Article
|
Open Access
Featured
-
-
Article |
 Complementary Alu sequences mediate enhancer–promoter selectivity
Complementary Alu sequences mediate enhancer–promoter selectivity
Using RNA in situ conformation sequencing technology, the role of Alu elements in mediating the interaction between enhancers and promoters is shown.
- Liang Liang
- , Changchang Cao
- & Yuanchao Xue
-
Article
| Open Access In vivo single-molecule analysis reveals COOLAIR RNA structural diversity
In vivo single-molecule analysis reveals COOLAIR RNA structural diversity
The structures of single COOLAIR RNA isoforms change in abundance and shape in response to external conditions; structural mutation of these isoforms altered FLC expression and flowering time, consistent with a regulatory role of the COOLAIR structure in FLC transcription.
- Minglei Yang
- , Pan Zhu
- & Yiliang Ding
-
Article |
 Targeting Xist with compounds that disrupt RNA structure and X inactivation
Targeting Xist with compounds that disrupt RNA structure and X inactivation
A molecule identified in a screen for compounds that bind the non-coding mouse RNA Xist blocks Xist-dependent X-chromosome inactivation, demonstrating the utility of this approach for identifying drugs that target RNA.
- Rodrigo Aguilar
- , Kerrie B. Spencer
- & Jeannie T. Lee
-
Article
| Open Access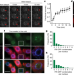 Cold-induced Arabidopsis FRIGIDA nuclear condensates for FLC repression
Cold-induced Arabidopsis FRIGIDA nuclear condensates for FLC repression
In Arabidopsis thaliana, downregulation of the floral repressor FLC in response to cold occurs through a mechanism in which the FLC activator FRIGIDA is sequestered into biomolecular condensates away from the FLC promoter.
- Pan Zhu
- , Clare Lister
- & Caroline Dean
-
Article |
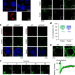 NORAD-induced Pumilio phase separation is required for genome stability
NORAD-induced Pumilio phase separation is required for genome stability
The noncoding RNA NORAD maintains genome stability in mammalian cells by sequestering Pumilio proteins in phase-separated compartments.
- Mahmoud M. Elguindy
- & Joshua T. Mendell
-
Article |
 MIR-NATs repress MAPT translation and aid proteostasis in neurodegeneration
MIR-NATs repress MAPT translation and aid proteostasis in neurodegeneration
The natural antisense transcript MAPT-AS1 interferes with translation of mRNA transcript into tau protein in the brain and may represent a general mechanism for controlling levels of intrinsically disordered proteins, with particular relevance for neurodegeneration.
- Roberto Simone
- , Faiza Javad
- & Rohan de Silva
-
Article |
 Structures of telomerase at several steps of telomere repeat synthesis
Structures of telomerase at several steps of telomere repeat synthesis
Cryo-electron microscopy structures of Tetrahymena telomerase with telomeric DNA at several steps of nucleotide addition provide insights into the structural basis of telomere repeat synthesis.
- Yao He
- , Yaqiang Wang
- & Juli Feigon
-
Article |
 Non-coding deletions identify Maenli lncRNA as a limb-specific En1 regulator
Non-coding deletions identify Maenli lncRNA as a limb-specific En1 regulator
The long non-coding RNA locus Maenli controls mouse limb development by regulating En1 activity, and the absence of the homolgous MAENLI locus is associated with severe congenital limb defects in humans.
- Lila Allou
- , Sara Balzano
- & Andrea Superti-Furga
-
Article |
 RAD51-dependent recruitment of TERRA lncRNA to telomeres through R-loops
RAD51-dependent recruitment of TERRA lncRNA to telomeres through R-loops
Telomeric-repeat-containing RNA is recruited to telomeres by a mechanism that involves the DNA recombinase RAD51 and the formation of DNA–RNA hybrids, or R-loops—a process similar to that involved in homology-directed DNA repair.
- Marianna Feretzaki
- , Michaela Pospisilova
- & Joachim Lingner
-
Article |
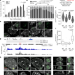 A protein assembly mediates Xist localization and gene silencing
A protein assembly mediates Xist localization and gene silencing
A protein condensate formed by multivalent interactions between the long non-coding RNA Xist and specific RNA-binding proteins drives the compartmentalization required to perpetuate gene silencing on the inactive X chromosome.
- Amy Pandya-Jones
- , Yolanda Markaki
- & Kathrin Plath
-
Article |
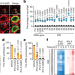 Nucleolar RNA polymerase II drives ribosome biogenesis
Nucleolar RNA polymerase II drives ribosome biogenesis
RNA polymerase II has an unexpected function in the nucleolus, helping to drive the expression of ribosomal RNA and to protect nucleolar structure through a mechanism involving triplex R-loop structures.
- Karan J. Abraham
- , Negin Khosraviani
- & Karim Mekhail
-
Article |
 RIC-seq for global in situ profiling of RNA–RNA spatial interactions
RIC-seq for global in situ profiling of RNA–RNA spatial interactions
RNA in situ conformation sequencing (RIC-seq) enables the generation of three-dimensional interaction maps of RNA in cells, which sheds light on the interactions and regulatory functions of RNA.
- Zhaokui Cai
- , Changchang Cao
- & Yuanchao Xue
-
Article |
 U1 snRNP regulates chromatin retention of noncoding RNAs
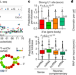
U1 snRNP regulates chromatin retention of noncoding RNAs
Long noncoding RNAs and certain unstable transcripts tend to localize to chromatin, in a process that is shown here to depend on an RNA motif that recognizes the small nuclear ribonuclear protein U1, and to rely on transcription.
- Yafei Yin
- , J. Yuyang Lu
- & Xiaohua Shen
-
Article |
 DNA-PKcs has KU-dependent function in rRNA processing and haematopoiesis
DNA-PKcs has KU-dependent function in rRNA processing and haematopoiesis
The catalytic subunit of DNA-PK autophosphorylates and contributes to ribosome biogenesis and haematopoiesis by binding to the U3 small nucleolar RNA.
- Zhengping Shao
- , Ryan A. Flynn
- & Eliezer Calo
-
Article |
 SPEN integrates transcriptional and epigenetic control of X-inactivation
SPEN integrates transcriptional and epigenetic control of X-inactivation
The transcriptional repressor SPEN bridges the non-coding RNA Xist to transcription machinery, histone deacetylases and chromatin remodelling factors to initiate X-chromosome inactivation.
- François Dossin
- , Inês Pinheiro
- & Edith Heard
-
Article |
 Developmental dynamics of lncRNAs across mammalian organs and species
Developmental dynamics of lncRNAs across mammalian organs and species
A transcriptome dataset from seven organs and seven mammalian species throughout development is used to analyse the expression of long noncoding RNAs in tissues within and between species, and at different stages of organ development.
- Ioannis Sarropoulos
- , Ray Marin
- & Henrik Kaessmann
-
Letter |
 Arabidopsis FLL2 promotes liquid–liquid phase separation of polyadenylation complexes
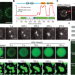
Arabidopsis FLL2 promotes liquid–liquid phase separation of polyadenylation complexes
A genetic screen for factors required by the Arabidopsis RNA-binding protein FCA identifies FLL2 as necessary in the formation of FCA nuclear bodies, and thus a role for FLL2 in liquid–liquid phase separation.
- Xiaofeng Fang
- , Liang Wang
- & Caroline Dean
-
Letter |
 The NORAD lncRNA assembles a topoisomerase complex critical for genome stability
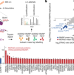
The NORAD lncRNA assembles a topoisomerase complex critical for genome stability
The long non-coding RNA NORAD interacts with proteins involved in DNA replication and repair, and controls the ability of RBMX to form a ribonucleoprotein complex that helps to maintain genomic stability.
- Mathias Munschauer
- , Celina T. Nguyen
- & Eric S. Lander
-
Letter |
 Sequences enriched in Alu repeats drive nuclear localization of long RNAs in human cells
Sequences enriched in Alu repeats drive nuclear localization of long RNAs in human cells
A sequence that is frequently found in Alu elements drives the localization of some long RNAs to the nucleus in human cells.
- Yoav Lubelsky
- & Igor Ulitsky
-
Article |
 An atlas of human long non-coding RNAs with accurate 5′ ends
An atlas of human long non-coding RNAs with accurate 5′ ends
A catalogue of human long non-coding RNA genes and their expression profiles across samples from major human primary cell types, tissues and cell lines.
- Chung-Chau Hon
- , Jordan A. Ramilowski
- & Alistair R. R. Forrest
-
Letter |
 mTORC1 and muscle regeneration are regulated by the LINC00961-encoded SPAR polypeptide
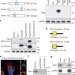
mTORC1 and muscle regeneration are regulated by the LINC00961-encoded SPAR polypeptide
The polypeptide SPAR is encoded by a long non-coding RNA, localizes to the late endosome and lysosome, and regulates muscle regeneration by inhibiting mTORC1.
- Akinobu Matsumoto
- , Alessandra Pasut
- & Pier Paolo Pandolfi
-
Letter |
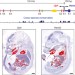 Transcription of the non-coding RNA upperhand controls Hand2 expression and heart development
Transcription of the non-coding RNA upperhand controls Hand2 expression and heart development
Transcription of a long non-coding RNA, known as upperhand (Uph) located upstream of the HAND2 transcription factor is required to maintain transcription of the Hand2 gene by RNA polymerase, and blockade of Uph expression leads to heart defects and embryonic lethality in mice.
- Kelly M. Anderson
- , Douglas M. Anderson
- & Eric N. Olson
-
Letter |
 Local regulation of gene expression by lncRNA promoters, transcription and splicing
Local regulation of gene expression by lncRNA promoters, transcription and splicing
Various cis-regulatory functions of genomic loci that produce long non-coding RNAs are revealed, including instances where their promoters have enhancer-like activity and the lncRNA transcripts themselves are not required for activity.
- Jesse M. Engreitz
- , Jenna E. Haines
- & Eric S. Lander
-
Article |
 m6A RNA methylation promotes XIST-mediated transcriptional repression
m6A RNA methylation promotes XIST-mediated transcriptional repression
The methylation of adenosine residues on the long non-coding RNA XIST is essential for X-chromosome transcriptional repression during female mammalian development.
- Deepak P. Patil
- , Chun-Kan Chen
- & Samie R. Jaffrey
-
Letter |
 Feedback modulation of cholesterol metabolism by the lipid-responsive non-coding RNA LeXis
Feedback modulation of cholesterol metabolism by the lipid-responsive non-coding RNA LeXis
The activation of lipid X receptors (LXRs) in mouse liver not only promotes cholesterol efflux but also inhibits cholesterol synthesis simultaneously; this is mediated by the lipid-responsive long non-coding RNA LeXis, which is induced by a Western diet and orchestrates crosstalk between LXRs and the cholesterol biosynthetic pathway.
- Tamer Sallam
- , Marius C. Jones
- & Peter Tontonoz
-
Letter |
 Melanoma addiction to the long non-coding RNA SAMMSON
Melanoma addiction to the long non-coding RNA SAMMSON
A known oncogene, MITF, resides in a region of chromosome 3 that is amplified in melanomas and associated with poor prognosis; now, a long non-coding RNA gene, SAMMSON, is shown to also lie in this region, to also act as a melanoma-specific survival oncogene, and to be a promising therapeutic target for anti-melanoma therapy.
- Eleonora Leucci
- , Roberto Vendramin
- & Jean-Christophe Marine
-
Letter |
 Integrator mediates the biogenesis of enhancer RNAs
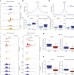
Integrator mediates the biogenesis of enhancer RNAs
This study demonstrates a role for the Integrator complex in the stimulus-dependent induction of eRNAs and their 3′ processing; together with previously known roles of Integrator in transcription elongation and RNA processing, these results indicate that Integrator has broad functions in the regulation of eukaryotic gene expression.
- Fan Lai
- , Alessandro Gardini
- & Ramin Shiekhattar
-
Letter |
 The Xist lncRNA interacts directly with SHARP to silence transcription through HDAC3
The Xist lncRNA interacts directly with SHARP to silence transcription through HDAC3
The mechanisms by which Xist, a long non-coding RNA, silences one X chromosome in female mammals are unknown; here a mass spectrometry-based approach is developed to identify several proteins that interact directly with Xist, including the transcriptional repressor SHARP that is required for transcriptional silencing through the histone deacetylase HDAC3.
- Colleen A. McHugh
- , Chun-Kan Chen
- & Mitchell Guttman
-
Letter |
 A long noncoding RNA protects the heart from pathological hypertrophy
A long noncoding RNA protects the heart from pathological hypertrophy
Here, a long noncoding RNA, termed Mhrt, is identified in the loci of myosin heavy chain (Myh) genes in mice and shown to be capable of suppressing cardiomyopathy in the animals, as well as being repressed in diseased human hearts.
- Pei Han
- , Wei Li
- & Ching-Pin Chang
-
Letter |
 High-resolution Xist binding maps reveal two-step spreading during X-chromosome inactivation
High-resolution Xist binding maps reveal two-step spreading during X-chromosome inactivation
During mammalian X-chromosome inactivation, the Xist long noncoding RNA coats the future inactive X chromosome and recruits polycomb repressive complex 2 to a nucleation site, but how Xist spreads silencing across the entire X chromosome is unclear; here high-resolution maps of Xist binding sites across the X chromosome are generated and show that Xist does not spread across the inactive X chromosome uniformly but in two steps, initially targeting gene-rich islands before later spreading to intervening gene-poor domains.
- Matthew D. Simon
- , Stefan F. Pinter
- & Jeannie T. Lee
-
Article |
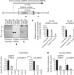 DNMT1-interacting RNAs block gene-specific DNA methylation
DNMT1-interacting RNAs block gene-specific DNA methylation
RNAs are shown to interact with DNA methyltransferase 1 and prevent DNA methylation of genes at their specific locus, providing evidence that active transcription directly regulates DNA methylation levels.
- Annalisa Di Ruscio
- , Alexander K. Ebralidze
- & Daniel G. Tenen
-
Letter |
 lncRNA-dependent mechanisms of androgen-receptor-regulated gene activation programs
lncRNA-dependent mechanisms of androgen-receptor-regulated gene activation programs
A study of prostate cancer cells reveals a transcriptional activation role for long non-coding RNAs (PRNCR1 and PCGEM1) that bind to the androgen receptor, and is also observed for the truncated androgen receptor characteristic of many aggressive prostate cancers.
- Liuqing Yang
- , Chunru Lin
- & Michael G. Rosenfeld
-
Letter |
 Promoter directionality is controlled by U1 snRNP and polyadenylation signals
Promoter directionality is controlled by U1 snRNP and polyadenylation signals
Asymmetric sequence determinants flanking gene transcription start sites are shown to control directionality of transcription elongation in mammalian cells by regulating promoter-proximal cleavage and polyadenylation.
- Albert E. Almada
- , Xuebing Wu
- & Phillip A. Sharp
-
Letter |
 Extensive transcriptional heterogeneity revealed by isoform profiling
Extensive transcriptional heterogeneity revealed by isoform profiling
Variation among RNA transcript isoforms can be generated from alternative start and polyadenylation sites, and results in RNAs and proteins with different properties being generated from the same genomic sequence; here a new method termed transcript isoform sequencing is described in yeast, and the method allows a fuller exploration of transcriptome diversity across the compact yeast genome.
- Vicent Pelechano
- , Wu Wei
- & Lars M. Steinmetz
-
Letter |
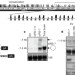 Natural RNA circles function as efficient microRNA sponges
Natural RNA circles function as efficient microRNA sponges
A natural circular RNA termed ciRS-7 is shown to function as a negative regulator of microRNA; ciRS-7 acts as an efficient sponge for the microRNA miR-7, and is resistant to the usual microRNA-mediated degradation pathway of exonucleolytic RNA decay.
- Thomas B. Hansen
- , Trine I. Jensen
- & Jørgen Kjems
-
Letter |
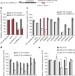 Activating RNAs associate with Mediator to enhance chromatin architecture and transcription
Activating RNAs associate with Mediator to enhance chromatin architecture and transcription
A class of long non-coding RNA (lncRNA) with enhancer-like activity is found to associate with the co-activator complex Mediator and promote its genomic association and enzymatic activity; together with Mediator, the lncRNAs also help to maintain the chromosomal architecture of active regulatory elements.
- Fan Lai
- , Ulf A. Orom
- & Ramin Shiekhattar
-
Letter |
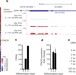 Control of somatic tissue differentiation by the long non-coding RNA TINCR
Control of somatic tissue differentiation by the long non-coding RNA TINCR
The human long non-coding RNA TINCR binds to STAU1 and controls epidermal differentiation by stabilizing key differentiation mRNAs, by means of a TINCR-binding motif found enriched in epidermal differentiation genes.
- Markus Kretz
- , Zurab Siprashvili
- & Paul A. Khavari
-
Letter |
 Long non-coding antisense RNA controls Uchl1 translation through an embedded SINEB2 repeat
Long non-coding antisense RNA controls Uchl1 translation through an embedded SINEB2 repeat
Antisense Uchl1, a long non-coding RNA that is an antisense transcript for the Uchl1 gene, upregulates UCHL1 protein levels through the combined action of an overlapping sequence at its 5′ end and an embedded SINEB2 element.
- Claudia Carrieri
- , Laura Cimatti
- & Stefano Gustincich
-
Research Highlights |
Noncoding RNAs decapped
-
Review Article |
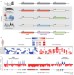 Modular regulatory principles of large non-coding RNAs
Modular regulatory principles of large non-coding RNAs
- Mitchell Guttman
- & John L. Rinn
-
Letter |
 A long noncoding RNA maintains active chromatin to coordinate homeotic gene expression
A long noncoding RNA maintains active chromatin to coordinate homeotic gene expression
Long intergenic non-coding RNAs (lincRNAs) have been implicated in both gene silencing and activation, and could be a means for long-range control of gene expression. Here a lincRNA termed HOTTIP is identified at the 5′ tip of the HOXA locus that coordinates the activation of multiple 5′ HOXA genes. Chromosomal looping brings HOTTIP into the proximity of its target genes, where it seems to be required to facilitate histone H3 lysine 4 trimethylation and gene transcription.
- Kevin C. Wang
- , Yul W. Yang
- & Howard Y. Chang
-
Letter |
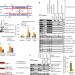 lncRNAs transactivate STAU1-mediated mRNA decay by duplexing with 3′ UTRs via Alu elements
lncRNAs transactivate STAU1-mediated mRNA decay by duplexing with 3′ UTRs via Alu elements
Staufen 1 (STAU1) protein binds regions of dsRNA in the 3′ UTR of mRNAs and promotes their degradation, a process known as SMD (Staufen-mediated mRNA decay). Although a specific stem-loop binding site had been defined for one SMD target, it was unclear how STAU1 was directed to other SMD targets that lack this structure. This paper reports that pairing of Alu element sequences in long non-coding RNAs (lncRNAs) and in the 3′ UTR of the SMD target generates a dsRNA structure that STAU1 recognizes. This result highlights a new function for lncRNAs.
- Chenguang Gong
- & Lynne E. Maquat
-
Research Highlights |
Genetics: Long and the short of it

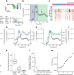 RNA-mediated symmetry breaking enables singular olfactory receptor choice
RNA-mediated symmetry breaking enables singular olfactory receptor choice