Featured
-
-
Article
| Open Access Specific, sensitive and quantitative protein detection by in-gel fluorescence
Specific, sensitive and quantitative protein detection by in-gel fluorescence
Recombinant proteins in complex solutions are typically detected with tag-specific antibodies in Western blots. Here, the author describes an antibody-free alternative in which tagged proteins are detected directly in polyacrylamide gels via fluorophore-labelling of the tagged protein using a ligase.
- Adrian C. D. Fuchs
-
Article
| Open Access The unifying catalytic mechanism of the RING-between-RING E3 ubiquitin ligase family
The unifying catalytic mechanism of the RING-between-RING E3 ubiquitin ligase family
RBR E3 ubiquitin ligases utilise a 2-step catalytic mechanism previously defined for only few of the RBR family members. Here, the authors examine the poorly studied RBRs HOIL-1 and RNF216 to define general principles of RBR catalysis and regulation and identify specific functional differences.
- Xiangyi S. Wang
- , Thomas R. Cotton
- & Bernhard C. Lechtenberg
-
Article
| Open Access A cryptic third active site in cyanophycin synthetase creates primers for polymerization
A cryptic third active site in cyanophycin synthetase creates primers for polymerization
Cyanophycin synthetase CphA1 polymerizes Asp and Arg into the nitrogen reserve polymer cyanophycin using two active sites. Sharon et al. show CphA1 has a cryptic 3rd active site that cleaves cyanophycin into primers for self-sufficient biosynthesis.
- Itai Sharon
- , Sharon Pinus
- & T. Martin Schmeing
-
Article
| Open Access Structures of LIG1 that engage with mutagenic mismatches inserted by polβ in base excision repair
Structures of LIG1 that engage with mutagenic mismatches inserted by polβ in base excision repair
Ligase I seals two ends of DNA during DNA repair and replication. Here — solving structures of Ligase I with imperfect DNA harboring mismatches incorporated by DNA polymerase — the authors reveal how the repair proteins maintain fidelity at final ligation step.
- Qun Tang
- , Mitchell Gulkis
- & Melike Çağlayan
-
Article
| Open Access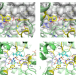 Structural basis of resistance to herbicides that target acetohydroxyacid synthase
Structural basis of resistance to herbicides that target acetohydroxyacid synthase
Acetohydroxyacid synthase (AHAS) is the target of more than 50 commercial herbicides, with many site-of-action resistance isolates identified in weeds. Here, the authors report the structural and kinetic characterizations to explain the effect AHAS mutations have on herbicide potency.
- Thierry Lonhienne
- , Yan Cheng
- & Luke W. Guddat
-
Article
| Open Access RING domains act as both substrate and enzyme in a catalytic arrangement to drive self-anchored ubiquitination
RING domains act as both substrate and enzyme in a catalytic arrangement to drive self-anchored ubiquitination
The mechanism by which RING E3-anchored ubiquitin chains are formed is not well understood. Here, the authors solve a crystal structure of the RING E3 enzyme TRIM21 trapped in the process of self-anchored chain elongation and provide biochemical and cellular insights into the mechanism of ubiquitin conjugation.
- Leo Kiss
- , Dean Clift
- & Leo C. James
-
Article
| Open Access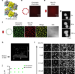 Phase separation by the polyhomeotic sterile alpha motif compartmentalizes Polycomb Group proteins and enhances their activity
Phase separation by the polyhomeotic sterile alpha motif compartmentalizes Polycomb Group proteins and enhances their activity
The conserved SAM motif of Polycomb Repressive Complex 1 subunit Ph has been shown to play an important role in chromatin organization. Here, the authors study the effect of Ph SAM on chromatin in vitro, showing that it induces the formation of concentrated, phase-separated condensates, which enhance the ubiquitin ligase activity of PRC1.
- Elias Seif
- , Jin Joo Kang
- & Nicole J. Francis
-
Article
| Open Access Polyanions provide selective control of APC/C interactions with the activator subunit
Polyanions provide selective control of APC/C interactions with the activator subunit
Activation of APC/C depends on transient interactions with an activator subunit. Here, the authors show that cellular polyanions trigger APC/C-activator dissociation and that high-affinity substrates block this effect, suggesting that substrate availability controls APC/C-activator binding.
- Arda Mizrak
- & David O. Morgan
-
Article
| Open Access A tri-ionic anchor mechanism drives Ube2N-specific recruitment and K63-chain ubiquitination in TRIM ligases
A tri-ionic anchor mechanism drives Ube2N-specific recruitment and K63-chain ubiquitination in TRIM ligases
The E3 ligase TRIM21 targets a broad range of pathogenic proteins using a unique mechanism. Here, the authors combine structural, biochemical and cell biological approaches to elucidate how TRIM21 selectively recruits its canonical E2 partner enzyme Ube2N and catalyzes ubiquitination.
- Leo Kiss
- , Jingwei Zeng
- & Leo C. James
-
Article
| Open Access Arkadia/RNF111 is a SUMO-targeted ubiquitin ligase with preference for substrates marked with SUMO1-capped SUMO2/3 chain
Arkadia/RNF111 is a SUMO-targeted ubiquitin ligase with preference for substrates marked with SUMO1-capped SUMO2/3 chain
The cellular functions of poly-SUMO chains of different compositions are not fully understood. Here, the authors characterize Arkadia/RNF111 as a SUMO-targeted ubiquitin ligase that recognizes proteins with hybrid SUMO1-capped SUMO2/3 chains and targets them for proteasomal degradation.
- Annie M. Sriramachandran
- , Katrin Meyer-Teschendorf
- & R. Jürgen Dohmen
-
Article
| Open Access A switch element in the autophagy E2 Atg3 mediates allosteric regulation across the lipidation cascade
A switch element in the autophagy E2 Atg3 mediates allosteric regulation across the lipidation cascade
Autophagy mediated by the conjugation pathway for ubiquitin-like proteins plays a key role in controlling homeostasis in eukaryotic cells. Here the authors provide a molecular basis for allosteric activation of the E2 ligase Atg3, uncovering the mechanism underlying Atg8 lipidation and a novel mechanism regulating E1-E2-E3-mediated ubiquitin-like protein conjugation.
- Yumei Zheng
- , Yu Qiu
- & Brenda A. Schulman
-
Article
| Open Access A multi-lock inhibitory mechanism for fine-tuning enzyme activities of the HECT family E3 ligases
A multi-lock inhibitory mechanism for fine-tuning enzyme activities of the HECT family E3 ligases
HECT type E3 ligases are key regulators of cell growth and proliferation. Here the authors present the crystal structures of the Nedd4 family E3 ligase WWP1 in a closed and semi-open state and in combination with mutagenesis experiments identify a multi-lock regulatory mechanism that allows the fine-tuning of activities of Nedd4 family E3 ligases.
- Zhen Wang
- , Ziheng Liu
- & Wenyu Wen
-
Article
| Open Access Structural basis of specific H2A K13/K15 ubiquitination by RNF168
Structural basis of specific H2A K13/K15 ubiquitination by RNF168
Ubiquitination of histone H2A can occur on distinct lysine residues, but how each site is recognised by the specific E3 ligase remains poorly understood. Here the authors demonstrate that the E3 ligase RNF168 binds the acidic patch on the nucleosome surface, directing the E2 to the target lysine K13/K15.
- Velten Horn
- , Michael Uckelmann
- & Hugo van Ingen
-
Article
| Open Access An E2-ubiquitin thioester-driven approach to identify substrates modified with ubiquitin and ubiquitin-like molecules
An E2-ubiquitin thioester-driven approach to identify substrates modified with ubiquitin and ubiquitin-like molecules
Ubiquitination and ubiquitin-like modifications of proteins regulate multiple cellular processes but identifying substrates of specific E2 and E3 enzymes remains challenging. Here, the authors conjugate E2 enzymes with enrichable ubiquitin derivatives to identify substrates of specific E2/E3 pairs by mass spectrometry.
- Gábor Bakos
- , Lu Yu
- & Jörg Mansfeld
-
Article
| Open Access Oligomerization-primed coiled-coil domain interaction with Ubc13 confers processivity to TRAF6 ubiquitin ligase activity
Oligomerization-primed coiled-coil domain interaction with Ubc13 confers processivity to TRAF6 ubiquitin ligase activity
Ubiquitin ligase TRAF6 catalyzes assembly of free polyubiquitin chains for TAK1 activation in the IL-1R/TLR pathways, but the mechanism underlying its processivity is unclear. Here, the authors show that TRAF6 coiled-coil oligomerization domain primes its interaction with Ubc13/Ub~Ubc13 to confer processivity.
- Lin Hu
- , Jiafeng Xu
- & Zongping Xia
-
Article
| Open Access Ufd2p synthesizes branched ubiquitin chains to promote the degradation of substrates modified with atypical chains
Ufd2p synthesizes branched ubiquitin chains to promote the degradation of substrates modified with atypical chains
How ubiquitination affects the proteins it modifies varies according to the type of linkage between ubiquitin moieties. Here, Liuet al. show how yeast Udf2p promotes K48 linkage formation onto K29-linked chains to generate branched K29-K48 ubiquitin chains that target its substrate to the proteasome.
- Chao Liu
- , Weixiao Liu
- & Wei Li
-
Article
| Open Access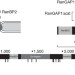 The RanBP2/RanGAP1*SUMO1/Ubc9 SUMO E3 ligase is a disassembly machine for Crm1-dependent nuclear export complexes
The RanBP2/RanGAP1*SUMO1/Ubc9 SUMO E3 ligase is a disassembly machine for Crm1-dependent nuclear export complexes
Continuous cycles of nucleo-cytoplasmic transport require disassembly of transport receptor-Ran-GTP complexes in the cytoplasm. Here the authors elucidate the specific function of the RanBP2 complex in the disassembly process.
- Tobias Ritterhoff
- , Hrishikesh Das
- & Frauke Melchior
-
Article
| Open Access The dynamic organization of fungal acetyl-CoA carboxylase
The dynamic organization of fungal acetyl-CoA carboxylase
Acetyl-CoA carboxylases are central regulatory hubs of fatty acid metabolism and are important targets for drug development in obesity and cancer. Here, the authors demonstrate that the regulation of these highly dynamic enzymes in fungi is governed by a mechanism based on phosphorylation-dependent conformational variability.
- Moritz Hunkeler
- , Edward Stuttfeld
- & Timm Maier
-
Article
| Open Access MDM2 E3 ligase-mediated ubiquitination and degradation of HDAC1 in vascular calcification
MDM2 E3 ligase-mediated ubiquitination and degradation of HDAC1 in vascular calcification
Vascular calcification (VC) increases morbidity and mortality in cardiovascular and metabolic diseases. Here, Kwon et al.show that calcification stimuli induce MDM2- mediated ubiquitination and proteasomal degradation of HDAC1, suggesting a possible therapeutic strategy for treatment of VC patients.
- Duk-Hwa Kwon
- , Gwang Hyeon Eom
- & Hyun Kook
-
Article
| Open Access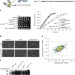 Targeting of SUMO substrates to a Cdc48–Ufd1–Npl4 segregase and STUbL pathway in fission yeast
Targeting of SUMO substrates to a Cdc48–Ufd1–Npl4 segregase and STUbL pathway in fission yeast
The small ubiquitin-like modifier SUMO can be conjugated to hundreds of protein species to affect their stability or activity. Here the authors use a quantitative proteomics approach to identify sumoylated proteins modified by the STUbL and Ufd1 pathways in fission yeast.
- Julie Bonne Køhler
- , Triin Tammsalu
- & Geneviève Thon
-
Article
| Open Access Degradation of the ABA co-receptor ABI1 by PUB12/13 U-box E3 ligases
Degradation of the ABA co-receptor ABI1 by PUB12/13 U-box E3 ligases
Signaling by the plant hormone abscisic acid (ABA) is regulated by the ABI1 protein phosphatase. Here Kong et al.propose that ABA signaling is fine-tuned by ubiquitination of ABI1 which promotes ABI degradation in response to ABA.
- Lingyao Kong
- , Jinkui Cheng
- & Zhizhong Gong
-
Article |
 The fidelity of the ligation step determines how ends are resolved during nonhomologous end joining
The fidelity of the ligation step determines how ends are resolved during nonhomologous end joining
DNA double strand breaks result in various types of damaged termini and are resolved by non-homologous end joining, but how cells coordinate the different steps that occur during repair is not clear. Here the authors show that a DNA ligase coordinates processing prior to the ligation step to limit errors.
- Crystal A. Waters
- , Natasha T. Strande
- & Dale A. Ramsden
-
Article |
 Potential for interdependent development of tRNA determinants for aminoacylation and ribosome decoding
Potential for interdependent development of tRNA determinants for aminoacylation and ribosome decoding
Aminoacyl-transfer RNA synthetases are conserved between bacteria and eukaryotes; however, bacterial enzymes cannot acylate eukaryote tRNAs. Now, fusion of a human and bacterial enzyme is shown to overcome the species barrier and confer tRNA specificity during both codon selection and proofreading on the ribosome.
- Cuiping Liu
- , Howard Gamper
- & Ya-Ming Hou

 Auranofin targets UBA1 and enhances UBA1 activity by facilitating ubiquitin trans-thioesterification to E2 ubiquitin-conjugating enzymes
Auranofin targets UBA1 and enhances UBA1 activity by facilitating ubiquitin trans-thioesterification to E2 ubiquitin-conjugating enzymes