Featured
-
-
Article
| Open Access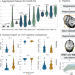 Automated temporalis muscle quantification and growth charts for children through adulthood
Automated temporalis muscle quantification and growth charts for children through adulthood
Temporalis muscle thickness is a promising marker of lean muscle mass but has had limited utility due to its unknown normal growth trajectory and lack of standardized measurement. Here, the authors develop an automated deep learning pipeline to accurately measure temporalis muscle thickness from routine brain magnetic resonance imaging.
- Anna Zapaishchykova
- , Kevin X. Liu
- & Benjamin H. Kann
-
Article
| Open Access Spatial-linked alignment tool (SLAT) for aligning heterogenous slices
Spatial-linked alignment tool (SLAT) for aligning heterogenous slices
Spatial omics technologies reveal the organisation of cells in various biological systems. Here, authors propose SLAT, a graph-based algorithm for aligning heterogenous data across technologies, modalities and timepoints, enabling spatiotemporal reconstruction of complex developmental processes.
- Chen-Rui Xia
- , Zhi-Jie Cao
- & Ge Gao
-
Article
| Open Access CamoTSS: analysis of alternative transcription start sites for cellular phenotypes and regulatory patterns from 5' scRNA-seq data
CamoTSS: analysis of alternative transcription start sites for cellular phenotypes and regulatory patterns from 5' scRNA-seq data
Five-prime single-cell RNA-seq, especially the read 1, has precise capture of transcription start sites (TSS), but such information is often overlooked. Here, authors present a computational method suite, CamoTSS, to precisely identify TSS and quantify its expression, enabling effective detection of alternative TSS usage in different biological processes.
- Ruiyan Hou
- , Chung-Chau Hon
- & Yuanhua Huang
-
Article
| Open Access trRosettaRNA: automated prediction of RNA 3D structure with transformer network
trRosettaRNA: automated prediction of RNA 3D structure with transformer network
Here, authors develop trRosettaRNA, a deep learning-based approach for predicting RNA 3D structures. Blind tests demonstrate that the automated predictions compete effectively with top human predictions on natural RNAs.
- Wenkai Wang
- , Chenjie Feng
- & Jianyi Yang
-
Article
| Open Access Speos: an ensemble graph representation learning framework to predict core gene candidates for complex diseases
Speos: an ensemble graph representation learning framework to predict core gene candidates for complex diseases
Understanding phenotype-genotype relationships is a grand challenge of current biological research. Here, the authors use graph representation learning to identify human genes which display key characteristics of core genes for five complex diseases.
- Florin Ratajczak
- , Mitchell Joblin
- & Matthias Heinig
-
Article
| Open Access Evolutionary design of explainable algorithms for biomedical image segmentation
Evolutionary design of explainable algorithms for biomedical image segmentation
Deep learning frameworks require large human-annotated datasets for training and the resulting ‘black box’ models are difficult to interpret. Here, the authors present Kartezio; a modular Cartesian Genetic Programming-based computational strategy that generates fully transparent and easily interpretable image processing pipelines.
- Kévin Cortacero
- , Brienne McKenzie
- & Sylvain Cussat-Blanc
-
Article
| Open Access LensAge index as a deep learning-based biological age for self-monitoring the risks of age-related diseases and mortality
LensAge index as a deep learning-based biological age for self-monitoring the risks of age-related diseases and mortality
Age is closely related to health, but chronologically defined age often disagrees with biological age. Here, the authors develop an indicator of biological age - LensAge index - to reveal individuals’ aging level, and it can be implemented with smartphones, showing potential for self-monitoring of aging.
- Ruiyang Li
- , Wenben Chen
- & Haotian Lin
-
Article
| Open Access Fetal biometry and amniotic fluid volume assessment end-to-end automation using Deep Learning
Fetal biometry and amniotic fluid volume assessment end-to-end automation using Deep Learning
Fetal biometry and amniotic fluid volume are essential but strenuous measurements in fetal ultrasound screening. Here, the authors show that deep learning models can automate these measurements with high accuracy, using a large and diverse dataset of Moroccan fetal ultrasound images.
- Saad Slimani
- , Salaheddine Hounka
- & El Houssine Bouyakhf
-
Article
| Open Access Proteomics reveal biomarkers for diagnosis, disease activity and long-term disability outcomes in multiple sclerosis
Proteomics reveal biomarkers for diagnosis, disease activity and long-term disability outcomes in multiple sclerosis
Precise biomarkers for multiple sclerosis prognosis are vital for treatment decisions. Here, the authors identify specific proteins in cerebrospinal fluid that can predict short-term disease activity and long-term disability outcomes in persons with multiple sclerosis.
- Julia Åkesson
- , Sara Hojjati
- & Mika Gustafsson
-
Article
| Open Access A systematic study of key elements underlying molecular property prediction
A systematic study of key elements underlying molecular property prediction
AI has become a crucial tool for drug discovery, but how to properly represent molecules for data-driven property prediction is still an open question. Here the authors evaluate 62,820 models to highlight existing challenges, the impact of activity cliffs, and the crucial role of dataset size.
- Jianyuan Deng
- , Zhibo Yang
- & Fusheng Wang
-
Article
| Open Access Neuropathologist-level integrated classification of adult-type diffuse gliomas using deep learning from whole-slide pathological images
Neuropathologist-level integrated classification of adult-type diffuse gliomas using deep learning from whole-slide pathological images
Determining glioma types directly from whole-slide images (WSIs) would be of great diagnostic utility. Here, the authors develop a deep learning model to identify diffuse glioma types from WSIs according to the 2021 WHO classification across multiple cohorts and with interpretable features.
- Weiwei Wang
- , Yuanshen Zhao
- & Zhenyu Zhang
-
Article
| Open Access Deep flanking sequence engineering for efficient promoter design using DeepSEED
Deep flanking sequence engineering for efficient promoter design using DeepSEED
Designing promoters with desired properties is crucial in synthetic biology. Here, authors introduce DeepSEED, an AI-aided flanking sequence optimisation framework which combines expert knowledge with deep learning techniques to efficiently design promoters in both eukaryotic and prokaryotic cells.
- Pengcheng Zhang
- , Haochen Wang
- & Xiaowo Wang
-
Article
| Open Access A pharmacophore-guided deep learning approach for bioactive molecular generation
A pharmacophore-guided deep learning approach for bioactive molecular generation
Designing novel molecules with desired bioactivity is a critical challenge in drug discovery, particularly for novel or understudied targets. The authors propose a pharmacophore-guided deep learning approach PGMG to generate diverse active-like molecules with limited activity data.
- Huimin Zhu
- , Renyi Zhou
- & Min Li
-
Article
| Open Access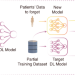 A unified method to revoke the private data of patients in intelligent healthcare with audit to forget
A unified method to revoke the private data of patients in intelligent healthcare with audit to forget
Revoking personal private data is one of the basic human rights. Here, the authors show AFS, a unified method to revoke patients’ private data from pre-trained deep learning models.
- Juexiao Zhou
- , Haoyang Li
- & Xin Gao
-
Article
| Open Access scBridge embraces cell heterogeneity in single-cell RNA-seq and ATAC-seq data integration
scBridge embraces cell heterogeneity in single-cell RNA-seq and ATAC-seq data integration
Multi-omics data integration can be challenging in the event of cell heterogeneity. Here, the authors present scBridge, a method that exploits heterogeneous omics differences, to progressively integrate cells and narrows omics gap, leading to promising integration and label transfer results.
- Yunfan Li
- , Dan Zhang
- & Xi Peng
-
Article
| Open Access Large-scale capture of hidden fluorescent labels for training generalizable markerless motion capture models
Large-scale capture of hidden fluorescent labels for training generalizable markerless motion capture models
Deep learning-based models for tracking behavior are often constrained by manual annotation. Here, authors present GlowTrack, an approach using fluorescence to generate large and diverse training sets that improve model robustness and tracking coverage.
- Daniel J. Butler
- , Alexander P. Keim
- & Eiman Azim
-
Article
| Open Access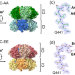 Determining subunit-subunit interaction from statistics of cryo-EM images: observation of nearest-neighbor coupling in a circadian clock protein complex
Determining subunit-subunit interaction from statistics of cryo-EM images: observation of nearest-neighbor coupling in a circadian clock protein complex
Deciphering interactions between subunits in protein complexes is an important problem. By combining cryo-EM imaging and statistical modeling, Han and colleagues reveal a significant cooperativity between subunits in the clock protein hexamer KaiC.
- Xu Han
- , Dongliang Zhang
- & Qi Ouyang
-
Article
| Open Access DeepSlice: rapid fully automatic registration of mouse brain imaging to a volumetric atlas
DeepSlice: rapid fully automatic registration of mouse brain imaging to a volumetric atlas
Navigating the complex structure of the brain poses a challenge to neuroscientists. Here, the authors have trained an AI (DeepSlice) that can automatically register brain images with speed and accuracy, thus simplifying this process.
- Harry Carey
- , Michael Pegios
- & Simon McMullan
-
Article
| Open Access Translating genomic tools to Raman spectroscopy analysis enables high-dimensional tissue characterization on molecular resolution
Translating genomic tools to Raman spectroscopy analysis enables high-dimensional tissue characterization on molecular resolution
Spatial transcriptomics of histological sections have revolutionized basic research, while the actual biomolecular composition of the sample has fallen behind. Here, the authors propose a novel approach to analyze untargeted spatiomolecular Raman spectroscopy data through bioinformatic tools developed for transcriptomic analyses, and integrate them with additional Omics techniques.
- Manuel Sigle
- , Anne-Katrin Rohlfing
- & Meinrad Paul Gawaz
-
Article
| Open Access Integrating end-to-end learning with deep geometrical potentials for ab initio RNA structure prediction
Integrating end-to-end learning with deep geometrical potentials for ab initio RNA structure prediction
Here the authors developed an open-source program (DRfold) for RNA tertiary structure prediction from sequence. Through a unique combination of end-to-end learning and geometry restraint guided simulations, the method demonstrates advantage over peer methods.
- Yang Li
- , Chengxin Zhang
- & Yang Zhang
-
Article
| Open Access Machine learning coarse-grained potentials of protein thermodynamics
Machine learning coarse-grained potentials of protein thermodynamics
Understanding protein dynamics is a complex scientific challenge. Here, authors construct coarse-grained molecular potentials using artificial neural networks, significantly accelerating protein dynamics simulations while preserving their thermodynamics.
- Maciej Majewski
- , Adrià Pérez
- & Gianni De Fabritiis
-
Article
| Open Access DNA methylation profiling to determine the primary sites of metastatic cancers using formalin-fixed paraffin-embedded tissues
DNA methylation profiling to determine the primary sites of metastatic cancers using formalin-fixed paraffin-embedded tissues
Molecular tests that can determine the tissue of origin of cancers of unknown primary (CUP) are still needed. Here, the authors develop a DNA methylation profiling assay and a machine learning classifier to predict the origin of metastatic tumours in CUP patients using formalin-fixed, paraffin embedded samples.
- Shirong Zhang
- , Shutao He
- & Hongcang Gu
-
Article
| Open Access Transfer Learning with Kernel Methods
Transfer Learning with Kernel Methods
Transfer learning can be applied in computer vision and natural language processing to utilize knowledge from a source task to improve performance on a target task. The authors propose a framework for transfer learning with kernel methods for improved image classification and virtual drug screening.
- Adityanarayanan Radhakrishnan
- , Max Ruiz Luyten
- & Caroline Uhler
-
Article
| Open Access Mining multi-center heterogeneous medical data with distributed synthetic learning
Mining multi-center heterogeneous medical data with distributed synthetic learning
Here the authors present Distributed Synthetic Learning, a system that addresses data privacy, isolated data islands, and heterogeneity concerns in healthcare analytics by learning to generate state-of-the-art synthetic data for downstream tasks.
- Qi Chang
- , Zhennan Yan
- & Dimitris N. Metaxas
-
Article
| Open Access Prediction of base editor off-targets by deep learning
Prediction of base editor off-targets by deep learning
Base editors can induce unwanted off-target effects. Here the authors design libraries of gRNA-off-target pairs and perform a screen to obtain editing efficiencies for ABE and CBE: they use the datasets to train DL models (ABEdeepoff and CBEdeepoff) which can predict mutation tolerance at potential off-targets.
- Chengdong Zhang
- , Yuan Yang
- & Yongming Wang
-
Article
| Open Access Interpreting biologically informed neural networks for enhanced proteomic biomarker discovery and pathway analysis
Interpreting biologically informed neural networks for enhanced proteomic biomarker discovery and pathway analysis
Deep neural networks hold significant promise in capturing the complexity of biological systems. However, they suffer from a lack of interpretability. Here, authors present a generalizable method for developing, interpreting, and visualizing biologically informed neural networks for proteomics data.
- Erik Hartman
- , Aaron M. Scott
- & Johan Malmström
-
Article
| Open Access Projecting RNA measurements onto single cell atlases to extract cell type-specific expression profiles using scProjection
Projecting RNA measurements onto single cell atlases to extract cell type-specific expression profiles using scProjection
Many expression deconvolution approaches have been developed to estimate % RNA contributions of diverse cell types to mixed RNA measurements. Here, the authors have developed a complementary approach called scProjection to recover cell type-specific expression profiles from mixed RNA measurements.
- Nelson Johansen
- , Hongru Hu
- & Gerald Quon
-
Article
| Open Access DECIMER.ai: an open platform for automated optical chemical structure identification, segmentation and recognition in scientific publications
DECIMER.ai: an open platform for automated optical chemical structure identification, segmentation and recognition in scientific publications
Chemical structures are typically published as nonmachine-readable images in scientific literature. Here, the authors present DECIMER.ai, an open platform for translating chemical structures in publications into machine-readable representations.
- Kohulan Rajan
- , Henning Otto Brinkhaus
- & Christoph Steinbeck
-
Article
| Open Access APOGEE 2: multi-layer machine-learning model for the interpretable prediction of mitochondrial missense variants
APOGEE 2: multi-layer machine-learning model for the interpretable prediction of mitochondrial missense variants
APOGEE 2 is a machine-learning tool for assessing the fragility of the mitochondrial genome, evaluating genetic variant pathogenicity and ultimately enhancing our understanding of the clinical heterogeneity of mitochondrial genetic diseases.
- Salvatore Daniele Bianco
- , Luca Parca
- & Tommaso Mazza
-
Article
| Open Access A deep learning method for replicate-based analysis of chromosome conformation contacts using Siamese neural networks
A deep learning method for replicate-based analysis of chromosome conformation contacts using Siamese neural networks
Siamese neural networks are a powerful deep learning approach for image analysis. Here, the authors adapt this method to the replicate-based analysis of Hi-C data and find that it successfully discriminates technical noise from biological variation.
- Ediem Al-jibury
- , James W. D. King
- & Daniel Rueckert
-
Article
| Open Access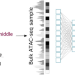 Whole genome deconvolution unveils Alzheimer’s resilient epigenetic signature
Whole genome deconvolution unveils Alzheimer’s resilient epigenetic signature
The authors present a deep learning method that deconvolutes ATAC-seq samples into cell type-specific chromatin accessibility profiles. Applied on 191 samples, the method unveils cell type-specific pathways and nominates potential epigenetic mediators underlying resilience to Alzheimer’s disease.
- Eloise Berson
- , Anjali Sreenivas
- & Thomas J. Montine
-
Article
| Open Access Deep transfer learning for inter-chain contact predictions of transmembrane protein complexes
Deep transfer learning for inter-chain contact predictions of transmembrane protein complexes
Membrane proteins are encoded by approximately a quarter of human genes. Here, the authors propose a deep transfer learning method for predicting inter-chain residue-residue contacts of transmembrane protein complexes.
- Peicong Lin
- , Yumeng Yan
- & Sheng-You Huang
-
Article
| Open Access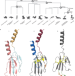 Domain loss enabled evolution of novel functions in the snake three-finger toxin gene superfamily
Domain loss enabled evolution of novel functions in the snake three-finger toxin gene superfamily
3-finger toxins are unique to the venoms of caenophidian snakes. This study traces the evolution of these toxins in snakes, highlighting a key shift from membrane-bound to secretory proteins. This transformation, involving the loss of a membrane-anchoring domain and changes in gene expression, paved the way for their venomous function.
- Ivan Koludarov
- , Tobias Senoner
- & Burkhard Rost
-
Article
| Open Access A machine-learning approach to human ex vivo lung perfusion predicts transplantation outcomes and promotes organ utilization
A machine-learning approach to human ex vivo lung perfusion predicts transplantation outcomes and promotes organ utilization
Ex vivo perfusion is a unique platform to study isolated human lungs. Here, authors show that a machine learning model, InsighTx, derived from data generated during ex vivo lung perfusion can accurately predict transplant outcomes and increase organ utilization rates.
- Andrew T. Sage
- , Laura L. Donahoe
- & Shaf Keshavjee
-
Article
| Open Access Experimental validation of the free-energy principle with in vitro neural networks
Experimental validation of the free-energy principle with in vitro neural networks
Empirical applications of the free-energy principle entail a commitment to a particular process theory. Here, the authors reverse engineered generative models from neural responses of in vitro networks and demonstrated that the free-energy principle could predict how neural networks reorganized in response to external stimulation.
- Takuya Isomura
- , Kiyoshi Kotani
- & Karl J. Friston
-
Article
| Open Access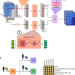 Getting personal with epigenetics: towards individual-specific epigenomic imputation with machine learning
Getting personal with epigenetics: towards individual-specific epigenomic imputation with machine learning
The authors present eDICE, an attention-based model that enables accurate imputation of missing portions of the observed epigenetic landscape, and show that eDICE can be used to predict individualspecific epigenomic variation in the EN-TEx dataset.
- Alex Hawkins-Hooker
- , Giovanni Visonà
- & Gabriele Schweikert
-
Article
| Open Access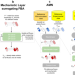 A neural-mechanistic hybrid approach improving the predictive power of genome-scale metabolic models
A neural-mechanistic hybrid approach improving the predictive power of genome-scale metabolic models
Mechanistic models estimate the phenotype of microorganisms in different environments but may have limited predictive capabilities. Here, authors develop trainable hybrid models with improved predictability using mechanistic insights and smaller training sets than conventional machine learning techniques.
- Léon Faure
- , Bastien Mollet
- & Jean-Loup Faulon
-
Article
| Open Access Segmenting functional tissue units across human organs using community-driven development of generalizable machine learning algorithms
Segmenting functional tissue units across human organs using community-driven development of generalizable machine learning algorithms
Constructing the human reference atlas requires integration and analysis of massive amounts of data. Here the authors report the setup and results of the Hacking the Human Body machine learning algorithm development competition hosted by the Human Biomolecular Atlas and the Human Protein Atlas teams.
- Yashvardhan Jain
- , Leah L. Godwin
- & Katy Börner
-
Article
| Open Access Identification of transcriptional programs using dense vector representations defined by mutual information with GeneVector
Identification of transcriptional programs using dense vector representations defined by mutual information with GeneVector
In single-cell RNA-seq analyses, it would be critical to measure the relationships between genes. Here, the authors develop a framework for single-cell dimensionality reduction that incorporates gene-specific relationships - GeneVector -, and use it for tasks such as annotating cell types and analysing pathway variation after treatment.
- Nicholas Ceglia
- , Zachary Sethna
- & Andrew McPherson
-
Article
| Open Access Detecting shortcut learning for fair medical AI using shortcut testing
Detecting shortcut learning for fair medical AI using shortcut testing
Diagnosing shortcut learning in clinical models is difficult, as sensitive attributes may be causally linked with disease. Using multitask learning, the authors propose a method to directly test for the presence of shortcut learning in clinical ML systems.
- Alexander Brown
- , Nenad Tomasev
- & Jessica Schrouff
-
Article
| Open Access Next generation pan-cancer blood proteome profiling using proximity extension assay
Next generation pan-cancer blood proteome profiling using proximity extension assay
Comprehensive and scalable proteomic profiling of plasma samples can improve the screening and diagnosis of cancer patients. Here, the authors use the Olink Proximity Extension Assay technology to characterise the plasma proteomes of 1477 patients across twelve cancer types, and use machine learning to obtain a protein panel for cancer classification.
- María Bueno Álvez
- , Fredrik Edfors
- & Mathias Uhlén
-
Article
| Open Access Deep structured learning for variant prioritization in Mendelian diseases
Deep structured learning for variant prioritization in Mendelian diseases
In individuals with rare, monogenic disorders it often remains challenging to identify the disease-causing genetic variants among numerous potential candidates. Here, the authors develop a neural network ensemble for variant pathogenicity prediction, specifically for this type of disorder.
- Matt C. Danzi
- , Maike F. Dohrn
- & Stephan Züchner
-
Article
| Open Access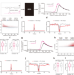 Exon-intron boundary inhibits m6A deposition, enabling m6A distribution hallmark, longer mRNA half-life and flexible protein coding
Exon-intron boundary inhibits m6A deposition, enabling m6A distribution hallmark, longer mRNA half-life and flexible protein coding
m6A mRNA modification is not typically found near splice junctions in mRNAs. Here the authors show exon-intron boundary inhibits m6A deposition at ~100 nt region nearby splice site, enabling m6A distribution hallmark, more stable mRNA and flexible protein coding.
- Zhiyuan Luo
- , Qilian Ma
- & Shengdong Ke
-
Article
| Open Access Discovering functionally important sites in proteins
Discovering functionally important sites in proteins
An important step in understanding and using proteins is to identify the residues that are important for function. The authors present a machine-learning based method to predict functional sites that leverages and combines the information available in protein sequences and structures.
- Matteo Cagiada
- , Sandro Bottaro
- & Kresten Lindorff-Larsen
-
Article
| Open Access Multi-batch single-cell comparative atlas construction by deep learning disentanglement
Multi-batch single-cell comparative atlas construction by deep learning disentanglement
Comparing single-cell RNA-seq and ATAC-seq data from multiple batches is challenging due to technical artifacts. Here, the authors propose a method that disentangles technical and biological effects, facilitating batch-confounded chromatin and gene expression state discovery and enhancing the analysis of perturbation effects on cell populations.
- Allen W. Lynch
- , Myles Brown
- & Clifford A. Meyer
-
Article
| Open Access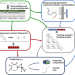 Turnover number predictions for kinetically uncharacterized enzymes using machine and deep learning
Turnover number predictions for kinetically uncharacterized enzymes using machine and deep learning
The turnover numbers of most enzyme-catalyzed reactions are unknown. Kroll et al. developed a general model that can predict turnover numbers even for enzymes dissimilar to those used for training, outperforming existing models.
- Alexander Kroll
- , Yvan Rousset
- & Martin J. Lercher
-
Article
| Open Access Spatial cellular architecture predicts prognosis in glioblastoma
Spatial cellular architecture predicts prognosis in glioblastoma
Intra-tumoral heterogeneity and cell-state plasticity contribute to the development of therapeutic resistance in glioblastoma (GBM). Here the authors use two deep learning models to predict spatial transcriptional programs and prognosis from histology images in GBM.
- Yuanning Zheng
- , Francisco Carrillo-Perez
- & Olivier Gevaert
-
Article
| Open Access Large depth-of-field ultra-compact microscope by progressive optimization and deep learning
Large depth-of-field ultra-compact microscope by progressive optimization and deep learning
Traditional optical microscope, while bulky, often fails to deliver optimal performance. Here, the authors have engineered an integrated microscope of 0.15 cm3 in volume and a weight of 0.5 g, which outperforms a commercial microscope and can be seamlessly integrated with a smartphone.
- Yuanlong Zhang
- , Xiaofei Song
- & Qionghai Dai
-
Article
| Open Access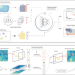 Leveraging spatial transcriptomics data to recover cell locations in single-cell RNA-seq with CeLEry
Leveraging spatial transcriptomics data to recover cell locations in single-cell RNA-seq with CeLEry
Cell location information is important for understanding how tissue is spatially organized. Here, the authors develop CeLEry, a machine learning method that aims to recover cell locations for single-cell RNA-seq data by leveraging information learned from spatial transcriptomics.
- Qihuang Zhang
- , Shunzhou Jiang
- & Mingyao Li

 CLOOME: contrastive learning unlocks bioimaging databases for queries with chemical structures
CLOOME: contrastive learning unlocks bioimaging databases for queries with chemical structures