Article
|
Open Access
Featured
-
-
Article
| Open Access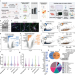 Integrated proteomics reveals autophagy landscape and an autophagy receptor controlling PKA-RI complex homeostasis in neurons
Integrated proteomics reveals autophagy landscape and an autophagy receptor controlling PKA-RI complex homeostasis in neurons
The health of brain cells is known to depend on functional autophagy, but the details are unclear. Here, the authors perform systematic proteomic profiling of human and mouse neurons, delineating the landscape of autophagy degradation in brain.
- Xiaoting Zhou
- , You-Kyung Lee
- & Zhenyu Yue
-
Article
| Open Access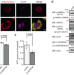 Mitochondrial protein C15ORF48 is a stress-independent inducer of autophagy that regulates oxidative stress and autoimmunity
Mitochondrial protein C15ORF48 is a stress-independent inducer of autophagy that regulates oxidative stress and autoimmunity
Stress-independent autophagy is less understood than stress-induced autophagy and is important for thymic self-tolerance. Here the authors show that a mitochondrial protein C15ORF48 is important for stress-independent autophagy and alters glutathione metabolism and C15orf48 knockout mice develop autoimmunity and changes to thymic epithelial cells.
- Yuki Takakura
- , Moeka Machida
- & Noritaka Yamaguchi
-
Article
| Open Access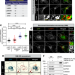 DENND6A links Arl8b to a Rab34/RILP/dynein complex, regulating lysosomal positioning and autophagy
DENND6A links Arl8b to a Rab34/RILP/dynein complex, regulating lysosomal positioning and autophagy
Small GTPases such as Rabs control the positioning of lysosomes. Here, the authors unveil a molecular cascade orchestrated by Arl8/DENND6A/Rab34 that regulates lysosome location, impacting autophagy.
- Rahul Kumar
- , Maleeha Khan
- & Peter S. McPherson
-
Article
| Open Access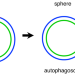 Experimental determination and mathematical modeling of standard shapes of forming autophagosomes
Experimental determination and mathematical modeling of standard shapes of forming autophagosomes
Autophagosome formation involves membrane morphological changes. Here, authors statistically determined average shapes of forming autophagosomes from 3D electron micrographs and established a theoretical model that quantitatively reproduces them.
- Yuji Sakai
- , Satoru Takahashi
- & Noboru Mizushima
-
Article
| Open Access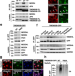 TMEM55B links autophagy flux, lysosomal repair, and TFE3 activation in response to oxidative stress
TMEM55B links autophagy flux, lysosomal repair, and TFE3 activation in response to oxidative stress
Lysosomes are critical regulators of cellular homeostasis. Here, the authors report that the lysosomal protein TMEM55B orchestrates cellular response to acute oxidative stress by coordinating autophagosome degradation, lysosomal repair, and activation of transcriptional stress responses.
- Eutteum Jeong
- , Rose Willett
- & Rosa Puertollano
-
Article
| Open Access NEMO reshapes the α-Synuclein aggregate interface and acts as an autophagy adapter by co-condensation with p62
NEMO reshapes the α-Synuclein aggregate interface and acts as an autophagy adapter by co-condensation with p62
Selective autophagy helps to degrade aggregated proteins accumulating in neurodegenerative diseases. Here, the authors show that NEMO, a ubiquitin binding protein previously linked to innate immune signaling, is recruited to misfolded proteins and promotes their autophagic clearance by forming condensates with the autophagy receptor p62.
- Nikolas Furthmann
- , Verian Bader
- & Konstanze F. Winklhofer
-
Article
| Open Access Autophagy of OTUD5 destabilizes GPX4 to confer ferroptosis-dependent kidney injury
Autophagy of OTUD5 destabilizes GPX4 to confer ferroptosis-dependent kidney injury
Understanding the role of GPX4 in cell ferroptosis at the interface of the inner cortex and medulla is crucial in the context of renal injury. Here, the authors demonstrate that the OTUD5 interaction with GPX4 is key in resisting ischemia/reperfusion-induced ferroptosis in renal cells, offering a new strategy for treating acute kidney injury.
- Li-Kai Chu
- , Xu Cao
- & Jun Liu
-
Article
| Open Access The AMPK-Sirtuin 1-YAP axis is regulated by fluid flow intensity and controls autophagy flux in kidney epithelial cells
The AMPK-Sirtuin 1-YAP axis is regulated by fluid flow intensity and controls autophagy flux in kidney epithelial cells
Urinary flow is sensed by renal cells but its intensity is dysregulated in renal diseases. Here, the authors report that physiological flow inhibits YAP to promote autophagy, while pathological flow leads to YAP activation and autophagy inhibition.
- Aurore Claude-Taupin
- , Pierre Isnard
- & Nicolas Dupont
-
Article
| Open Access Local membrane source gathering by p62 body drives autophagosome formation
Local membrane source gathering by p62 body drives autophagosome formation
Phase separated p62 body plays pivotal roles in autophagy. Here, the authors describe a spatial membrane gathering mode by which p62 body functions in autophagosome formation.
- Xuezhao Feng
- , Daxiao Sun
- & Na Mi
-
Article
| Open Access C9orf72-catalyzed GTP loading of Rab39A enables HOPS-mediated membrane tethering and fusion in mammalian autophagy
C9orf72-catalyzed GTP loading of Rab39A enables HOPS-mediated membrane tethering and fusion in mammalian autophagy
The HOPS complex mediates membrane tethering and autophagosome-lysosome fusion. Here, the authors biochemically reconstitute the mammalian HOPS in protoliposomes and propose a model of complex assembly that depends on Rab2 and Rab39A.
- Shen Zhang
- , Mindan Tong
- & Qing Zhong
-
Article
| Open Access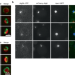 A mechanism that ensures non-selective cytoplasm degradation by autophagy
A mechanism that ensures non-selective cytoplasm degradation by autophagy
How membrane morphology is regulated during autophagosome formation remains elusive. Here, authors reveal a mechanism by which the forming autophagosomal membrane expands with a large opening for non-selective sequestration of the cytoplasm.
- Tetsuya Kotani
- , Yuji Sakai
- & Hitoshi Nakatogawa
-
Article
| Open Access CD44 connects autophagy decline and ageing in the vascular endothelium
CD44 connects autophagy decline and ageing in the vascular endothelium
Mechanisms underlying the connection between autophagy decline and vascular endothelial cell (VEC) ageing remain unclear. Here, the authors identify a key role for CD44 in controlling autophagy and ageing in VECs, and this function is conserved in nematodes.
- Lu Zhang
- , Peichang Yang
- & Shiwei Ma
-
Article
| Open Access A conserved membrane curvature-generating protein is crucial for autophagosome formation in fission yeast
A conserved membrane curvature-generating protein is crucial for autophagosome formation in fission yeast
Rop1 is the single representaive of a subfamily of the membrane-curvature generating REEPs in fission yeast. Wang et al. show that Rop1 is crucial for the macroautophagy of organelles and cytosolic proteins, facilitating autophagosome formation.
- Ning Wang
- , Yoko Shibata
- & Tom A. Rapoport
-
Article
| Open Access ATPase activity of DFCP1 controls selective autophagy
ATPase activity of DFCP1 controls selective autophagy
The endoplasmic reticulum protein DFCP1 is found on omegasomes implicated in autophagosome biogenesis, but its function has remained unknown. Here, Nähse et al. show that DFCP1 is an ATPase that mediates selective autophagy by promoting constriction of large omegasomes.
- Viola Nähse
- , Camilla Raiborg
- & Harald Stenmark
-
Article
| Open Access The mechanisms to dispose of misfolded proteins in the endoplasmic reticulum of adipocytes
The mechanisms to dispose of misfolded proteins in the endoplasmic reticulum of adipocytes
Endoplasmic reticulum (ER)-associated degradation (ERAD) and ER-phagy are two central degradative mechanisms in the ER. Here the authors describe the sequence of events underlying the disposition of misfolded ER proteins by ERAD and ER-phagy.
- Shuangcheng Alivia Wu
- , Chenchen Shen
- & Ling Qi
-
Article
| Open Access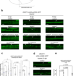 LC3-associated phagocytosis promotes glial degradation of axon debris after injury in Drosophila models
LC3-associated phagocytosis promotes glial degradation of axon debris after injury in Drosophila models
Glia are housekeepers of the nervous system that eliminate neuronal debris after injury. Here, the authors show that LC3-associated phagocytosis in Drosophila glia promotes debris clearance after wing nerve injury and recovery after traumatic brain injury.
- Áron Szabó
- , Virág Vincze
- & Gábor Juhász
-
Article
| Open Access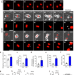 TRABID inhibition activates cGAS/STING-mediated anti-tumor immunity through mitosis and autophagy dysregulation
TRABID inhibition activates cGAS/STING-mediated anti-tumor immunity through mitosis and autophagy dysregulation
cGAS/STING activation is linked to the induction of anti-tumor immune responses. Here the authors report a role for the deubiquitinating enzyme TRABID in regulating mitotic cell division and suppressing anti-tumor immunity, suggesting that TRABID inhibition induces micronuclei and activates cGAS/STING pathway.
- Yu-Hsuan Chen
- , Han-Hsiun Chen
- & Ruey-Hwa Chen
-
Article
| Open Access Redefining the role of AMPK in autophagy and the energy stress response
Redefining the role of AMPK in autophagy and the energy stress response
According to the current understanding in the field, AMPK promotes autophagy by activating ULK1 during energy stress. Here, authors show that AMPK is indeed a negative regulator of ULK1 and it suppresses autophagy in energy depleted cells.
- Ji-Man Park
- , Da-Hye Lee
- & Do-Hyung Kim
-
Article
| Open Access De novo lipogenesis fuels adipocyte autophagosome and lysosome membrane dynamics
De novo lipogenesis fuels adipocyte autophagosome and lysosome membrane dynamics
The function of de novo lipogenesis (DNL) in adipocytes has been a mystery as it contributes little to fat storage in these cells. Here, the authors show that DNL is a critical source of fatty acids for membrane-expanding processes like autophagy.
- Leslie A. Rowland
- , Adilson Guilherme
- & Michael P. Czech
-
Article
| Open Access MYTHO is a novel regulator of skeletal muscle autophagy and integrity
MYTHO is a novel regulator of skeletal muscle autophagy and integrity
Here, the authors identify a novel regulator of autophagy, skeletal muscle mass and integrity named MYTHO. Silencing MYTHO protects against muscle atrophy in a wide range of acute catabolic conditions, while prolonged silencing causes a severe myopathy.
- Jean-Philippe Leduc-Gaudet
- , Anais Franco-Romero
- & Gilles Gouspillou
-
Article
| Open Access Autophagy inhibition prevents lymphatic malformation progression to lymphangiosarcoma by decreasing osteopontin and Stat3 signaling
Autophagy inhibition prevents lymphatic malformation progression to lymphangiosarcoma by decreasing osteopontin and Stat3 signaling
Lymphatic malformation (LM) is a rare, non-malignant vascular abnormality that can progress to lymphangiosarcoma (LAS). The authors use genetic mouse models to show that autophagy inhibition blocks the progression of LM to LAS by decreasing osteopontin expression and Jak/Stat signalling.
- Fuchun Yang
- , Shiva Kalantari
- & Jun-Lin Guan
-
Article
| Open Access Modulating glycosphingolipid metabolism and autophagy improves outcomes in pre-clinical models of myeloma bone disease
Modulating glycosphingolipid metabolism and autophagy improves outcomes in pre-clinical models of myeloma bone disease
Here, the authors show that the glycosylceramide synthesis inhibitor and FDA approved drug Eliglustat inhibits autophagic degradation of TRAF3 which is a key step for osteoclast differentiation and thereby improves myeloma bone lesions.
- Houfu Leng
- , Hanlin Zhang
- & Nicole J. Horwood
-
Article
| Open Access The UFM1 system regulates ER-phagy through the ufmylation of CYB5R3
The UFM1 system regulates ER-phagy through the ufmylation of CYB5R3
The UFM1 system, a ubiquitin-like conjugation system is crucial for endoplasmic reticulum (ER) homeostasis. Here, authors found that CYB5R3 is covalently conjugated with UFM1, which becomes a signal for ER-phagy, a selective autophagy of ER.
- Ryosuke Ishimura
- , Afnan H. El-Gowily
- & Masaaki Komatsu
-
Article
| Open Access The cholesterol transport protein GRAMD1C regulates autophagy initiation and mitochondrial bioenergetics
The cholesterol transport protein GRAMD1C regulates autophagy initiation and mitochondrial bioenergetics
The functions of specific lipids in autophagosome biogenesis are not entirely clear. Here, the authors show that the ER protein GRAMD1C, a cholesterol transport protein, suppresses autophagy initiation and has roles in mitochondrial cholesterol homeostasis and respiration.
- Matthew Yoke Wui Ng
- , Chara Charsou
- & Anne Simonsen
-
Article
| Open Access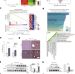 Induction of the hepatic aryl hydrocarbon receptor by alcohol dysregulates autophagy and phospholipid metabolism via PPP2R2D
Induction of the hepatic aryl hydrocarbon receptor by alcohol dysregulates autophagy and phospholipid metabolism via PPP2R2D
Alcohol consumption promotes neutral fat accumulation in the liver. Here, the authors report that alcohol induces aryl hydrocarbon receptor AhR in the liver, and hepatocyte-specific AhR deletion protects against alcohol induced accumulation potentially via transcriptional regulation of Protein phosphatase 2 regulatory subunit Bdelta and subsequent effects on autophagy.
- Yun Seok Kim
- , Bongsub Ko
- & Sang Geon Kim
-
Article
| Open Access Phosphatidylinositol-4-phosphate controls autophagosome formation in Arabidopsis thaliana
Phosphatidylinositol-4-phosphate controls autophagosome formation in Arabidopsis thaliana
Autophagosomes are specialized vesicles that target and deliver cargo to the lytic vacuole. Here the authors show that plasma-membrane derived lipid phosphatidylinositol-4-phosphate supports the assembly and expansion of autophagosomes in Arabidopsis
- Rodrigo Enrique Gomez
- , Clément Chambaud
- & Amélie Bernard
-
Article
| Open Access Compounds activating VCP D1 ATPase enhance both autophagic and proteasomal neurotoxic protein clearance
Compounds activating VCP D1 ATPase enhance both autophagic and proteasomal neurotoxic protein clearance
Several neurodegenerative diseases are characterized by the aggregation of cytoplasmic proteins. Here, the authors demonstrate that the small molecule SMER28 activates VCP, which enhances both autophagic and proteasomal clearance of aggregate-prone proteins.
- Lidia Wrobel
- , Sandra M. Hill
- & David C. Rubinsztein
-
Article
| Open Access Endosomal LC3C-pathway selectively targets plasma membrane cargo for autophagic degradation
Endosomal LC3C-pathway selectively targets plasma membrane cargo for autophagic degradation
Autophagy can selectively target cargo for degradation. Here the authors map the proximal interactome of ATG8-paralogs LC3B and LC3C uncovering an LC3C-Endocytic-Associated-Pathway that selectively recruits internalized plasma membrane cargo, Met and transferrin receptors, to nascent autophagosomes.
- Paula P. Coelho
- , Geoffrey G. Hesketh
- & Morag Park
-
Article
| Open Access Structural mechanism of protein recognition by the FW domain of autophagy receptor Nbr1
Structural mechanism of protein recognition by the FW domain of autophagy receptor Nbr1
Nbr1 recognizes cargos in selective autophagy. Here, authors show filamentous yeast Nbr1 binds Ams1 via an FW domain, and the cryo-EM structure reveals that Nbr1 recognizes the N-terminal di-glycine and tetrameric assembly of Ams1.
- Jianxiu Zhang
- , Ying-Ying Wang
- & Keqiong Ye
-
Article
| Open Access LC3B is an RNA-binding protein to trigger rapid mRNA degradation during autophagy
LC3B is an RNA-binding protein to trigger rapid mRNA degradation during autophagy
LC3/ATG8 plays an essential role in autophagy. Here the authors show that LC3B exhibits RNA-binding ability and induces rapid degradation of target mRNAs via autophagic activation, highlighting the interplay between autophagy and RNA biology.
- Hyun Jung Hwang
- , Hongseok Ha
- & Yoon Ki Kim
-
Article
| Open Access Oxygen-sensitive methylation of ULK1 is required for hypoxia-induced autophagy
Oxygen-sensitive methylation of ULK1 is required for hypoxia-induced autophagy
Hypoxia induces mitochondrial clearance and autophagy, although the upstream mechanisms are not well defined. Here, the authors identify that oxygen-sensitive methylation of the key autophagy regulator ULK1 promotes ULK1 activation and subsequent autophagosome formation and mitochondrial clearance.
- Jingyi Li
- , Tao Zhang
- & Rui Liu
-
Article
| Open Access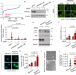 Kansl1 haploinsufficiency impairs autophagosome-lysosome fusion and links autophagic dysfunction with Koolen-de Vries syndrome in mice
Kansl1 haploinsufficiency impairs autophagosome-lysosome fusion and links autophagic dysfunction with Koolen-de Vries syndrome in mice
Here the authors show that the Koolen-de Vries syndrome associated gene KANSL1 modulates autophagosome-lysosome fusion via transcriptional regulation of autophagosomal gene Syntaxin17, and that 13-cis retinoic acid can reverses mitophagic defects and neurobehavioural abnormalities of mice lacking Kansl1.
- Ting Li
- , Dingyi Lu
- & Xin Pan
-
Article
| Open Access The AUTOTAC chemical biology platform for targeted protein degradation via the autophagy-lysosome system
The AUTOTAC chemical biology platform for targeted protein degradation via the autophagy-lysosome system
Targeted protein degradation is a promising approach for basic research and therapeutic applications. Here, the authors develop a targeted protein degradation platform called AUTOTAC to degrade oncoproteins and neurodegeneration-associated proteins via the p62-dependent autophagy-lysosome system.
- Chang Hoon Ji
- , Hee Yeon Kim
- & Yong Tae Kwon
-
Article
| Open Access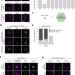 Spatial control of avidity regulates initiation and progression of selective autophagy
Spatial control of avidity regulates initiation and progression of selective autophagy
The molecular principles governing the initiation of autophagosome formation are not clearly understood. Here we show that the vacuolar protein Vac8 coordinates this process by promoting an avidity-driven assembly of several autophagy factors.
- David M. Hollenstein
- , Mariya Licheva
- & Claudine Kraft
-
Article
| Open Access The autophagy protein ATG9A enables lipid mobilization from lipid droplets
The autophagy protein ATG9A enables lipid mobilization from lipid droplets
ATG9A is transmembrane autophagic machinery protein that delivers phospholipids to expanding autophagosomes. Mailler et al. show that ATG9A is required to mobilize lipids from lipid droplets for autophagosome expansion as well as mitochondrial fatty acid import and β-oxidation.
- Elodie Mailler
- , Carlos M. Guardia
- & Juan S. Bonifacino
-
Article
| Open Access Reconstitution defines the roles of p62, NBR1 and TAX1BP1 in ubiquitin condensate formation and autophagy initiation
Reconstitution defines the roles of p62, NBR1 and TAX1BP1 in ubiquitin condensate formation and autophagy initiation
Misfolded proteins are ubquitinated and subsequently condensed by cargo receptors for selective autophagy. Here, the authors use in vitro reconstitution to elegantly dissect how the receptors p62/SQSTM1, NBR1 and TAX1BP1 contribute to p62-ubiquitin condensate formation and degradation by autophagy.
- Eleonora Turco
- , Adriana Savova
- & Sascha Martens
-
Article
| Open Access Gαq activation modulates autophagy by promoting mTORC1 signaling
Gαq activation modulates autophagy by promoting mTORC1 signaling
Nutrient status in the cell regulates autophagy via mTORC1 activity. Here, the authors show that the ubiquitous G protein subunit Gαq contributes to nutrient sensing by promoting formation of an mTOR-p62-Raptor complex in replete conditions, modulating autophagy.
- Sofía Cabezudo
- , Maria Sanz-Flores
- & Catalina Ribas
-
Article
| Open Access VCP maintains nuclear size by regulating the DNA damage-associated MDC1–p53–autophagy axis in Drosophila
VCP maintains nuclear size by regulating the DNA damage-associated MDC1–p53–autophagy axis in Drosophila
Cells maintain a constant cytoplasm to nucleus volume ratio, although the role of DNA damage is not well explored. Here, the authors use Drosophila to connect TER94, the fly homolog of VCP, to disruption of DNA damage repair, leading to ubiquitinated Mu2 protein accumulation and enlarged nuclei.
- Ya-Chu Chang
- , Yu-Xiang Peng
- & Tzu-Kang Sang
-
Article
| Open Access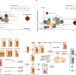 SARS-CoV-2-mediated dysregulation of metabolism and autophagy uncovers host-targeting antivirals
SARS-CoV-2-mediated dysregulation of metabolism and autophagy uncovers host-targeting antivirals
Viruses manipulate host cell pathways to support infection. Here the authors show that SARS-CoV-2 infection modulates cellular metabolism and limits autophagy, and identify druggable host pathways for virus inhibition.
- Nils C. Gassen
- , Jan Papies
- & Marcel A. Müller
-
Article
| Open Access NEK9 regulates primary cilia formation by acting as a selective autophagy adaptor for MYH9/myosin IIA
NEK9 regulates primary cilia formation by acting as a selective autophagy adaptor for MYH9/myosin IIA
Ciliogenesis is a tightly regulated process, although the role of selective autophagy is unclear. Here, the authors show NIMA-related kinase 9 controls actin network stabilization and subsequently ciliogenesis by targeting myosin MYH9 for autophagic degradation via GABARAP interaction.
- Yasuhiro Yamamoto
- , Haruka Chino
- & Noboru Mizushima
-
Article
| Open Access Model-based analysis uncovers mutations altering autophagy selectivity in human cancer
Model-based analysis uncovers mutations altering autophagy selectivity in human cancer
Although autophagy has been linked to tumourigenesis, it is unclear how genomic alterations affect autophagy selectivity in tumours. Here, the authors establish a pipeline that integrates computational and experimental approaches to show that altered autophagy selectivity is frequent in cancer cells and link glycogen autophagy with tumourigenesis.
- Zhu Han
- , Weizhi Zhang
- & Da Jia
-
Article
| Open Access MAEA is an E3 ubiquitin ligase promoting autophagy and maintenance of haematopoietic stem cells
MAEA is an E3 ubiquitin ligase promoting autophagy and maintenance of haematopoietic stem cells
Haematopoietic stem cells (HSCs) are metabolically quiescent, with balanced myeloid and lymphoid potential. Here the authors show that MAEA is required in HSCs for ubiquitination and downregulation of surface cytokine receptors via autophagy; MAEA loss leads to impaired HSC quiescence and a myeloproliferative disorder.
- Qiaozhi Wei
- , Sandra Pinho
- & Paul S. Frenette
-
Article
| Open Access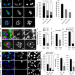 α-Catenin levels determine direction of YAP/TAZ response to autophagy perturbation
α-Catenin levels determine direction of YAP/TAZ response to autophagy perturbation
Autophagy regulates multiple pathways including YAP/TAZ of the Hippo pathway, but precise mechanisms are unclear as autophagy may either activate or inhibit YAP/TAZ. Here, the authors show that autophagy can either activate or regulate YAP/TAZ via dynamic negative feedback loops involving alpha-catenin.
- Mariana Pavel
- , So Jung Park
- & David C. Rubinsztein
-
Article
| Open Access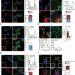 Oxidation inhibits autophagy protein deconjugation from phagosomes to sustain MHC class II restricted antigen presentation
Oxidation inhibits autophagy protein deconjugation from phagosomes to sustain MHC class II restricted antigen presentation
LC3-associated phagocytosis (LAP) is a non-canonical use of the autophagy machinery that can contribute to immune responses. Here, the authors describe the mechanism by which ROS production regulates LAPosome stabilization sustaining MHC II dependent antigen presentation.
- Laure-Anne Ligeon
- , Maria Pena-Francesch
- & Christian Münz
-
Article
| Open Access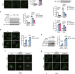 VPS34 K29/K48 branched ubiquitination governed by UBE3C and TRABID regulates autophagy, proteostasis and liver metabolism
VPS34 K29/K48 branched ubiquitination governed by UBE3C and TRABID regulates autophagy, proteostasis and liver metabolism
Autophagy and the ubiquitin–proteasome system (UPS) are cellular quality control processes, but their coordination remains unclear. Here, the authors show that branched ubiquitination of VPS34 functions as a switch between UPS and autophagy and has an important role in lipid metabolism in the liver.
- Yu-Hsuan Chen
- , Tzu-Yu Huang
- & Ruey-Hwa Chen
-
Article
| Open Access An autophagy enhancer ameliorates diabetes of human IAPP-transgenic mice through clearance of amyloidogenic oligomer
An autophagy enhancer ameliorates diabetes of human IAPP-transgenic mice through clearance of amyloidogenic oligomer
Islet amyloid polypeptide (IAPP) deposition is associated with islet cell loss in diabetes. Here the authors show that a small molecule autophagy enhancer reduces IAPP accumulation in vitro, and also improves glucose tolerance in hIAPP+ mice fed high-fat diet, accompanied by reduced hIAPP accumulation, in vivo.
- Jinyoung Kim
- , Kihyoun Park
- & Myung-Shik Lee
-
Article
| Open Access p62/SQSTM1-droplet serves as a platform for autophagosome formation and anti-oxidative stress response
p62/SQSTM1-droplet serves as a platform for autophagosome formation and anti-oxidative stress response
Liquid-liquid phase separation of p62/SQSTM1 has been previously described, although the significance in vivo remains unclear. Here the authors show p62 droplets contain ubiquitin, autophagy-related proteins and Keap1 to serve as platform of not only autophagosome formation but also Nrf2 activation.
- Shun Kageyama
- , Sigurdur Runar Gudmundsson
- & Masaaki Komatsu
-
Article
| Open Access Regulation of cytokine signaling through direct interaction between cytokine receptors and the ATG16L1 WD40 domain
Regulation of cytokine signaling through direct interaction between cytokine receptors and the ATG16L1 WD40 domain
The WD40 domain of ATG16L1 is thought to be involved in non-canonical autophagy. Here the authors screen peptide libraries and identify interactions between this domain and the IL-2Rγ and IL-10RB receptors, indicating endosomal regulation of cytokine signalling by non-canonical autophagy.
- Inmaculada Serramito-Gómez
- , Emilio Boada-Romero
- & Felipe X. Pimentel-Muiños
-
Article
| Open Access Wipi3 is essential for alternative autophagy and its loss causes neurodegeneration
Wipi3 is essential for alternative autophagy and its loss causes neurodegeneration
Unlike canonical macroautophagy, alternative autophagy does not require the factors Atg5 and Atg7. Here, the authors show that Wipi3 is essential for alternative autophagy, but not for canonical autophagy, and that Wipi3 functions to maintain neuronal cells via mechanisms different from those of canonical autophagy.
- Hirofumi Yamaguchi
- , Shinya Honda
- & Shigeomi Shimizu

 Structural basis for the intracellular regulation of ferritin degradation
Structural basis for the intracellular regulation of ferritin degradation