (ERVs) is essential for genome and transcriptome integrity. Here, the authors show that the ATPase function of the chromatin remodeler SMARCAD1 facilitates the binding of KAP1 to ERVs and is required for their repression in embryonic stem cells.
- Parysatis Sachs
- , Dong Ding
- & Jacqueline E. Mermoud
-
Article
| Open Access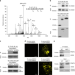 Dual regulation of Arabidopsis AGO2 by arginine methylation
Dual regulation of Arabidopsis AGO2 by arginine methylation
AGO2 is a core component of the RNAi machinery and contributes to plant immunity during bacterial infection. Here the authors show that AGO2 activity is suppressed by arginine methylation which not only promotes AGO2 degradation but also recruits TSN proteins to degrade AGO2-associated small RNAs.
- Po Hu
- , Hongwei Zhao
- & Hailing Jin
-
Article
| Open Access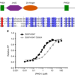 Histone H3 binding to the PHD1 domain of histone demethylase KDM5A enables active site remodeling
Histone H3 binding to the PHD1 domain of histone demethylase KDM5A enables active site remodeling
The demethylase activity of KDM5A is allosterically enhanced by binding of histone H3 to its PHD1 reader domain, through an unknown mechanism. Here the authors show that the PHD1 domain drives ligand-induced allosteric stimulation by stabilizing the binding of substrate to the catalytic domain.
- James E. Longbotham
- , Cynthia M. Chio
- & Danica Galonić Fujimori
-
Article
| Open Access A chemical biology toolbox to study protein methyltransferases and epigenetic signaling
A chemical biology toolbox to study protein methyltransferases and epigenetic signaling
Protein methyltransferases (PMTs) are epigenetic regulatory enzymes with significant therapeutic relevance. Here the authors describe a collection of chemical inhibitors and antagonists to modulate most of the key methylation marks on histones H3 and H4, and use the collection to study of the role of PMTs in mouse and human T cell differentiation.
- Sebastian Scheer
- , Suzanne Ackloo
- & Cheryl H. Arrowsmith
-
Article
| Open Access Cardiac specific PRMT1 ablation causes heart failure through CaMKII dysregulation
Cardiac specific PRMT1 ablation causes heart failure through CaMKII dysregulation
The mechanisms that regulate the activity of Ca2 +/calmodulin-dependent protein kinase II (CaMKII) in the context of heart failure are incompletely understood. Here the authors show that protein arginine methyltransferase 1 (PRMT1) prevents cardiac hyperactivation of CaMKII and heart failure development by methylating CaMKII at arginine residues 9 and 275.
- Jung-Hoon Pyun
- , Hyun-Ji Kim
- & Jong-Sun Kang
-
Article
| Open Access Histone H4K20 methylation mediated chromatin compaction threshold ensures genome integrity by limiting DNA replication licensing
Histone H4K20 methylation mediated chromatin compaction threshold ensures genome integrity by limiting DNA replication licensing
Cell cycle and replication need to be tightly regulated to ensure genome stability in mammalian cells. Here the authors provide a link between chromatin structure and DNA replication regulation by showing that chromatin compaction limits replication licensing thereby promoting genome integrity.
- Muhammad Shoaib
- , David Walter
- & Claus S. Sørensen
-
Article
| Open Access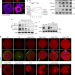 CFP1 coordinates histone H3 lysine-4 trimethylation and meiotic cell cycle progression in mouse oocytes
CFP1 coordinates histone H3 lysine-4 trimethylation and meiotic cell cycle progression in mouse oocytes
The transcription-independent function of trimethylation of histone H3 (H3K4me) in cell division is unclear. Here, Heng-Yu Fan and colleagues report that CFP1, a subunit of the H3K4 methyltransferase, is required for oocyte meiosis, being phosphorylated and degraded during cell cycle transition.
- Qian-Qian Sha
- , Xing-Xing Dai
- & Heng-Yu Fan
-
Article
| Open Access A complex of C9ORF72 and p62 uses arginine methylation to eliminate stress granules by autophagy
A complex of C9ORF72 and p62 uses arginine methylation to eliminate stress granules by autophagy
Many Amyotrophic Lateral Sclerosis (ALS)-linked mutations cause accumulation of stress granules, and most ALS cases are caused by repeat expansions in C9ORF72. Here the authors show that C9ORF72 and the autophagy receptor p62 interact to associate with proteins symmetrically dimethylated on arginines such as FUS, to eliminate stress granules by autophagy.
- Maneka Chitiprolu
- , Chantal Jagow
- & Derrick Gibbings

 Methylation of ESCRT-III components regulates the timing of cytokinetic abscission
Methylation of ESCRT-III components regulates the timing of cytokinetic abscission