Featured
-
-
Article
| Open Access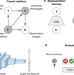 Learning representations for image-based profiling of perturbations
Learning representations for image-based profiling of perturbations
Assessing cell phenotypes in image-based assays requires solid computational methods for transforming images into quantitative data. Here, the authors present a strategy for learning representations of treatment effects from high-throughput imaging, following a causal interpretation.
- Nikita Moshkov
- , Michael Bornholdt
- & Juan C. Caicedo
-
Article
| Open Access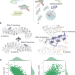 Design of target specific peptide inhibitors using generative deep learning and molecular dynamics simulations
Design of target specific peptide inhibitors using generative deep learning and molecular dynamics simulations
Here the authors report a computational approach which integrates deep learning and structural modelling to design target-specific peptides. They apply this to β-catenin and NF-κB essential modulator, resulting in improved binding, highlighting the efficacy of this strategy.
- Sijie Chen
- , Tong Lin
- & Xiaolin Cheng
-
Article
| Open Access Large language models streamline automated machine learning for clinical studies
Large language models streamline automated machine learning for clinical studies
A knowledge gap persists between machine learning developers and clinicians. Here, the authors show that the Advanced Data Analysis extension of ChatGPT could bridge this gap and simplify complex data analyses, making them more accessible to clinicians.
- Soroosh Tayebi Arasteh
- , Tianyu Han
- & Sven Nebelung
-
Article
| Open Access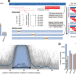 Efficient encoding of large antigenic spaces by epitope prioritization with Dolphyn
Efficient encoding of large antigenic spaces by epitope prioritization with Dolphyn
Profiling antibody responses to vast antigenic spaces has been challenging using programmable phage display (PhIP-Seq). Here, authors develop a methodology for compressing large proteomic spaces and have discovered human antibodies targeting gut bacteria-infecting phages.
- Anna-Maria Liebhoff
- , Thiagarajan Venkataraman
- & H. Benjamin Larman
-
Article
| Open Access Machine learning-based extrachromosomal DNA identification in large-scale cohorts reveals its clinical implications in cancer
Machine learning-based extrachromosomal DNA identification in large-scale cohorts reveals its clinical implications in cancer
‘Extrachromosomal DNA has been previously linked to tumour progression and heterogeneity, but its potential as a cancer biomarker has not been fully explored. Here, the authors develop a computational framework to refine genomic subtypes and predict response to immunotherapy in gastrointestinal cancer.
- Shixiang Wang
- , Chen-Yi Wu
- & Qi Zhao
-
Article
| Open Access A signal processing and deep learning framework for methylation detection using Oxford Nanopore sequencing
A signal processing and deep learning framework for methylation detection using Oxford Nanopore sequencing
The authors present DeepMod2, a deep-learning based computational method that allows fast and accurate detection of DNA methylation and epihaplotypes from Oxford Nanopore sequencing data.
- Mian Umair Ahsan
- , Anagha Gouru
- & Kai Wang
-
Article
| Open Access A deep-learning-based framework for identifying and localizing multiple abnormalities and assessing cardiomegaly in chest X-ray
A deep-learning-based framework for identifying and localizing multiple abnormalities and assessing cardiomegaly in chest X-ray
Accurate localization of abnormalities is crucial in the interpretation of chest X-rays. Here the authors present a deep learning framework for simultaneous localization of 14 thoracic abnormalities and calculation of cardiothoracic ratio, based on large X-ray dataset with bounding boxes created via a human-in-the-loop approach.
- Weijie Fan
- , Yi Yang
- & Dong Zhang
-
Article
| Open Access Regression-based Deep-Learning predicts molecular biomarkers from pathology slides
Regression-based Deep-Learning predicts molecular biomarkers from pathology slides
Cancer biomarkers are often continuous measurements, which poses challenges for their prediction using classification-based deep learning. Here, the authors develop a regression-based deep learning method to predict continuous biomarkers - such as the homologous repair deficiency score - from cancer histopathology images.
- Omar S. M. El Nahhas
- , Chiara M. L. Loeffler
- & Jakob Nikolas Kather
-
Article
| Open Access Predicting DNA structure using a deep learning method
Predicting DNA structure using a deep learning method
In this work, the authors report a deep learning method, Deep DNAshape, to predict the influence of flanking regions on three-dimensional DNA structure and in structural readout mechanisms of protein-DNA binding.
- Jinsen Li
- , Tsu-Pei Chiu
- & Remo Rohs
-
Article
| Open Access A multicenter clinical AI system study for detection and diagnosis of focal liver lesions
A multicenter clinical AI system study for detection and diagnosis of focal liver lesions
Early detection and accurate diagnosis of focal liver lesions are crucial for effective treatment and prognosis. Here, the authors present a fully automated diagnostic system that leverages multi-phase CT scans and clinical features, for diagnosing liver lesions.
- Hanning Ying
- , Xiaoqing Liu
- & Xiujun Cai
-
Article
| Open Access MAIVeSS: streamlined selection of antigenically matched, high-yield viruses for seasonal influenza vaccine production
MAIVeSS: streamlined selection of antigenically matched, high-yield viruses for seasonal influenza vaccine production
Vaccines combat global influenza threats, relying on timely selection of optimal seed viruses. Here, authors introduce MAIVeSS, a machine learning assisted framework to streamline vaccine seed virus selection using genomic sequence, expediting seasonal flu vaccine production and supply.
- Cheng Gao
- , Feng Wen
- & Xiu-Feng Wan
-
Article
| Open Access Orientation-invariant autoencoders learn robust representations for shape profiling of cells and organelles
Orientation-invariant autoencoders learn robust representations for shape profiling of cells and organelles
In image analysis, the shape properties of cells/organelles should be unaffected by image orientation. Conventional autoencoder (AE) methods can be sensitive to orientation. Here, the authors develop an unsupervised AE method that learns robust, orientation-invariant representations.
- James Burgess
- , Jeffrey J. Nirschl
- & Serena Yeung-Levy
-
Article
| Open Access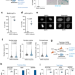 Detection of senescence using machine learning algorithms based on nuclear features
Detection of senescence using machine learning algorithms based on nuclear features
Identifying senescence is complicated by a lack of universal markers. Here, Duran et al. use nuclear morphology features to devise machine-learning classifiers that detect senescence in cell lines and liver sections of patients and mouse models of aging and disease.
- Imanol Duran
- , Joaquim Pombo
- & Jesús Gil
-
Article
| Open Access The impacts of active and self-supervised learning on efficient annotation of single-cell expression data
The impacts of active and self-supervised learning on efficient annotation of single-cell expression data
Cell type annotation for single-cell data is challenging. Here, authors explore active and self-supervised learning and introduce adaptive reweighting as a tailored heuristic, demonstrating competitive performance and showing that incorporating prior knowledge enhances cell type annotation accuracy.
- Michael J. Geuenich
- , Dae-won Gong
- & Kieran R. Campbell
-
Article
| Open Access SiFT: uncovering hidden biological processes by probabilistic filtering of single-cell data
SiFT: uncovering hidden biological processes by probabilistic filtering of single-cell data
Cells simultaneously encode multiple signals, some harder to recover. Here, authors introduce SiFT (Signal FilTering), a kernel-based projection method, revealing underlying biological processes in single-cell data.
- Zoe Piran
- & Mor Nitzan
-
Article
| Open Access Segment anything in medical images
Segment anything in medical images
Segmentation is an important fundamental task in medical image analysis. Here the authors show a deep learning model for efficient and accurate segmentation across a wide range of medical image modalities and anatomies.
- Jun Ma
- , Yuting He
- & Bo Wang
-
Article
| Open Access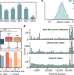 Using big sequencing data to identify chronic SARS-Coronavirus-2 infections
Using big sequencing data to identify chronic SARS-Coronavirus-2 infections
Chronic SARS-CoV-2 infections have been hypothesised to be sources of new variants. Here, the authors use large-scale genome sequencing data to identify mutations predictive of chronic infections, which may therefore be relevant in future variants.
- Sheri Harari
- , Danielle Miller
- & Adi Stern
-
Article
| Open Access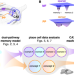 Distinguishing examples while building concepts in hippocampal and artificial networks
Distinguishing examples while building concepts in hippocampal and artificial networks
While the hippocampus is well-known to store specific memories, it can also learn common features that are shared across individual memories. Here, the authors show how this ability arises from dual input pathways and how it can inspire better machine learning methods.
- Louis Kang
- & Taro Toyoizumi
-
Article
| Open Access Clinical application of tumour-in-normal contamination assessment from whole genome sequencing
Clinical application of tumour-in-normal contamination assessment from whole genome sequencing
Assessing tumour contamination in normal samples is critical for accurate variant calling in cancer samples. Here, the authors develop TINC, a computational method to determine the level of tumour in normal contamination, and demonstrate its application in the Genomics England 100,000 Genomes Project dataset.
- Jonathan Mitchell
- , Salvatore Milite
- & Giulio Caravagna
-
Article
| Open Access PROST: quantitative identification of spatially variable genes and domain detection in spatial transcriptomics
PROST: quantitative identification of spatially variable genes and domain detection in spatial transcriptomics
Understanding biological mechanisms requires a thorough exploration of spatiotemporal transcriptional patterns in complex tissues. Here, authors present PROST to quantify spatial gene expression patterns and detect spatial domains using spatial transcriptomics data of varying resolutions.
- Yuchen Liang
- , Guowei Shi
- & Zhonghui Tang
-
Article
| Open Access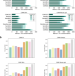 Effective binning of metagenomic contigs using contrastive multi-view representation learning
Effective binning of metagenomic contigs using contrastive multi-view representation learning
Here, the authors present COMEBin, a metagenomics binning method based on contrastive multi-view representation learning that uses data augmentation to generate multiple fragments (views) of each contig, resulting in high-quality embeddings of heterogeneous features. COMEBin outperforms state-of-the art binning methods, particularly in recovering near-complete genomes from real environmental samples.
- Ziye Wang
- , Ronghui You
- & Shanfeng Zhu
-
Article
| Open Access BIDCell: Biologically-informed self-supervised learning for segmentation of subcellular spatial transcriptomics data
BIDCell: Biologically-informed self-supervised learning for segmentation of subcellular spatial transcriptomics data
Subcellular in situ spatial transcriptomics offers the promise to address biological problems that were previously inaccessible but requires accurate cell segmentation to uncover insights. Here, authors present BIDCell, a biologically informed, deep learning-based cell segmentation framework.
- Xiaohang Fu
- , Yingxin Lin
- & Jean Y. H. Yang
-
Article
| Open Access Gene-SGAN: discovering disease subtypes with imaging and genetic signatures via multi-view weakly-supervised deep clustering
Gene-SGAN: discovering disease subtypes with imaging and genetic signatures via multi-view weakly-supervised deep clustering
Many diseases can display distinct brain imaging phenotypes across individuals, potentially reflecting disease subtypes. However, biological interpretability is limited if the derived subtypes are not associated with genetic drivers or susceptibility factors. Here, the authors describe a deep-learning method that links imaging phenotypes with genetic factors, thereby conferring genetic correlations to the disease subtypes.
- Zhijian Yang
- , Junhao Wen
- & Christos Davatzikos
-
Article
| Open Access MarsGT: Multi-omics analysis for rare population inference using single-cell graph transformer
MarsGT: Multi-omics analysis for rare population inference using single-cell graph transformer
Identifying rare cell populations is key to understanding cancer progression and response to therapy. Here, authors introduce MarsGT, an end-to-end deep learning model for rare cell population identification from scMulti-omics data.
- Xiaoying Wang
- , Maoteng Duan
- & Qin Ma
-
Article
| Open Access Radiomic tractometry reveals tract-specific imaging biomarkers in white matter
Radiomic tractometry reveals tract-specific imaging biomarkers in white matter
Diffusion MRI is used for tract-specific microstructural analysis of the white matter. Here, the authors introduce radiomic tractometry (RadTract), enhancing tractometry with radiomics-based imaging biomarkers for improved predictive modelling.
- Peter Neher
- , Dusan Hirjak
- & Klaus Maier-Hein
-
Article
| Open Access Improving deep neural network generalization and robustness to background bias via layer-wise relevance propagation optimization
Improving deep neural network generalization and robustness to background bias via layer-wise relevance propagation optimization
Image background features can undesirably affect deep networks’ decisions. Here, the authors show that the optimization of Layer-wise Relevance Propagation explanation heatmaps can hinder such influence, improving out-of-distribution generalization.
- Pedro R. A. S. Bassi
- , Sergio S. J. Dertkigil
- & Andrea Cavalli
-
Article
| Open Access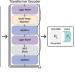 ECOLE: Learning to call copy number variants on whole exome sequencing data
ECOLE: Learning to call copy number variants on whole exome sequencing data
Copy number variants (CNV) are shown to contribute to the etiology of various genetic disorders. Here, authors present ECOLE, a deep learning-based somatic and germline CNV caller for WES data. Utilising a variant of the transformer architecture, the model is trained to call CNVs per exon.
- Berk Mandiracioglu
- , Furkan Ozden
- & A. Ercument Cicek
-
Article
| Open Access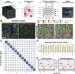 MAPS: pathologist-level cell type annotation from tissue images through machine learning
MAPS: pathologist-level cell type annotation from tissue images through machine learning
Current cell annotation methods using high-plex spatial proteomics data are resource intensive and demand iterative expert input. Here, the authors present MAPS (Machine learning for Analysis of Proteomics in Spatial biology), an approach that facilitates rapid and precise cell type identification with human-level accuracy from spatial proteomics data.
- Muhammad Shaban
- , Yunhao Bai
- & Faisal Mahmood
-
Article
| Open Access Deep learning-driven fragment ion series classification enables highly precise and sensitive de novo peptide sequencing
Deep learning-driven fragment ion series classification enables highly precise and sensitive de novo peptide sequencing
Accurate and high-throughput sequencing methods for proteins are lacking. Here the authors report Spectralis which improves de novo peptide sequencing using a convolutional layer that connects peaks in spectra spaced by amino acid masses, fragment ion series classification and a peptide-spectrum match confidence score.
- Daniela Klaproth-Andrade
- , Johannes Hingerl
- & Julien Gagneur
-
Article
| Open Access Merizo: a rapid and accurate protein domain segmentation method using invariant point attention
Merizo: a rapid and accurate protein domain segmentation method using invariant point attention
Proteins contain modular structural and functional units called domains. Here, the authors have developed Merizo, a deep learning method for domain segmentation applicable to experimental structures as well as those generated by AlphaFold2.
- Andy M. Lau
- , Shaun M. Kandathil
- & David T. Jones
-
Article
| Open Access Deep learning-based phenotyping reclassifies combined hepatocellular-cholangiocarcinoma
Deep learning-based phenotyping reclassifies combined hepatocellular-cholangiocarcinoma
Combined hepatocellular-cholangiocarcinomas (cHCC-CCA) are challenging to diagnose, as they exhibit features of hepatocellular carcinoma (HCC) and intrahepatic cholangiocarcinoma (ICCA). Here, the authors use deep learning to re-classify cHCC-CCA tumours into HCC or ICCA based on histopathology images.
- Julien Calderaro
- , Narmin Ghaffari Laleh
- & Jakob Nikolas Kather
-
Article
| Open Access Accurate prediction of protein assembly structure by combining AlphaFold and symmetrical docking
Accurate prediction of protein assembly structure by combining AlphaFold and symmetrical docking
Current methods to predict structures of proteins cannot handle large assemblies with complex symmetries. Here, the authors demonstrate that structures of proteins with cubic symmetries can be accurately predicted with a method combining AlphaFold with symmetrical assembly simulations.
- Mads Jeppesen
- & Ingemar André
-
Article
| Open Access GNTD: reconstructing spatial transcriptomes with graph-guided neural tensor decomposition informed by spatial and functional relations
GNTD: reconstructing spatial transcriptomes with graph-guided neural tensor decomposition informed by spatial and functional relations
Reconstructing transcriptome-wide spatially-resolved gene expressions requires modelling nonlinear patterns and spatial structures in RNA profiling data. Here, authors introduce a graph-guided neural hierarchical tensor decomposition model that incorporates spatial and functional relations for the task.
- Tianci Song
- , Charles Broadbent
- & Rui Kuang
-
Article
| Open Access DeepRTAlign: toward accurate retention time alignment for large cohort mass spectrometry data analysis
DeepRTAlign: toward accurate retention time alignment for large cohort mass spectrometry data analysis
Retention time (RT) alignment is a crucial step in large cohort proteomics and metabolomics studies. Here, the authors introduce DeepRTAlign, a deep learning tool for RT alignment that shows high identification sensitivity and quantitative accuracy.
- Yi Liu
- , Yun Yang
- & Cheng Chang
-
Article
| Open Access High-throughput target trial emulation for Alzheimer’s disease drug repurposing with real-world data
High-throughput target trial emulation for Alzheimer’s disease drug repurposing with real-world data
Target trial emulation (TTE) simulates randomized controlled trials using real world data (RWD). Here, authors show the effectiveness of different TTE strategies to identify drug candidates that could be potentially repurposed to Alzheimer’s disease using two large scale RWD warehouses.
- Chengxi Zang
- , Hao Zhang
- & Fei Wang
-
Article
| Open Access STalign: Alignment of spatial transcriptomics data using diffeomorphic metric mapping
STalign: Alignment of spatial transcriptomics data using diffeomorphic metric mapping
Spatial transcriptomics (ST) enables gene expression characterisation within tissue sections, but comparing across sections and technologies remains challenging. Here, authors develop STalign to spatially align ST data and demonstrate applications including aligning to common coordinate frameworks.
- Kalen Clifton
- , Manjari Anant
- & Jean Fan
-
Article
| Open Access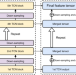 Accurate de novo peptide sequencing using fully convolutional neural networks
Accurate de novo peptide sequencing using fully convolutional neural networks
De novo peptide sequencing allows the identification of peptides without requiring target databases. Here, the authors present PepNet, a convolutional neural network model for accurate de novo peptide sequencing that is capable of analysing large-scale proteomics data.
- Kaiyuan Liu
- , Yuzhen Ye
- & Haixu Tang
-
Article
| Open Access Autoencoder neural networks enable low dimensional structure analyses of microbial growth dynamics
Autoencoder neural networks enable low dimensional structure analyses of microbial growth dynamics
Here, the authors apply autoencoder neural networks to show that microbial growth dynamics can be compressed into low-dimensional representations and reconstructed with high fidelity, facilitating quantitative predictions and deduction of potential mechanisms.
- Yasa Baig
- , Helena R. Ma
- & Lingchong You
-
Article
| Open Access Augmenting interpretable models with large language models during training
Augmenting interpretable models with large language models during training
Prediction and interpretation tasks may be challenging in high-stakes applications, such as medical decision-making, or systems with compute-limited hardware. The authors introduce an augmented framework for leveraging the knowledge learned by Large Language Models to build interpretable models which are both accurate and efficient.
- Chandan Singh
- , Armin Askari
- & Jianfeng Gao
-
Article
| Open Access Integrating spatial and single-cell transcriptomics data using deep generative models with SpatialScope
Integrating spatial and single-cell transcriptomics data using deep generative models with SpatialScope
Spatial transcriptomics (ST) is transforming tissue analysis but has limitations. Here, authors introduce SpatialScope, an integrated approach combining scRNA-seq and ST data using deep generative models, enabling comprehensive spatial characterisation at transcriptome-wide single-cell resolution.
- Xiaomeng Wan
- , Jiashun Xiao
- & Can Yang
-
Article
| Open Access ZeroBind: a protein-specific zero-shot predictor with subgraph matching for drug-target interactions
ZeroBind: a protein-specific zero-shot predictor with subgraph matching for drug-target interactions
Existing drug-target interaction (DTI) prediction methods generally fail to generalize well unseen proteins and drugs. Here the authors report a protein-specific meta-learning framework, ZeroBind, with subgraph matching for predicting protein-drug interactions from their structures.
- Yuxuan Wang
- , Ying Xia
- & Xiaoyong Pan
-
Article
| Open Access PhenoSV: interpretable phenotype-aware model for the prioritization of genes affected by structural variants
PhenoSV: interpretable phenotype-aware model for the prioritization of genes affected by structural variants
Here, authors present PhenoSV, a phenotype-aware machine-learning model for the functional interpretation of various types of structural variants (SVs) and genes within or outside SVs, facilitating the extraction of biological insights from coding and noncoding SVs.
- Zhuoran Xu
- , Quan Li
- & Kai Wang
-
Article
| Open Access Data-driven grading of acute graft-versus-host disease
Data-driven grading of acute graft-versus-host disease
Acute GVHD severity grading is based on target organ assessments. Here, the authors show that data-driven grading can identify 12 distinct grades with specific aGVHD phenotypes, which are associated with clinical outcomes, and that their method outperformed conventional gradings.
- Evren Bayraktar
- , Theresa Graf
- & Amin T. Turki
-
Article
| Open Access Plasma proteomic profiles predict individual future health risk
Plasma proteomic profiles predict individual future health risk
The predictive capability of future health risk using plasma proteomic profiles remains largely unexplored. Using 1461 proteins collected from 50k individuals, authors show proteins can derive much better or equivalent performance than established clinical indicators for more than 40 endpoints.
- Jia You
- , Yu Guo
- & Jin-Tai Yu
-
Article
| Open Access A fully integrated, standalone stretchable device platform with in-sensor adaptive machine learning for rehabilitation
A fully integrated, standalone stretchable device platform with in-sensor adaptive machine learning for rehabilitation
Methods for the wireless, continuous monitoring and analysis of activities directly from the throat skin have not been developed. Here, the authors present a stretchable device platform that provides wireless measurements and machine learning-based analysis of vibrations and muscle electrical activities from the throat.
- Hongcheng Xu
- , Weihao Zheng
- & Libo Gao
-
Article
| Open Access Robust mapping of spatiotemporal trajectories and cell–cell interactions in healthy and diseased tissues
Robust mapping of spatiotemporal trajectories and cell–cell interactions in healthy and diseased tissues
The integration of spatial, imaging, and sequencing information enables the mapping of cellular dynamics within a tissue. Here, authors show three algorithms in stLearn software to accurately reveal spatial trajectory, detect cell-cell interactions, and impute missing data.
- Duy Pham
- , Xiao Tan
- & Quan H. Nguyen
-
Article
| Open Access Paired single-cell multi-omics data integration with Mowgli
Paired single-cell multi-omics data integration with Mowgli
Mowgli is a novel paired single-cell multi-omics integration method leveraging matrix factorization and Optimal Transport. In-depth benchmarking demonstrates promising cell clustering results and improved biological interpretability.
- Geert-Jan Huizing
- , Ina Maria Deutschmann
- & Laura Cantini
-
Article
| Open Access Global pathogenomic analysis identifies known and candidate genetic antimicrobial resistance determinants in twelve species
Global pathogenomic analysis identifies known and candidate genetic antimicrobial resistance determinants in twelve species
A global analysis of antimicrobial resistance (AMR) across 27,155 genomes and 69 drugs reveals patterns in AMR gene transfer between species and identifies 142 AMR gene candidates, two of which were tested and confirmed as contributing to AMR.
- Jason C. Hyun
- , Jonathan M. Monk
- & Bernhard O. Palsson
-
Article
| Open Access ProRefiner: an entropy-based refining strategy for inverse protein folding with global graph attention
ProRefiner: an entropy-based refining strategy for inverse protein folding with global graph attention
Inverse Protein Folding is a critical component of protein design. Here, authors introduce ProRefiner, a deep-learning model for IPF that exhibits both high performance and memory efficiency, thereby contributing to advancements in protein design.
- Xinyi Zhou
- , Guangyong Chen
- & Pheng Ann Heng

 scCASE: accurate and interpretable enhancement for single-cell chromatin accessibility sequencing data
scCASE: accurate and interpretable enhancement for single-cell chromatin accessibility sequencing data