Featured
-
-
Article
| Open Access Cell Painting-based bioactivity prediction boosts high-throughput screening hit-rates and compound diversity
Cell Painting-based bioactivity prediction boosts high-throughput screening hit-rates and compound diversity
Identifying active compounds for a target is time- and resource-intensive. Here, the authors show that deep learning models trained on Cell Painting and single-point activity data, can reliably predict compound activity across diverse targets while maintaining high hit rates and scaffold diversity.
- Johan Fredin Haslum
- , Charles-Hugues Lardeau
- & Erik Müllers
-
Article
| Open Access Microenvironmental reorganization in brain tumors following radiotherapy and recurrence revealed by hyperplexed immunofluorescence imaging
Microenvironmental reorganization in brain tumors following radiotherapy and recurrence revealed by hyperplexed immunofluorescence imaging
Improved imaging techniques are required to help advance our understanding of the complex role of the tumour microenvironment (TME). Here, the authors develop a high-throughput, highly multiplexed tissue visualisation workflow and demonstrate its utility by characterising the response of the TME to radiotherapy in preclinical models of glioblastoma.
- Spencer S. Watson
- , Benoit Duc
- & Johanna A. Joyce
-
Article
| Open Access Teacher-student collaborated multiple instance learning for pan-cancer PDL1 expression prediction from histopathology slides
Teacher-student collaborated multiple instance learning for pan-cancer PDL1 expression prediction from histopathology slides
PDL1 expression is a common biomarker for immunotherapy response in cancer, and it is usually quantified using immunohistochemistry. Here, the authors develop a weakly supervised learning approach combining multiple instance learning and a teacher-student framework to predict PDL1 expression from histopathological imaging.
- Darui Jin
- , Shangying Liang
- & Xiangzhi Bai
-
Article
| Open Access DeepDOF-SE: affordable deep-learning microscopy platform for slide-free histology
DeepDOF-SE: affordable deep-learning microscopy platform for slide-free histology
Histopathology can be limited by the time-consuming and labour-intensive preparation of slides from resected tissue. Here, the authors report DeepDOF-SE, a deep-learning-enabled microscope to rapidly scan intact tissue at cellular resolution without the need for physical sectioning.
- Lingbo Jin
- , Yubo Tang
- & Ashok Veeraraghavan
-
Article
| Open Access Mapping cell-to-tissue graphs across human placenta histology whole slide images using deep learning with HAPPY
Mapping cell-to-tissue graphs across human placenta histology whole slide images using deep learning with HAPPY
Placenta histopathology for maternal and newborn health is highly specialised and time consuming. Here, authors present a deep learning pipeline for quantifying cells and tissues in placenta whole slide images, revealing biological and clinical insights.
- Claudia Vanea
- , Jelisaveta Džigurski
- & Christoffer Nellåker
-
Article
| Open Access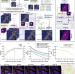 SpiDe-Sr: blind super-resolution network for precise cell segmentation and clustering in spatial proteomics imaging
SpiDe-Sr: blind super-resolution network for precise cell segmentation and clustering in spatial proteomics imaging
Imaging mass cytometry (IMC) is a powerful single-cell resolution platform for targeted spatial proteomics, but it can be constrained by imaging noise and resolution. Here, the authors propose SpiDe-Sr, a super-resolution network embedded with a denoising module for IMC spatial resolution enhancement.
- Rui Chen
- , Jiasu Xu
- & Xianting Ding
-
Article
| Open Access DeepETPicker: Fast and accurate 3D particle picking for cryo-electron tomography using weakly supervised deep learning
DeepETPicker: Fast and accurate 3D particle picking for cryo-electron tomography using weakly supervised deep learning
Picking particles of biological macromolecules is critical for solving their structures in situ using cryo-electron tomograms. Here, authors develop DeepETPicker, a deep learning-based tool for fast, accurate, and automated picking of three-dimensional particles.
- Guole Liu
- , Tongxin Niu
- & Ge Yang
-
Article
| Open Access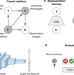 Learning representations for image-based profiling of perturbations
Learning representations for image-based profiling of perturbations
Assessing cell phenotypes in image-based assays requires solid computational methods for transforming images into quantitative data. Here, the authors present a strategy for learning representations of treatment effects from high-throughput imaging, following a causal interpretation.
- Nikita Moshkov
- , Michael Bornholdt
- & Juan C. Caicedo
-
Article
| Open Access A deep-learning-based framework for identifying and localizing multiple abnormalities and assessing cardiomegaly in chest X-ray
A deep-learning-based framework for identifying and localizing multiple abnormalities and assessing cardiomegaly in chest X-ray
Accurate localization of abnormalities is crucial in the interpretation of chest X-rays. Here the authors present a deep learning framework for simultaneous localization of 14 thoracic abnormalities and calculation of cardiothoracic ratio, based on large X-ray dataset with bounding boxes created via a human-in-the-loop approach.
- Weijie Fan
- , Yi Yang
- & Dong Zhang
-
Article
| Open Access Regression-based Deep-Learning predicts molecular biomarkers from pathology slides
Regression-based Deep-Learning predicts molecular biomarkers from pathology slides
Cancer biomarkers are often continuous measurements, which poses challenges for their prediction using classification-based deep learning. Here, the authors develop a regression-based deep learning method to predict continuous biomarkers - such as the homologous repair deficiency score - from cancer histopathology images.
- Omar S. M. El Nahhas
- , Chiara M. L. Loeffler
- & Jakob Nikolas Kather
-
Article
| Open Access DeepFocus: fast focus and astigmatism correction for electron microscopy
DeepFocus: fast focus and astigmatism correction for electron microscopy
High-throughput electron microscopy demands minimal human intervention and high image quality. Here, authors introduce DeepFocus, a data-driven method for aberration correction in electron microscopy, robust for low SNR images, fast and easily adaptable to microscopes and samples. Peer Review Information: Nature Communications thanks Yang Zhang and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
- P. J. Schubert
- , R. Saxena
- & J. Kornfeld
-
Article
| Open Access BIDCell: Biologically-informed self-supervised learning for segmentation of subcellular spatial transcriptomics data
BIDCell: Biologically-informed self-supervised learning for segmentation of subcellular spatial transcriptomics data
Subcellular in situ spatial transcriptomics offers the promise to address biological problems that were previously inaccessible but requires accurate cell segmentation to uncover insights. Here, authors present BIDCell, a biologically informed, deep learning-based cell segmentation framework.
- Xiaohang Fu
- , Yingxin Lin
- & Jean Y. H. Yang
-
Article
| Open Access Improving deep neural network generalization and robustness to background bias via layer-wise relevance propagation optimization
Improving deep neural network generalization and robustness to background bias via layer-wise relevance propagation optimization
Image background features can undesirably affect deep networks’ decisions. Here, the authors show that the optimization of Layer-wise Relevance Propagation explanation heatmaps can hinder such influence, improving out-of-distribution generalization.
- Pedro R. A. S. Bassi
- , Sergio S. J. Dertkigil
- & Andrea Cavalli
-
Article
| Open Access Petascale pipeline for precise alignment of images from serial section electron microscopy
Petascale pipeline for precise alignment of images from serial section electron microscopy
Segmentation accuracy of serial section electron microscopy (ssEM) images can be limited by the step of aligning 2D section images to create a 3D image stack. Here the authors report a computational pipeline for aligning ssEM images and apply this to a whole fly brain dataset.
- Sergiy Popovych
- , Thomas Macrina
- & H. Sebastian Seung
-
Article
| Open Access Evolutionary design of explainable algorithms for biomedical image segmentation
Evolutionary design of explainable algorithms for biomedical image segmentation
Deep learning frameworks require large human-annotated datasets for training and the resulting ‘black box’ models are difficult to interpret. Here, the authors present Kartezio; a modular Cartesian Genetic Programming-based computational strategy that generates fully transparent and easily interpretable image processing pipelines.
- Kévin Cortacero
- , Brienne McKenzie
- & Sylvain Cussat-Blanc
-
Article
| Open Access Revealing invisible cell phenotypes with conditional generative modeling
Revealing invisible cell phenotypes with conditional generative modeling
Biological research relies on observing cell phenotypes, often obscured by natural variability. Here, the authors use generative modelling to unveil hidden changes triggered by infections, mutations, or drugs, allowing for accessible discovery of biomarkers.
- Alexis Lamiable
- , Tiphaine Champetier
- & Auguste Genovesio
-
Article
| Open Access DeepSlice: rapid fully automatic registration of mouse brain imaging to a volumetric atlas
DeepSlice: rapid fully automatic registration of mouse brain imaging to a volumetric atlas
Navigating the complex structure of the brain poses a challenge to neuroscientists. Here, the authors have trained an AI (DeepSlice) that can automatically register brain images with speed and accuracy, thus simplifying this process.
- Harry Carey
- , Michael Pegios
- & Simon McMullan
-
Article
| Open Access Demonstration of an AI-driven workflow for autonomous high-resolution scanning microscopy
Demonstration of an AI-driven workflow for autonomous high-resolution scanning microscopy
Modern microscopes can image a sample with sub-Angstrom and sub-picosecond resolutions, but this often requires analysis of tremendously large datasets. Here, the authors demonstrate that an autonomous experiment can yield over a 70% reduction in dataset size while still producing high-fidelity images of the sample.
- Saugat Kandel
- , Tao Zhou
- & Mathew J. Cherukara
-
Article
| Open Access Self-supervised learning with application for infant cerebellum segmentation and analysis
Self-supervised learning with application for infant cerebellum segmentation and analysis
Neuroimaging of the cerebellum in infants has been challenging. Here the authors describe a framework for cerebellum MRI segmentation in infants up to 2 years.
- Yue Sun
- , Limei Wang
- & Li Wang
-
Article
| Open Access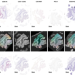 Virtual alignment of pathology image series for multi-gigapixel whole slide images
Virtual alignment of pathology image series for multi-gigapixel whole slide images
The spatial organization of a tumor affects how it grows and responds to treatment. Here, the authors present VALIS, a software to align sets of whole slide images (WSI) with state-of-the-art accuracy, enabling spatial studies of the tumor ecology.
- Chandler D. Gatenbee
- , Ann-Marie Baker
- & Alexander R. A. Anderson
-
Article
| Open Access Super-resolved trajectory-derived nanoclustering analysis using spatiotemporal indexing
Super-resolved trajectory-derived nanoclustering analysis using spatiotemporal indexing
Existing single-molecule localization microscopy analyses overlook important temporal information in living cells. Here, the authors report nanoscale spatiotemporal indexing clustering (NASTIC), which leverages a video game algorithm to fast-track the investigation of the complex temporal dynamics of protein clustering.
- Tristan P. Wallis
- , Anmin Jiang
- & Frédéric A. Meunier
-
Article
| Open Access Spatial analysis with SPIAT and spaSim to characterize and simulate tissue microenvironments
Spatial analysis with SPIAT and spaSim to characterize and simulate tissue microenvironments
Spatial proteomic data serve to provide cell-level location information for the extraction of biological features from tissues, but analyzing such data can be difficult. Here the authors report the development of SPIAT for data analyses and spaSim for simulation and validation of methods to help bridge the gap between the technology and its translation.
- Yuzhou Feng
- , Tianpei Yang
- & Anna S. Trigos
-
Article
| Open Access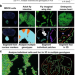 The PECAn image and statistical analysis pipeline identifies Minute cell competition genes and features
The PECAn image and statistical analysis pipeline identifies Minute cell competition genes and features
The 3D nature of clones makes sample image analysis challenging. Here the authors report PECAn, a pipeline for image processing and statistical analysis of complex multi-genotype 3D images, and apply this to the study of Minute cell competition in drosophila.
- Michael E. Baumgartner
- , Paul F. Langton
- & Eugenia Piddini
-
Article
| Open Access Privacy risks of whole-slide image sharing in digital pathology
Privacy risks of whole-slide image sharing in digital pathology
Access to Whole-Slide Images has become a cornerstone of the development of AI methods in pathology, for diagnostic use and research. Authors have developed model for privacy risks analysis and propose guidelines for safe sharing of WSI data.
- Petr Holub
- , Heimo Müller
- & Tomáš Brázdil
-
Article
| Open Access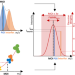 Spatial probabilistic mapping of metabolite ensembles in mass spectrometry imaging
Spatial probabilistic mapping of metabolite ensembles in mass spectrometry imaging
Spatial visualization of metabolites in tissues via mass spectrometry imaging can be prone to user perception bias. Here, the authors report the computational framework moleculaR that introduces probabilistic data-dependent molecular mapping of nonrandom spatial patterns of metabolite signals.
- Denis Abu Sammour
- , James L. Cairns
- & Carsten Hopf
-
Article
| Open Access Estimating conformational landscapes from Cryo-EM particles by 3D Zernike polynomials
Estimating conformational landscapes from Cryo-EM particles by 3D Zernike polynomials
Conformational heterogeneity is key to understand how macromolecular function and structure converge. Here, the authors propose an algorithm designed to estimate structural landscapes directly from Cryo-EM particle images.
- D. Herreros
- , R. R. Lederman
- & J. M. Carazo
-
Article
| Open Access Single-shot self-supervised object detection in microscopy
Single-shot self-supervised object detection in microscopy
Object detection using machine learning universally requires vast amounts of training datasets. Midtvedt et al. proposes a deep-learning method that enables detecting microscopic objects with sub-pixel accuracy from a single unlabeled image by exploiting the roto-translational symmetries of the problem.
- Benjamin Midtvedt
- , Jesús Pineda
- & Giovanni Volpe
-
Article
| Open Access Deep learning-based image analysis predicts PD-L1 status from H&E-stained histopathology images in breast cancer
Deep learning-based image analysis predicts PD-L1 status from H&E-stained histopathology images in breast cancer
Programmed death ligand-1 (PD-L1) has been recently adopted for breast cancer as a predictive biomarker for immunotherapies. Here, the authors show that PD-L1 expression can be predicted from H&E-stained images using deep learning.
- Gil Shamai
- , Amir Livne
- & Ron Kimmel
-
Article
| Open Access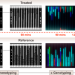 Rapid antibiotic susceptibility testing and species identification for mixed samples
Rapid antibiotic susceptibility testing and species identification for mixed samples
Rapid antibiotic susceptibility testing (AST) is needed. Here the authors report a method for phenotypic AST at the single cell level, using a microfluidic chip that allows for subsequent genotyping with in situ FISH; they apply this to a mixed sample of 7 species and 4 antibiotics.
- Vinodh Kandavalli
- , Praneeth Karempudi
- & Johan Elf
-
Article
| Open Access Adversarial attacks and adversarial robustness in computational pathology
Adversarial attacks and adversarial robustness in computational pathology
Artificial Intelligence can support diagnostic workflows in oncology, but they are vulnerable to adversarial attacks. Here, the authors show that convolutional neural networks are highly susceptible to white- and black-box adversarial attacks in clinically relevant classification tasks.
- Narmin Ghaffari Laleh
- , Daniel Truhn
- & Jakob Nikolas Kather
-
Article
| Open Access An analysis modality for vascular structures combining tissue-clearing technology and topological data analysis
An analysis modality for vascular structures combining tissue-clearing technology and topological data analysis
Understanding blood and lymphatic vasculature networks is currently limited by existing imaging systems and quantification methods. Here the authors use the tissue clearing method CUBIC to generate 3D images, machine learning to capture the signals, and extract geometric features by topological data analysis.
- Kei Takahashi
- , Ko Abe
- & Kohei Miyazono
-
Article
| Open Access Deep learning image segmentation reveals patterns of UV reflectance evolution in passerine birds
Deep learning image segmentation reveals patterns of UV reflectance evolution in passerine birds
Here, the authors develop software that uses photographs of birds to extract information on plumage UV reflectance. They use these data to show that UV reflectance is phylogenetically conserved and associated with the light environment.
- Yichen He
- , Zoë K. Varley
- & Christopher R. Cooney
-
Article
| Open Access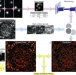 Fast DNA-PAINT imaging using a deep neural network
Fast DNA-PAINT imaging using a deep neural network
DNA-PAINT image acquisition is limited by speed. Here the authors use the neural network DeepSTORM to predict fluorophore positions from high emitter density DNA-PAINT data in order to achieve image acquisition in one minute; they demonstrate multi-colour and large-area imaging of semi-thin neuronal tissue.
- Kaarjel K. Narayanasamy
- , Johanna V. Rahm
- & Mike Heilemann
-
Article
| Open Access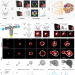 Multiscale light-sheet organoid imaging framework
Multiscale light-sheet organoid imaging framework
Live imaging of organoid growth remains a challenge: it requires long-term imaging of several samples simultaneously and dedicated analysis pipelines. Here the authors report an experimental and image processing framework to turn long-term light-sheet imaging of intestinal organoids into digital organoids.
- Gustavo de Medeiros
- , Raphael Ortiz
- & Prisca Liberali
-
Comment
| Open Access Addressing fairness in artificial intelligence for medical imaging
Addressing fairness in artificial intelligence for medical imaging
A plethora of work has shown that AI systems can systematically and unfairly be biased against certain populations in multiple scenarios. The field of medical imaging, where AI systems are beginning to be increasingly adopted, is no exception. Here we discuss the meaning of fairness in this area and comment on the potential sources of biases, as well as the strategies available to mitigate them. Finally, we analyze the current state of the field, identifying strengths and highlighting areas of vacancy, challenges and opportunities that lie ahead.
- María Agustina Ricci Lara
- , Rodrigo Echeveste
- & Enzo Ferrante
-
Article
| Open Access Enabling reactive microscopy with MicroMator
Enabling reactive microscopy with MicroMator
In microscopy, applications in which reactiveness is needed are multifarious. Here the authors report MicroMator, a Python software package for reactive experiments, which they use for applications requiring real-time tracking and light-targeting at the single-cell level.
- Zachary R. Fox
- , Steven Fletcher
- & Gregory Batt
-
Article
| Open Access Membrane marker selection for segmenting single cell spatial proteomics data
Membrane marker selection for segmenting single cell spatial proteomics data
Cell segmentation of single-cell spatial proteomics data remains a challenge and often relies on the selection of a membrane marker, which is not always known. Here, the authors introduce RAMCES, a method that selects the optimal membrane markers to use for more accurate cell segmentation.
- Monica T. Dayao
- , Maigan Brusko
- & Ziv Bar-Joseph
-
Article
| Open Access Integrating deep learning and unbiased automated high-content screening to identify complex disease signatures in human fibroblasts
Integrating deep learning and unbiased automated high-content screening to identify complex disease signatures in human fibroblasts
By coupling robotic cell culture systems with artificial intelligence–powered image analysis, Schiff et al. identify previously unseen characteristics of Parkinson’s disease in patient skin cells that distinguish them from healthy controls.
- Lauren Schiff
- , Bianca Migliori
- & Bjarki Johannesson
-
Article
| Open Access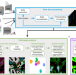 A SIMPLI (Single-cell Identification from MultiPLexed Images) approach for spatially-resolved tissue phenotyping at single-cell resolution
A SIMPLI (Single-cell Identification from MultiPLexed Images) approach for spatially-resolved tissue phenotyping at single-cell resolution
Current high-dimension imaging data analysis methods are technology-specific and require multiple tools, restricting analytical scalability and result reproducibility. Here the authors present SIMPLI, a software that overcomes these limitations for single-cell and pixel analysis of multiplexed images at spatial resolution.
- Michele Bortolomeazzi
- , Lucia Montorsi
- & Francesca D. Ciccarelli
-
Article
| Open Access Automatic mapping of multiplexed social receptive fields by deep learning and GPU-accelerated 3D videography
Automatic mapping of multiplexed social receptive fields by deep learning and GPU-accelerated 3D videography
High resolution descriptions of social interactions and their neural correlates are lacking. Here the authors report a pipeline enabling fully automatic multi-animal tracking during social encounters, together with simultaneous electrophysiological recordings, and show this works in low-light settings.
- Christian L. Ebbesen
- & Robert C. Froemke
-
Article
| Open Access 4polar-STORM polarized super-resolution imaging of actin filament organization in cells
4polar-STORM polarized super-resolution imaging of actin filament organization in cells
Single-molecule localisation microscopy does not give orientation information. Here the authors combine Stochastic Optical Reconstruction Microscopy (STORM) with single molecule orientation and wobbling measurements using four-polarisation image splitting, 4polar-STORM.
- Caio Vaz Rimoli
- , Cesar Augusto Valades-Cruz
- & Sophie Brasselet
-
Article
| Open Access ClusterMap for multi-scale clustering analysis of spatial gene expression
ClusterMap for multi-scale clustering analysis of spatial gene expression
In situ transcriptomics maps RNA expression patterns across intact tissues taking our understanding of gene expression to a new level. Here, the authors present a computational method that uncovers gene expression, cell niche, and tissue region patterns from 2D and 3D spatial transcriptomics.
- Yichun He
- , Xin Tang
- & Xiao Wang
-
Article
| Open Access Annotation-efficient deep learning for automatic medical image segmentation
Annotation-efficient deep learning for automatic medical image segmentation
Existing high-performance deep learning methods typically rely on large training datasets with high-quality manual annotations, which are difficult to obtain in many clinical applications. Here, the authors introduce an open-source framework to handle imperfect training datasets.
- Shanshan Wang
- , Cheng Li
- & Hairong Zheng
-
Article
| Open Access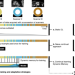 Dynamic memory to alleviate catastrophic forgetting in continual learning with medical imaging
Dynamic memory to alleviate catastrophic forgetting in continual learning with medical imaging
In clinical practice, the continuous progress of image acquisition technology or diagnostic procedures and evolving imaging protocols hamper the utility of machine learning, as prediction accuracy on new data deteriorates. Here, the authors propose a continual learning approach to deal with such domain shifts occurring at unknown time points.
- Matthias Perkonigg
- , Johannes Hofmanninger
- & Georg Langs
-
Article
| Open Access Sub-diffraction error mapping for localisation microscopy images
Sub-diffraction error mapping for localisation microscopy images
Determining the quality of localisation microscopy images is currently challenging. Here the authors report use of the Haar wavelet kernel analysis (HAWK) Method for the Assessment of Nanoscopy, termed HAWKMAN, to assess the reliability of localisation information.
- Richard J. Marsh
- , Ishan Costello
- & Susan Cox
-
Article
| Open Access The structural dynamics of macropinosome formation and PI3-kinase-mediated sealing revealed by lattice light sheet microscopy
The structural dynamics of macropinosome formation and PI3-kinase-mediated sealing revealed by lattice light sheet microscopy
Macropinocytosis is a cellular process for the uptake of extracellular fluid. Here, the authors use lattice light sheet microscopy to examine the spatial dynamics of the plasma membrane, PI3K activity, and structural differences of various macrophage cell types during macropinocytosis.
- Shayne E. Quinn
- , Lu Huang
- & Brandon L. Scott
-
Article
| Open Access Cell segmentation-free inference of cell types from in situ transcriptomics data
Cell segmentation-free inference of cell types from in situ transcriptomics data
Inaccurate cell segmentation has been the major problem for cell-type identification and tissue characterization of the in situ spatially resolved transcriptomics data. Here we show a robust cell segmentation-free computational framework (SSAM), for identifying cell types and tissue domains in 2D and 3D.
- Jeongbin Park
- , Wonyl Choi
- & Naveed Ishaque
-
Article
| Open Access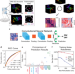 Deep learning connects DNA traces to transcription to reveal predictive features beyond enhancer–promoter contact
Deep learning connects DNA traces to transcription to reveal predictive features beyond enhancer–promoter contact
Recent advances in super-resolution microscopy have made it possible to measure chromatin 3D structure and transcription in thousands of single cells. Here, authors present a deep learning-based approach to characterise how chromatin structure relates to transcriptional state of individual cells and determine which structural features of chromatin regulation are important for gene expression state.
- Aparna R. Rajpurkar
- , Leslie J. Mateo
- & Alistair N. Boettiger
-
Article
| Open Access Regression plane concept for analysing continuous cellular processes with machine learning
Regression plane concept for analysing continuous cellular processes with machine learning
High-content screening prompted the development of software enabling discrete phenotypic analysis of single cells. Here, the authors show that supervised continuous machine learning can drive novel discoveries in diverse imaging experiments and present the Regression Plane module of Advanced Cell Classifier.
- Abel Szkalisity
- , Filippo Piccinini
- & Peter Horvath

 Double-negative B cells and DNASE1L3 colocalise with microbiota in gut-associated lymphoid tissue
Double-negative B cells and DNASE1L3 colocalise with microbiota in gut-associated lymphoid tissue