Featured
-
-
News & Views |
 Excessive mobility interrupted
Excessive mobility interrupted
Mobile DNA sequences called L1 contribute to the brain's genetic heterogeneity and may affect neuron function. The protein MeCP2, which is mutated in Rett syndrome, seems to regulate the activity of these genomic elements. See Letter p.443
- Lorenz Studer
-
Letter |
 Formation, regulation and evolution of Caenorhabditis elegans 3′UTRs
Formation, regulation and evolution of Caenorhabditis elegans 3′UTRs
When genes are transcribed into RNA, the polymerase extends beyond the end of the protein-coding portion to make the 3′ untranslated region (UTR); this region contains important regulatory sequences, such as microRNA binding sites, and facilitates translation. Long tracts of untemplated adenines are added to the 3′ UTR, and the standard method for sequencing the transcriptome is based on hybridization to the poly(A) tail. A new high-throughput approach to transcriptome sequencing is reported that avoids a known limitation of the poly(A) method; the method is used to provide a more accurate analysis of functional and evolutionary aspects of 3′ UTRs of the nematode.
- Calvin H. Jan
- , Robin C. Friedman
- & David P. Bartel
-
Letter |
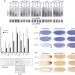 Maternal mRNA deadenylation and decay by the piRNA pathway in the early Drosophila embryo
Maternal mRNA deadenylation and decay by the piRNA pathway in the early Drosophila embryo
Piwi-associated RNAs (piRNAs) are small RNAs with several functions in the germline, such as repressing transposable elements and helping to maintain germline stem cells. Now, a function for piRNAs has been discovered outside the germline, in the fruitfly embryo. Specifically, piRNAs are required for the decay of the messenger RNA encoding the posterior morphogen Nanos. When piRNA-induced regulation is impaired, this mRNA is stabilized and developmental defects ensue.
- Christel Rouget
- , Catherine Papin
- & Martine Simonelig
-
Research Highlights |
Neurobiology: Powerless against Parkinson's
-
Letter |
 xnd-1 regulates the global recombination landscape in Caenorhabditis elegans
xnd-1 regulates the global recombination landscape in Caenorhabditis elegans
To facilitate their proper segregation, duplicated meiotic chromosomes are physically joined by crossovers. Crossover formation begins with the introduction of meiosis-specific double-strand breaks. These authors identify a new gene in Caenorhabditis elegans, xnd-1, that is required for crossover distribution on both the X and the autosomal chromosomes. Preliminary data suggest that xnd-1 does this by regulating acetylation of histone H2A on lysine 5.
- Cynthia R. Wagner
- , Lynnette Kuervers
- & Judith L. Yanowitz
-
Letter |
 Role of Tet proteins in 5mC to 5hmC conversion, ES-cell self-renewal and inner cell mass specification
Role of Tet proteins in 5mC to 5hmC conversion, ES-cell self-renewal and inner cell mass specification
TET1 is an enzyme that catalyses the conversion of 5-methylcytosine of DNA to 5-hydroxymethylcytosine, raising the possibility that it is involved in mediating DNA demethylation. These authors show that Tet1 is involved in mouse embryonic stem cell maintenance and specification of the inner cell mass. It is required to maintain both the expression of Nanog in embryonic stem cells and the Nanog promoter in a hypomethylated state, supporting a role for Tet1 in regulating DNA methylation.
- Shinsuke Ito
- , Ana C. D’Alessio
- & Yi Zhang
-
Article |
 Mammalian microRNAs predominantly act to decrease target mRNA levels
Mammalian microRNAs predominantly act to decrease target mRNA levels
MicroRNAs are known to affect the levels of both messenger RNA (mRNA) and protein. But as protein production is dependent on the presence of mRNA, it was not clear what the relative contributions of microRNA-mediated mRNA cleavage and translational repression were. These authors have parsed out the two mechanisms, and unexpectedly find that microRNAs function primarily by affecting mRNA levels rather than their translation. This suggests a reassessment of many previous conclusions is necessary.
- Huili Guo
- , Nicholas T. Ingolia
- & David P. Bartel
-
Letter |
 OncomiR addiction in an in vivo model of microRNA-21-induced pre-B-cell lymphoma
OncomiR addiction in an in vivo model of microRNA-21-induced pre-B-cell lymphoma
One model for cancer development posits that the proliferating cells in a tumour can become 'addicted' to activating mutations in an oncogene. With the realization that certain microRNAs promote tumorigenesis, it has been proposed that tumours may also become dependent on such 'oncomiRs'. Here, evidence is provided that the gene encoding microRNA-21 is an oncogene, and that in its absence, tumours undergo apoptosis and regress. Thus tumours can indeed become addicted to oncomiRs.
- Pedro P. Medina
- , Mona Nolde
- & Frank J. Slack
-
Article |
 From noncoding variant to phenotype via SORT1 at the 1p13 cholesterol locus
From noncoding variant to phenotype via SORT1 at the 1p13 cholesterol locus
A non-coding polymorphism at a locus associated with myocardial infarction in humans creates a CCAAT/enhancer binding protein transcription factor binding site and alters the hepatic expression of the SORT1 gene. These authors show that modulating Sort1 levels in mouse liver alters levels of plasma low-density lipoprotein cholesterol and very low-density lipoprotein, potentially explaining why polymorphisms at this locus are associated with heart disease.
- Kiran Musunuru
- , Alanna Strong
- & Daniel J. Rader
-
Letter |
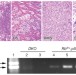 Rb regulates fate choice and lineage commitment in vivo
Rb regulates fate choice and lineage commitment in vivo
The retinoblastoma tumour suppressor protein pRb can suppress the activity of certain transcription factors and potentiate the activity of others, and has been shown to affect the differentiation of different cell lineages in vitro. These authors show that the Rb gene has a role in driving bone cell formation or brown adipose tissue formation in vivo.
- Eliezer Calo
- , Jose A. Quintero-Estades
- & Jacqueline A. Lees
-
Letter |
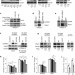 Pathogenic LRRK2 negatively regulates microRNA-mediated translational repression
Pathogenic LRRK2 negatively regulates microRNA-mediated translational repression
LRRK2 activity is dysregulated in Parkinson's disease, but how it contributes to the pathogenesis is unknown. These authors show that Drosophila LRRK2 interacts with the Argonaute component (dAgo1) of the RNA-induced silencing complex. This is associated with reduced levels of dAgo1, interaction between phospho-4E-BP1 and hAgo2, and impairment of microRNA-mediated repression. This leads to overexpression of the cell cycle genes e2f1 and dp, and consequent degeneration of dopaminergic neurons.
- Stephan Gehrke
- , Yuzuru Imai
- & Bingwei Lu
-
-
Letter |
 Regulation of myeloid leukaemia by the cell-fate determinant Musashi
Regulation of myeloid leukaemia by the cell-fate determinant Musashi
Chronic myelogenous leukaemia (CML) can progress from a chronic to an acute phase. These authors show in mouse models that leukaemia progression is controlled by the cell-fate regulator Musashi2, which in turn regulates Numb, Notch and p53 to block cellular differentiation. Musashi2 expression can be increased by aberrant transcription factors found in leukaemia, is observed during cancer progression in human CML patients and is associated with poorer prognosis.
- Takahiro Ito
- , Hyog Young Kwon
- & Tannishtha Reya
-
News |
 Mystery RNA spawns gene-activating peptides
Mystery RNA spawns gene-activating peptides
Short peptides that regulate fruitfly development are produced from 'junk' RNA.
- Heidi Ledford
-
Letter |
 Histone H4K20/H3K9 demethylase PHF8 regulates zebrafish brain and craniofacial development
Histone H4K20/H3K9 demethylase PHF8 regulates zebrafish brain and craniofacial development
PHF8 is a JmjC domain-containing protein, the gene for which has been linked to X-linked mental retardation (XLMR). These authors demonstrate PHF8 to be a histone demethylase with activity against H4K20me1. It has a role in regulating gene expression as well as in neuronal cell survival and craniofacial development in zebrafish. The results suggest there may be a link between histone methylation dynamics and XLMR.
- Hank H. Qi
- , Madathia Sarkissian
- & Yang Shi
-
Letter |
 PHF8 mediates histone H4 lysine 20 demethylation events involved in cell cycle progression
PHF8 mediates histone H4 lysine 20 demethylation events involved in cell cycle progression
These authors show that the JmjC domain-containing protein PHF8 has histone demethylase activity against H4K20me1 and is linked to two distinct events during cell cycle progression. PHF8 is recruited to the promoters of genes involved in the G1–S phase transition, where it removes H4K20me1 and contributes to gene activation, whereas dissociation of PHF8 from chromatin in prophase allows H4K20me1 to accumulate during mitosis.
- Wen Liu
- , Bogdan Tanasa
- & Michael G. Rosenfeld
-
Letter |
 A new DAF-16 isoform regulates longevity
A new DAF-16 isoform regulates longevity
The insulin/IGF-1 signalling (IIS) pathway is involved in various biological processes, including regulation of longevity. In the worm Caenorhabditis elegans, the transcription factor DAF-16a, one of two isoforms, has a major role in this pathway, regulating longevity, stress response and dauer diapause. These authors describe a new isoform, DAF-16d/f, which is also important in the regulation of lifespan. The DAF-16 isoforms functionally cooperate to fine-tune IIS-mediated processes in the context of a whole organism.
- Eun-Soo Kwon
- , Sri Devi Narasimhan
- & Heidi A. Tissenbaum
-
-
News & Views |
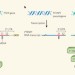 Decoy for microRNAs
Decoy for microRNAs
Pseudogenes are considered to be defunct relatives of known genes. But there is some surprising news: pseudogenes are functional and could have a role in the control of cancer
1 . Two experts discuss the significance of these findings for understanding the regulation of gene expression and cancer biology.- Isidore Rigoutsos
- & Frank Furnari
-
Letter |
 Small regulatory RNAs inhibit RNA polymerase II during the elongation phase of transcription
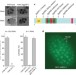
Small regulatory RNAs inhibit RNA polymerase II during the elongation phase of transcription
Small regulatory RNAs function both in the cytoplasm, inhibiting expression from messenger RNAs, and in the nucleus, silencing heterochromatin and preventing genome rearrangement. Now a new protein involved in RNA interference in the nucleus has been characterized. This protein, NRDE-2, associates with NRDE-3 and short interfering RNAs on nascent transcripts. This association prevents elongation of the transcripts by RNA polymerase II, making this a co-transcriptional form of gene silencing.
- Shouhong Guang
- , Aaron F. Bochner
- & Scott Kennedy
-
-
Letter |
 eIF5 has GDI activity necessary for translational control by eIF2 phosphorylation
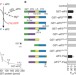
eIF5 has GDI activity necessary for translational control by eIF2 phosphorylation
The initiation of protein synthesis requires the eukaryotic translation initiation factor (eIF) 2, which uses energy from the hydrolysis of GTP. Another factor, eIF5, accelerates the GTP-hydrolysing activity of eIF2. Here, two other roles for eIF5 have been defined. One involves stabilizing GDP, the product of GTP hydrolysis, on eIF2. In its other role, eIF5 works with phosphorylated eIF2 to inhibit the guanine-nucleotide exchange factor eIF2B. These results clarify our understanding of how the initiation of translation is regulated.
- Martin D. Jennings
- & Graham D. Pavitt
-
News & Views |
 Breaking the second genetic code
Breaking the second genetic code
Diverse messenger RNAs, and thus proteins, can be generated from a single piece of DNA. A computational approach is helping to uncover complex combinatorial rules by which specific gene instructions are selected.
- J. Ramón Tejedor
- & Juan Valcárcel
-
Research Highlights |
Developmental biology: Hidden differences
-
Article |
 Adiponectin and AdipoR1 regulate PGC-1α and mitochondria by Ca2+ and AMPK/SIRT1
Adiponectin and AdipoR1 regulate PGC-1α and mitochondria by Ca2+ and AMPK/SIRT1
Adiponectin is a protein with anti-diabetic properties; its levels are decreased in obesity, insulin resistance and type 2 diabetes. It is shown here that mice with a muscle-specific disruption of adiponectin receptor 1 (AdipoR1) are insulin resistant and less able to endure exercise. The pathway downstream of receptor activation is delineated; the findings suggest that the decreased levels of adiponectin and AdipoR1 seen in obesity may have causal roles in the mitochondrial dysfunction and insulin resistance seen in diabetes.
- Masato Iwabu
- , Toshimasa Yamauchi
- & Takashi Kadowaki
-
Letter |
 NINJA connects the co-repressor TOPLESS to jasmonate signalling
NINJA connects the co-repressor TOPLESS to jasmonate signalling
In plants, the hormone jasmonoyl-isoleucine (JA-Ile) regulates growth, development and defence against pathogens. Proteins of the JAZ family repress JA-Ile-dependent gene expression, but the mechanism has been unclear. Here, an adaptor protein, NINJA, has been identified, which recruits co-repressor proteins that are known to mediate auxin-responsive gene expression as well. Hence these co-repressors are part of general repression complexes that are recruited to several different signalling pathways.
- Laurens Pauwels
- , Gemma Fernández Barbero
- & Alain Goossens
-
Letter |
 Identification of two evolutionarily conserved genes regulating processing of engulfed apoptotic cells
Identification of two evolutionarily conserved genes regulating processing of engulfed apoptotic cells
In multicellular organisms, apoptotic cells are removed from tissues by phagocytes, which recognize and engulf the dying cells. The molecular mechanisms underlying the subsequent degradation of the cells have been unclear. Here, two evolutionarily conserved genes have been identified that are required for such processing in Caenorhabditis elegans and mammals. An understanding of these events could lead to new treatments for diseases associated with poor engulfment and destruction of dying cells.
- Jason M. Kinchen
- & Kodi S. Ravichandran
-
-
Letter |
 Genetic analysis of variation in transcription factor binding in yeast
Genetic analysis of variation in transcription factor binding in yeast
Variation in the regulation of gene transcription between individuals is thought to be a major cause of phenotypic diversity. Here, individual differences in the binding of transcription-factor proteins are studied. A well-known transcription factor in the yeast pheromone pathway is used as an example, and the underlying genetic loci responsible for variation in its binding are mapped. The study reveals new insights into the mechanisms of gene regulation, and new regulators of the yeast pheromone pathway.
- Wei Zheng
- , Hongyu Zhao
- & Michael Snyder
-
Letter |
 Spatial control of EGF receptor activation by reversible dimerization on living cells
Spatial control of EGF receptor activation by reversible dimerization on living cells
Signalling through the epidermal growth factor receptor (EGFR) is preceded by its dimerization, which has typically been thought to occur through a ligand-induced conformational change. Here, the dimerization dynamics of individual EGFR molecules have been determined in living cells in real time, using a quantum-dot-based approach. Unliganded EGFR molecules undergo spontaneous and reversible dimerization; these pre-formed dimers are primed for ligand binding and signalling and are enriched at the cell periphery.
- Inhee Chung
- , Robert Akita
- & Ira Mellman
-
Letter |
 Transcription-independent ARF regulation in oncogenic stress-mediated p53 responses
Transcription-independent ARF regulation in oncogenic stress-mediated p53 responses
In response to oncogenic stress, the tumour suppressor ARF activates the p53 protein. ARF protein is highly stable in most human cell lines, so it has been thought that ARF activation occurs mainly at the level of transcription. Here, however, ARF is shown to be unstable in normal human cells but stable in cancer cells, through a transcription-independent mechanism. A ubiquitin ligase for ARF is identified and shown to promote ARF degradation in normal cells. This activity is prevented in cancer cells, stabilizing ARF.
- Delin Chen
- , Jing Shan
- & Wei Gu
-
Letter |
 SIRT3 regulates mitochondrial fatty-acid oxidation by reversible enzyme deacetylation
SIRT3 regulates mitochondrial fatty-acid oxidation by reversible enzyme deacetylation
During fasting SIRT3 is induced in liver and brown adipose tissue. One of SIRT3's substrates is shown to be long–chain acyl co-enzyme A dehydrogenase (LCAD). Without SIRT3 LCAD becomes hyperacetylated, which diminishes its activity, and reduces fatty acid oxidation. Mice without SIRT3 have all the hallmarks of fatty acid oxidation disorders during fasting, including reduced ATP levels and intolerance to cold. Thus, acetylation is a novel regulatory mechanism for fatty acid oxidation.
- Matthew D. Hirschey
- , Tadahiro Shimazu
- & Eric Verdin
-
Research Highlights |
Genetics: Male regulator switched
-
Research Highlights |
Neuroscience: Baby blues
-
News |
 Junk DNA holds clues to heart disease
Junk DNA holds clues to heart disease
Deleting a non-coding region leads to narrowing of arteries in mice.
- Janet Fang
-
News & Views |
 A chromatin thermostat
A chromatin thermostat
When environmental temperatures rise, plants seek help from their core molecular mechanisms to adapt. The chromatin protein H2A.Z, which regulates gene expression, is one such rescue molecule.
- Roger B. Deal
- & Steven Henikoff
-
Letter |
 FOXO-dependent regulation of innate immune homeostasis
FOXO-dependent regulation of innate immune homeostasis
Antimicrobial peptides (AMPs) are an important class of immune effector molecules which fight pathogen infections. AMP induction in Drosophila is regulated through the activation of the Toll and immune deficiency pathways; it is now shown that AMP activation can be achieved independently of these pathways by the transcription factor FOXO. In non-infected animals, AMP genes are activated in response to nuclear FOXO activity when induced by starvation.
- Thomas Becker
- , Gerrit Loch
- & Michael Hoch
-
Research Highlights |
 Molecular biology: Flowering time unravelled
Molecular biology: Flowering time unravelled

 The novel gene twenty-four defines a critical translational step in the Drosophila clock
The novel gene twenty-four defines a critical translational step in the Drosophila clock Extrinsic regulation of pluripotent stem cells
Extrinsic regulation of pluripotent stem cells