Featured
-
-
Article
| Open Access RS-1 enhances CRISPR/Cas9- and TALEN-mediated knock-in efficiency
RS-1 enhances CRISPR/Cas9- and TALEN-mediated knock-in efficiency
CRISPR/Cas9 and transcription activator-like effector nuclease (TALEN) are becoming major tools for genome editing. Here, Song et al. show that RS-1, a small-molecule enhancer for homology directed repair, increases the CRISPR/Cas9 and TALEN mediated knock-in efficiency both in vitro and in vivowith rabbit.
- Jun Song
- , Dongshan Yang
- & Jifeng Zhang
-
Article
| Open Access Complex disease and phenotype mapping in the domestic dog
Complex disease and phenotype mapping in the domestic dog
The domestic dog is an important model organism for our understanding of cancer and other diseases. Here the authors conduct a genome-wide association study across multiple breeds and identify novel loci significantly associated with several complex diseases and morphological traits.
- Jessica J. Hayward
- , Marta G. Castelhano
- & Adam R. Boyko
-
Article
| Open Access ssODN-mediated knock-in with CRISPR-Cas for large genomic regions in zygotes
ssODN-mediated knock-in with CRISPR-Cas for large genomic regions in zygotes
CRISPR-Cas9 is a powerful genome engineering tool but gene knock-in is limited by fragment size and efficiency of recombination. Here the authors used a modified strategy employing single-strand oligonucleotides to efficiently knock-in large DNA fragments and humanise native rat loci.
- Kazuto Yoshimi
- , Yayoi Kunihiro
- & Tomoji Mashimo
-
Article
| Open Access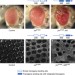 A Drosophila RNAi library modulates Hippo pathway-dependent tissue growth
A Drosophila RNAi library modulates Hippo pathway-dependent tissue growth
Drosophila RNAi libraries are commonly used to perform large-scale functional genetics screens in vivo. Here the authors find that a subset of lines from the VDRC KK RNAi line cause false-positive enhancement of the Hippo pathway, and provide a strain that can test whether a genetic screen of interest will be affected by this technical artefact.
- Joseph H.A. Vissers
- , Samuel A. Manning
- & Kieran F. Harvey
-
Article
| Open Access A generic strategy for CRISPR-Cas9-mediated gene tagging
A generic strategy for CRISPR-Cas9-mediated gene tagging
CRISPR-Cas9 has greatly enhanced genome engineering however gene tagging can still be cumbersome due to a requirement for homology donors. Here the authors introduce a generic system for gene tagging that does not require homology between the donor and the genomic target site.
- Daniel H. Lackner
- , Alexia Carré
- & Tilmann Bürckstümmer
-
Article
| Open Access PAM multiplicity marks genomic target sites as inhibitory to CRISPR-Cas9 editing
PAM multiplicity marks genomic target sites as inhibitory to CRISPR-Cas9 editing
A key step for the use of CRISPR-Cas9 in any study is the design of the guide RNA, however the underlying principles are still poorly understood. Here the authors show that multiple protospacer adjacent motif sequences are refractory to efficient targeting and repair.
- Abba Malina
- , Christopher J. F. Cameron
- & Jerry Pelletier
-
Article
| Open Access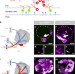 Dynamic labelling of neural connections in multiple colours by trans-synaptic fluorescence complementation
Dynamic labelling of neural connections in multiple colours by trans-synaptic fluorescence complementation
The ability to map the activity of synapses within a circuit will help in elucidating the neural basis of behaviour. Here, Macpherson et al. report new strategies to specifically label active synapses in Drosophila with multi-colour fluorescence tags.
- Lindsey J. Macpherson
- , Emanuela E. Zaharieva
- & Marco Gallio
-
Article
| Open Access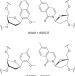 Identification of DNA lesions using a third base pair for amplification and nanopore sequencing
Identification of DNA lesions using a third base pair for amplification and nanopore sequencing
Genomic DNA lesions exist in low levels and cannot be amplified by standard PCR. Here, Riedlet al. report a method to amplify damaged DNA sites by replacing them via DNA repair with unnatural base pairs, which are subsequently identified by Sanger sequencing or α-hemolysin nanopore sequencing.
- Jan Riedl
- , Yun Ding
- & Cynthia J. Burrows
-
Article
| Open Access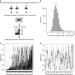 Genetic mapping uncovers cis-regulatory landscape of RNA editing
Genetic mapping uncovers cis-regulatory landscape of RNA editing
Adenosine-to-inosine (A-to-I) RNA editing plays an important role in neurological functions. Here, by a quantitative trait loci (QTL) mapping approach in 131 Drosophila melanogasterstrains, the authors identify 545 QTLs associated with differences in RNA editing.
- Gokul Ramaswami
- , Patricia Deng
- & Jin Billy Li
-
Article
| Open Access Outbred genome sequencing and CRISPR/Cas9 gene editing in butterflies
Outbred genome sequencing and CRISPR/Cas9 gene editing in butterflies
Butterflies are a promising system to study the genetics and evolution of morphological diversification, yet genomic and technological resources are limited. Here, the authors sequence genomes of two Papiliobutterflies and develop a CRISPR/Cas9 gene editing method for these species.
- Xueyan Li
- , Dingding Fan
- & Wen Wang
-
Article
| Open Access Cas9-Assisted Targeting of CHromosome segments CATCH enables one-step targeted cloning of large gene clusters
Cas9-Assisted Targeting of CHromosome segments CATCH enables one-step targeted cloning of large gene clusters
Genomic engineering often requires the cloning of long DNA segments that contain large gene clusters. Here, the authors describe an RNA-guided Cas9 nuclease assistedin vitrotechnique that allows the targeted cloning of near-arbitrary, long bacterial genomic sequences of up to 100 kb in a single step.
- Wenjun Jiang
- , Xuejin Zhao
- & Ting F. Zhu
-
Article
| Open Access Rapid and efficient one-step generation of paired gRNA CRISPR-Cas9 libraries
Rapid and efficient one-step generation of paired gRNA CRISPR-Cas9 libraries
CRISPR-Cas9 is a powerful tool for genome editing; however, difficulties in generating pools of paired guide RNAs limit its applicability to large-scale screening experiments. Here the authors report a one-step method for rapid and efficient generation of pooled libraries of guide RNA pairs.
- Joana A. Vidigal
- & Andrea Ventura
-
Article
| Open Access A genome-wide association study identifies multiple loci for variation in human ear morphology
A genome-wide association study identifies multiple loci for variation in human ear morphology
The shape of the pinna varies widely in the general human population but the genetic basis of this variation is unknown. Here Adhikari et al. conduct a genome-wide association study in Latin Americans and discover seven gene regions influencing pinna morphology, including EDAR and TBX15.
- Kaustubh Adhikari
- , Guillermo Reales
- & Andrés Ruiz-Linares
-
Article
| Open Access Rh D blood group conversion using transcription activator-like effector nucleases
Rh D blood group conversion using transcription activator-like effector nucleases
Group O/RhD− blood can be safely transfused to any recipient and methods for converting other blood groups into this group hold therapeutic potential. By using programmable nucleases, here the authors edit the gene that determines the RhD blood group and convert the RhD+ into RhD− erythroid progenitor cells.
- Young-Hoon Kim
- , Hyun O. Kim
- & Hyongbum Kim
-
Article
| Open Access Context influences on TALE–DNA binding revealed by quantitative profiling
Context influences on TALE–DNA binding revealed by quantitative profiling
TALE proteins are popular tools for genome engineering because they can recognize specific DNA sequences, however off-target effects are a routine problem. Here Rogers and Barrera et al. comprehensively map TALE–DNA interactions to develop a computational model to predict binding specificity.
- Julia M. Rogers
- , Luis A. Barrera
- & Martha L. Bulyk
-
Article
| Open Access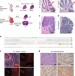 Somatic CRISPR/Cas9-mediated tumour suppressor disruption enables versatile brain tumour modelling
Somatic CRISPR/Cas9-mediated tumour suppressor disruption enables versatile brain tumour modelling
Gene transfer is a powerful technique to investigate the mechanistic basis of tumorigenesis. Here Zuckermann et al. adapt CRISPR/Cas9 genome editing to target potential oncogenes somatically in vivo, establishing a fast and convenient system for validating novel genetic candidates.
- Marc Zuckermann
- , Volker Hovestadt
- & Jan Gronych
-
Article
| Open Access Effective heritable gene knockdown in zebrafish using synthetic microRNAs
Effective heritable gene knockdown in zebrafish using synthetic microRNAs
Zebrafish is a model system for which for no reliable heritable gene silencing method is available. Here the authors provide a system for heritable miRNA-mediated knockdown and demonstrate tunable silencing of the smn1gene that recapitulate different forms of spinal muscular atrophy.
- Jean Giacomotto
- , Silke Rinkwitz
- & Thomas S. Becker
-
Article
| Open Access Drosophila glucome screening identifies Ck1alpha as a regulator of mammalian glucose metabolism
Drosophila glucome screening identifies Ck1alpha as a regulator of mammalian glucose metabolism
Diabetes is associated with aberrations in glucose metabolism. Here the authors perform a genomic screen in fruit flies to identify new regulators of fly glucose metabolism, and show that mice lacking the murine homologue of one of their hits, the protein kinase CK1alpha, in the adipose lineage develop diabetes.
- Rupali Ugrankar
- , Eric Berglund
- & Jonathan M. Graff
-
Article
| Open Access Targeted DNA degradation using a CRISPR device stably carried in the host genome
Targeted DNA degradation using a CRISPR device stably carried in the host genome
The ability to contain and destroy synthetically engineered microorganisms is an important consideration with environmental, industrial and intellectual property implications. Here Caliando et al. design and demonstrate a stably integrated CRISPR-based system for targeted DNA destruction.
- Brian J. Caliando
- & Christopher A. Voigt
-
Article |
 eQTL mapping identifies insertion- and deletion-specific eQTLs in multiple tissues
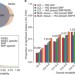
eQTL mapping identifies insertion- and deletion-specific eQTLs in multiple tissues
Expression quantitative trait loci (eQTLs) may provide insight into the functional mechanisms underlying disease risk variants. Here the authors characterize INDEL-specific eQTLs in several tissues and show that these can have both tissue-specific and tissue-consistent effects.
- Jinyan Huang
- , Jun Chen
- & Liming Liang
-
Article
| Open Access High-resolution genetic mapping of maize pan-genome sequence anchors
High-resolution genetic mapping of maize pan-genome sequence anchors
Structural variations in crop genomes are thought to be responsible for significant differences in phenotype and they can be well-represented by a ‘pan-genome’. Here, Lu et al.develop an approach to genetically map pan-genome sequence anchors using 14,129 inbred lines of maize, showing structural variation is a significant source of adaptive variation.
- Fei Lu
- , Maria C. Romay
- & Edward S. Buckler
-
Article |
 Use of the CRISPR/Cas9 system as an intracellular defense against HIV-1 infection in human cells
Use of the CRISPR/Cas9 system as an intracellular defense against HIV-1 infection in human cells
The CRISPR/Cas9 system can be used for genome editing. Here, Liao et al. show that the system can be adapted to inhibit HIV expression and replication, excise the integrated HIV genome and provide long-term protection against new infections in human cells, including pluripotent stem cells.
- Hsin-Kai Liao
- , Ying Gu
- & Juan Carlos Izpisua Belmonte
-
Article |
 Multiplex CRISPR/Cas9-based genome editing for correction of dystrophin mutations that cause Duchenne muscular dystrophy
Multiplex CRISPR/Cas9-based genome editing for correction of dystrophin mutations that cause Duchenne muscular dystrophy
Duchenne muscular dystrophy is caused by mutations in the dystrophin gene. Here, Ousterout et al. use multiplexed CRISPR/Cas9 genome editing to excise a large portion of the gene that carries over 60% of known dystrophin mutations. They show that this excision restores dystrophin expression in patient-derived cells.
- David G. Ousterout
- , Ami M. Kabadi
- & Charles A. Gersbach
-
Article
| Open Access Dense genotyping of immune-related susceptibility loci reveals new insights into the genetics of psoriatic arthritis
Dense genotyping of immune-related susceptibility loci reveals new insights into the genetics of psoriatic arthritis
Psoriatic arthritis (PsA) is a chronic inflammatory arthritis with a significant genetic component. Here, the authors analyse immune-related genetic markers in 1,962 PsA patients and 8,923 controls to identify novel PsA risk loci and highlight distinct genetic differences between psoriasis and PsA.
- John Bowes
- , Ashley Budu-Aggrey
- & Anne Barton
-
Article
| Open Access A high-throughput optomechanical retrieval method for sequence-verified clonal DNA from the NGS platform
A high-throughput optomechanical retrieval method for sequence-verified clonal DNA from the NGS platform
One of the biggest bottlenecks in large-scale DNA synthesis is the retrieval of target clonal DNA from high-density sequencing platforms. Here, the authors present a method called ‘Sniper Cloning’ that allows for precise mapping of target clone features and rapid retrieval of targets for full utilization of DNA clones.
- Howon Lee
- , Hyoki Kim
- & Sunghoon Kwon
-
Article
| Open Access Targeted and genome-wide sequencing reveal single nucleotide variations impacting specificity of Cas9 in human stem cells
Targeted and genome-wide sequencing reveal single nucleotide variations impacting specificity of Cas9 in human stem cells
The microbial RNA-guided CRISPR/Cas9 system has robust genome-editing activities, but the off-target effects of the Cas9 nuclease have only recently begun to be analysed. Here the authors provide evidence for high specificity of the Cas9 nuclease on targeting of the Tafazzin gene in human-induced pluripotent stem cells and demonstrate the impact of single-nucleotide variations of the human genome on Cas9 specificity.
- Luhan Yang
- , Dennis Grishin
- & George Church
-
Article
| Open Access Microhomology-mediated end-joining-dependent integration of donor DNA in cells and animals using TALENs and CRISPR/Cas9
Microhomology-mediated end-joining-dependent integration of donor DNA in cells and animals using TALENs and CRISPR/Cas9
One challenge facing the use of programmable nucleases in genome engineering is the requirement for homologous recombination. Here, Nakade et al.harness microhomology-mediated end-joining as a means of inserting exogenous coding sequences into the genome using both TALEN and CRISPR/Cas9 technologies.
- Shota Nakade
- , Takuya Tsubota
- & Ken-ichi T. Suzuki
-
Article |
 Synthesizing AND gate genetic circuits based on CRISPR-Cas9 for identification of bladder cancer cells
Synthesizing AND gate genetic circuits based on CRISPR-Cas9 for identification of bladder cancer cells
Tools derived from synthetic biology offer powerful means to refine drug delivery and disease detection. Liu et al. engineer a logical AND gate using CRISPR-Cas9 to drive gene expression only cells in which two promoters are active, and use it to selectively inhibit the growth of bladder cancer cells in vitro.
- Yuchen Liu
- , Yayue Zeng
- & Zhiming Cai
-
Article |
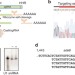 Implementation of the CRISPR-Cas9 system in fission yeast
Implementation of the CRISPR-Cas9 system in fission yeast
The fission yeast, Schizosaccharomyces pombe, is a valuable model organism, but the lack of a portable RNA Pol III promoter has prevented the implementation of the CRISPR/Cas9 system. Here the authors develop a CRISPR/Cas9 system that achieves selection-free specific mutagenesis with very high efficiencies in S. pombe.
- Jake Z. Jacobs
- , Keith M. Ciccaglione
- & Mikel Zaratiegui
-
Article
| Open Access Combining high-throughput phenotyping and genome-wide association studies to reveal natural genetic variation in rice
Combining high-throughput phenotyping and genome-wide association studies to reveal natural genetic variation in rice
Next-generation sequencing technology has made the generation of huge amounts of genetic data possible, but phenotype characterization remains slow and difficult. Here the authors develop a high-throughput phenotyping facility for rice that is able to accurately identify and characterize traits related to morphology, biomass and yield.
- Wanneng Yang
- , Zilong Guo
- & Lizhong Xiong
-
Article |
 Phenotypic characterization of missense polymerase-δ mutations using an inducible protein-replacement system
Phenotypic characterization of missense polymerase-δ mutations using an inducible protein-replacement system
The essential nature of replicative polymerases has hampered the study of polymerase-δ mutations found in colorectal cancer cells. Here, using polymerase-δ mutations as a proof of principle, the authors present an inducible single vector system that replaces any endogenous gene with an RNAi-resistant mutant version.
- Medini Manohar Ghodgaonkar
- , Patrick Kehl
- & Josef Jiricny
-
Article |
 Expansion of the CRISPR–Cas9 genome targeting space through the use of H1 promoter-expressed guide RNAs
Expansion of the CRISPR–Cas9 genome targeting space through the use of H1 promoter-expressed guide RNAs
Current CRISPR-mediated genome-editing methods are limited by the requirement for a specific +1 nucleotide when using the U6 promoter to drive guide RNA synthesis. Now, Ranganathan et al.report a modification of the CRISPR–Cas9 system that more than doubles the number of targetable CRISPR sites within the human genome.
- Vinod Ranganathan
- , Karl Wahlin
- & Donald J. Zack
-
Article
| Open Access Allele-specific genome editing and correction of disease-associated phenotypes in rats using the CRISPR–Cas platform
Allele-specific genome editing and correction of disease-associated phenotypes in rats using the CRISPR–Cas platform
The bacterial CRISPR–Cas system is increasingly used for genome editing in animal models. Here the authors utilize this system to target and edit specific coat colour alleles in rats and demonstrate the potential of this technology for the creation of genetically engineered animal models.
- K. Yoshimi
- , T. Kaneko
- & T. Mashimo
-
Article |
 Integrating sequence and array data to create an improved 1000 Genomes Project haplotype reference panel
Integrating sequence and array data to create an improved 1000 Genomes Project haplotype reference panel
1000 Genomes imputation can increase the power of genome-wide association studies to detect genetic variants associated with human traits and diseases. Here, the authors develop a method to integrate and analyse low-coverage sequence data and SNP array data, and show that it improves imputation performance.
- Olivier Delaneau
- , Jonathan Marchini
- & Leena Peltonenz
-
Article |
 Genomic mapping of phosphorothioates reveals partial modification of short consensus sequences
Genomic mapping of phosphorothioates reveals partial modification of short consensus sequences
Phosphorothioate (PT) DNA modifications are widespread in bacteria and play a critical role in cell physiology. Here, the authors develop two sequence-based technologies to map PT modifications across bacterial genomes.
- Bo Cao
- , Chao Chen
- & Peter C. Dedon
-
Article |
 Engineering human tumour-associated chromosomal translocations with the RNA-guided CRISPR–Cas9 system
Engineering human tumour-associated chromosomal translocations with the RNA-guided CRISPR–Cas9 system
CRISPR and Cas9 are endonucleases that are found in bacteria and have recently been exploited for genome engineering. Here, the authors use this system in cultured mammalian cells to engineer chromosomal translocations that are found in acute myeloid leukaemia and Ewing’s sarcoma.
- R. Torres
- , M. C. Martin
- & S. Rodriguez-Perales
-
Article |
 Targeted genomic rearrangements using CRISPR/Cas technology
Targeted genomic rearrangements using CRISPR/Cas technology
Genomic rearrangements have important functional consequences for cancer. Here, Choi and Meyerson use CRISPR/Cas technology to generate translocations and inversions that are known drivers of lung cancer, and demonstrate the utility of this technology for studying the role of genomic rearrangements in disease.
- Peter S. Choi
- & Matthew Meyerson
-
Article |
 A TAL effector repeat architecture for frameshift binding
A TAL effector repeat architecture for frameshift binding
Transcription activator-like effectors (TALEs) of pathogenic bacteria activate target genes in host plants to support infection. Here, the authors show that TALEs with single natural repeats of aberrant length tolerate one base pair deletions and may enable the bacteria to overcome natural plant resistance.
- Annekatrin Richter
- , Jana Streubel
- & Jens Boch
-
Article |
 Surrogate reporter-based enrichment of cells containing RNA-guided Cas9 nuclease-induced mutations
Surrogate reporter-based enrichment of cells containing RNA-guided Cas9 nuclease-induced mutations
RNA-guided endonucleases (RGENs) are promising tools for genome editing, although their limited activity poses a challenge. Here the authors use surrogate reporters to achieve enrichment of cells containing RGEN-induced mutations up to 11-fold, which could help facilitate the use of RGENs in biomedical research.
- Suresh Ramakrishna
- , Seung Woo Cho
- & Hyongbum Kim
-
Article |
 Genotyping with CRISPR-Cas-derived RNA-guided endonucleases
Genotyping with CRISPR-Cas-derived RNA-guided endonucleases
Cas9 RNA-guided engineered nucleases (RGENs) induce site-specific DNA cleavages in cultured cells and organisms and are used widely as genome-editing tools. Here, the authors develop an RGEN-based technology to genotype both RGEN-induced mutations and cancer-associated mutations in human cell lines.
- Jong Min Kim
- , Daesik Kim
- & Jin-Soo Kim
-
Article |
 Efficient genome engineering by targeted homologous recombination in mouse embryos using transcription activator-like effector nucleases
Efficient genome engineering by targeted homologous recombination in mouse embryos using transcription activator-like effector nucleases
Genetically engineered mice are an important aspect of human disease research. Here, the authors use artificial transcription activator-like effector-nucleases to generate a mouse line with a conditionally targeted allele and suggest that this method can be easily adapted to any gene in the mouse genome.
- Daniel Sommer
- , Annika E. Peters
- & Marc Beyer
-
Article |
 Using synthetic templates to design an unbiased multiplex PCR assay
Using synthetic templates to design an unbiased multiplex PCR assay
Immunosequencing enables cost-effective sequencing of repertoires of immune cells, but it often suffers from amplification biases when attempting cell quantification. Here, the authors present a powerful multiplex PCR assay that allows for quantitative and unbiased analysis of frequency of different T cell receptors.
- Christopher S. Carlson
- , Ryan O. Emerson
- & Harlan Robins
-
Article
| Open Access Zinc-finger nickase-mediated insertion of the lysostaphin gene into the beta-casein locus in cloned cows
Zinc-finger nickase-mediated insertion of the lysostaphin gene into the beta-casein locus in cloned cows
Zinc-finger nickases are programmable nucleases that can be used to generate site-specific single-strand breaks in DNA. Liu et al. use this technology to insert an antimicrobial gene into the endogenous beta-casein locus in cloned cows, with the aim of providing protection against mastitis.
- Xu Liu
- , Yongsheng Wang
- & Yong Zhang
-
Article
| Open Access Compact designer TALENs for efficient genome engineering
Compact designer TALENs for efficient genome engineering
Transcription activator-like effector nucleases (TALENs) are dimeric 'molecular scissors' that can be readily engineered for gene-targeting applications. Beurdeley et al. develop a single-chain TALEN architecture having significant in vivoactivity in yeast, plant and mammalian systems.
- Marine Beurdeley
- , Fabian Bietz
- & George H. Silva
-
Article |
 Tet-mediated covalent labelling of 5-methylcytosine for its genome-wide detection and sequencing
Tet-mediated covalent labelling of 5-methylcytosine for its genome-wide detection and sequencing
A number of methylome sequencing technologies depend on affinity purification of methylated DNA. Zhang et al. demonstrate a click-chemistry-based protocol for covalently labelling 5-methylcytosine residues with biotin, providing enhanced sensitivity and specificity compared with antibody-based enrichment.
- Liang Zhang
- , Keith E. Szulwach
- & Chuan He
-
Article |
 Cleavage-based signal amplification of RNA
Cleavage-based signal amplification of RNA
RNA detection is important in biomedical research and largely relies on the reverse transcription–PCR reaction. Zhao et al.report an isothermal reaction, which involves cleavage by a DNAzyme and signal amplification, to simultaneously amplify and detect RNA.
- Yongyun Zhao
- , Li Zhou
- & Zhuo Tang
-
Article
| Open Access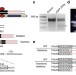 Non-transgenic genome modifications in a hemimetabolous insect using zinc-finger and TAL effector nucleases
Non-transgenic genome modifications in a hemimetabolous insect using zinc-finger and TAL effector nucleases
Hemimetabolous insects comprise many pests but introducing targeted mutations into these species has been difficult. This paper reports efficient targeted mutagenesis, and the generation of homozygous knockouts, in crickets based on zinc finger nucleases or transcription activator-like effector nucleases.
- Takahito Watanabe
- , Hiroshi Ochiai
- & Taro Mito
-
Article |
 Reprogramming within hours following nuclear transfer into mouse but not human zygotes
Reprogramming within hours following nuclear transfer into mouse but not human zygotes
The generation of human cell lines using somatic cell nuclear transfer has been difficult to achieve. In this study, Egliet al. show that while mouse eggs reprogram somatic cells within hours, human eggs arrest after nuclear transfer which may be due to a lack of genome transcription.
- Dieter Egli
- , Alice E. Chen
- & Kevin Eggan

 Multiplexed pancreatic genome engineering and cancer induction by transfection-based CRISPR/Cas9 delivery in mice
Multiplexed pancreatic genome engineering and cancer induction by transfection-based CRISPR/Cas9 delivery in mice