Featured
-
-
Article
| Open Access RAD51C-XRCC3 structure and cancer patient mutations define DNA replication roles
RAD51C-XRCC3 structure and cancer patient mutations define DNA replication roles
In this study, the authors present structures and functional analyses for the RAD51C-XRCC3 tumor suppressor complex, providing insights into recurrent mutations in cancer and Fanconi Anemia patients that uncover distinct DNA replication fork protection, restart and reversal regions.
- Michael A. Longo
- , Sunetra Roy
- & Katharina Schlacher
-
Article
| Open Access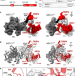 Extended DNA threading through a dual-engine motor module of the activating signal co-integrator 1 complex
Extended DNA threading through a dual-engine motor module of the activating signal co-integrator 1 complex
ASCC3 is a multi-functional helicase that contains two consecutive Ski2-like helicase units. Here, the authors show that ASCC3 can unwind DNA by threading one strand of a substrate duplex through both helicase units, supported by the TRIP4 protein.
- Junqiao Jia
- , Tarek Hilal
- & Markus C. Wahl
-
Article
| Open Access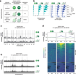 Unscheduled DNA replication in G1 causes genome instability and damage signatures indicative of replication collisions
Unscheduled DNA replication in G1 causes genome instability and damage signatures indicative of replication collisions
Reusswig et al. use engineered systems to force DNA replication in the G1 phase of the cell cycle. This unscheduled G1 replication shows hallmarks of S phase replication, but leads to over-replication and DNA breaks from replication collisions.
- Karl-Uwe Reusswig
- , Julia Bittmann
- & Boris Pfander
-
Article
| Open Access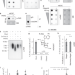 Mammalian N1-adenosine PARylation is a reversible DNA modification
Mammalian N1-adenosine PARylation is a reversible DNA modification
Poly-ADP-ribosylation (PARylation) is a well-known posttranslational modification of proteins. Here the authors show that beyond proteins also mammalian single-stranded DNA is PARylated in vitro and in vivo.
- Michael U. Musheev
- , Lars Schomacher
- & Christof Niehrs
-
Article
| Open Access Structural basis for APE1 processing DNA damage in the nucleosome
Structural basis for APE1 processing DNA damage in the nucleosome
AP endonuclease 1 (APE1) processes genomic AP sites during base excision repair. Here, the authors determine the structural mechanism used by APE1 to process nucleosomal AP sites, providing new insight into DNA repair in chromatin.
- Tyler M. Weaver
- , Nicole M. Hoitsma
- & Bret D. Freudenthal
-
Article
| Open Access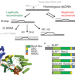 The toposiomerase IIIalpha-RMI1-RMI2 complex orients human Bloom’s syndrome helicase for efficient disruption of D-loops
The toposiomerase IIIalpha-RMI1-RMI2 complex orients human Bloom’s syndrome helicase for efficient disruption of D-loops
Human Bloom’s syndrome (BLM) helicase has a role in DNA repair, and BLM deficiency in humans is associated with chromosomal abnormalities. Here the authors employ solution biophysical assays to show BLM maintains a balance for disruption and stabilization of oligonucleotide-based D-loops. Interaction with the Topoisomerase IIIalpha-RMI1-RMI2 complex orients the activity toward D-loop disruption.
- Gábor M. Harami
- , János Pálinkás
- & Mihály Kovács
-
Article
| Open Access Structural evolution of a DNA repair self-resistance mechanism targeting genotoxic secondary metabolites
Structural evolution of a DNA repair self-resistance mechanism targeting genotoxic secondary metabolites
Microbial DNA glycosylases associated with the biosynthesis of DNA-damaging antibiotics have evolved self-resistance for their cognate natural products. Here, the authors provide evidence that cellular self-resistance is enabled by reduced affinity of the glycosylases for the excision products of the corresponding DNA lesions.
- Elwood A. Mullins
- , Jonathan Dorival
- & Brandt F. Eichman
-
Article
| Open Access Arginine methylation and ubiquitylation crosstalk controls DNA end-resection and homologous recombination repair
Arginine methylation and ubiquitylation crosstalk controls DNA end-resection and homologous recombination repair
Post-translational modifications are critical for regulating the DNA damage response. Here, the authors identify a methylation-deubiquitination crosstalk between methyltransferase PRMT1 and deubiquitinase USP11, showing that the enzymes regulate each other’s functions in DNA repair.
- Maria Pilar Sanchez-Bailon
- , Soo-Youn Choi
- & Clare C. Davies
-
Article
| Open Access CRISPR-Associated Primase-Polymerases are implicated in prokaryotic CRISPR-Cas adaptation
CRISPR-Associated Primase-Polymerases are implicated in prokaryotic CRISPR-Cas adaptation
CAPPs are putative Primase-Polymerases associated with CRISPR-Cas operons. Here, the authors show CAPPs genetic and physical association with Cas1 and Cas2, their capacity to function as DNA-dependent DNA primases and DNA polymerases, and that Cas1-Cas2 complex adjacent to CAPP has bona fide spacer integration activity.
- Katerina Zabrady
- , Matej Zabrady
- & Aidan J. Doherty
-
Article
| Open Access Histone Parylation factor 1 contributes to the inhibition of PARP1 by cancer drugs
Histone Parylation factor 1 contributes to the inhibition of PARP1 by cancer drugs
PARP1 is the target of clinically approved anti-cancer drugs whose in vivo efficacy has been hard to predict. Here the authors show how an altered active site formed between PARP1 and Histone PARylation Factor 1 (HPF1) changes the efficacy of some of these inhibitors.
- Johannes Rudolph
- , Genevieve Roberts
- & Karolin Luger
-
Article
| Open Access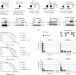 Warsaw Breakage Syndrome associated DDX11 helicase resolves G-quadruplex structures to support sister chromatid cohesion
Warsaw Breakage Syndrome associated DDX11 helicase resolves G-quadruplex structures to support sister chromatid cohesion
WABS patient derived cells display loss of sister chromatid cohesion. Here the authors by analyzing WABS patient derived cells, reveal a role of the DDX11 helicase in resolving G-Quadruplex structures to support sister chromatid cohesion.
- Janne J. M. van Schie
- , Atiq Faramarz
- & Job de Lange
-
Article
| Open Access Molecular determinants for dsDNA translocation by the transcription-repair coupling and evolvability factor Mfd
Molecular determinants for dsDNA translocation by the transcription-repair coupling and evolvability factor Mfd
Transcription-repair coupling factors (TRCFs) are large ATPases that mediate the preferential repair of the transcribed DNA strand. Here the authors reveal the cryo-EM structure of DNA-bound Mfd, the bacterial TRCF, and provide molecular insights into its mode of action.
- Christiane Brugger
- , Cheng Zhang
- & Alexandra M. Deaconescu
-
Article
| Open Access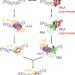 Single-molecule live-cell imaging visualizes parallel pathways of prokaryotic nucleotide excision repair
Single-molecule live-cell imaging visualizes parallel pathways of prokaryotic nucleotide excision repair
In Escherichia coli, the UvrAB damage sensor recognizes helix-distorting lesions by itself or via Mfd bound to stalled RNA polymerase. Here authors use single-molecule fluorescence imaging to quantify the kinetic signatures of interactions of UvrA with Mfd and UvrB in live cells.
- Harshad Ghodke
- , Han Ngoc Ho
- & Antoine M. van Oijen
-
Article
| Open Access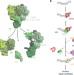 Single-molecule imaging reveals molecular coupling between transcription and DNA repair machinery in live cells
Single-molecule imaging reveals molecular coupling between transcription and DNA repair machinery in live cells
Mfd recognizes stalled transcriptional complexes at sites of lesions and recruits the nucleotide excision repair proteins (UvrAB) to the site. Here the authors use live cell imaging in E. coli to demonstrate that coordinated ATP hydrolysis by UvrA and loading of UvrB on DNA facilitate the dissociation of Mfd from the handoff complex.
- Han Ngoc Ho
- , Antoine M. van Oijen
- & Harshad Ghodke
-
Article
| Open Access PARP1 exhibits enhanced association and catalytic efficiency with γH2A.X-nucleosome
PARP1 exhibits enhanced association and catalytic efficiency with γH2A.X-nucleosome
The poly(ADP-ribose) polymerases play a key role in maintaining genomic integrity by detecting DNA damage and mediating repair. Here the authors characterize the kinetics of PARP1 binding to a variety of nucleosomes harbouring DNA double-strand breaks.
- Deepti Sharma
- , Louis De Falco
- & Curt A. Davey
-
Article
| Open Access Functional analysis of genetic variants in the high-risk breast cancer susceptibility gene PALB2
Functional analysis of genetic variants in the high-risk breast cancer susceptibility gene PALB2
PALB2 is an established breast cancer risk gene but the pathogenicity of many variants remains uncharacterised. Here, the authors present a cDNA-based system for the functional analysis of PALB2 variants of unknown significance.
- Rick A. C. M. Boonen
- , Amélie Rodrigue
- & Haico van Attikum
-
Article
| Open Access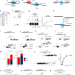 Molecular basis of microhomology-mediated end-joining by purified full-length Polθ
Molecular basis of microhomology-mediated end-joining by purified full-length Polθ
DNA polymerase θ is a polymerase-helicase essential for microhomology-mediated end-joining (MMEJ) or alternative end-joining of DNA. Here the authors use biochemical and biophysical methods to reveal how full-length human DNA polymerase θ performs MMEJ at the molecular level.
- Samuel J. Black
- , Ahmet Y. Ozdemir
- & Richard T. Pomerantz
-
Article
| Open Access Pol μ dGTP mismatch insertion opposite T coupled with ligation reveals promutagenic DNA repair intermediate
Pol μ dGTP mismatch insertion opposite T coupled with ligation reveals promutagenic DNA repair intermediate
Incorporation of mismatched nucleotides during DNA replication or repair can lead to mutagenesis. Here the authors reveal that DNA ligase can ligate NHEJ intermediates following incorporation of 8-oxodGTP or dGTP opposite T by DNA Polymerase mu (Pol mu) in vitro, which suggests that Pol mu could cause promutagenic mismatches during DSB repair.
- Melike Çağlayan
- & Samuel H. Wilson
-
Article
| Open Access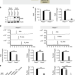 PARP2 mediates branched poly ADP-ribosylation in response to DNA damage
PARP2 mediates branched poly ADP-ribosylation in response to DNA damage
PARP1 and PARP2 of the PARP family enzymes are involved in DNA damage response. Here the authors report PARP2 activation mechanisms and its role in the formation of branched poly(ADP-ribose) chains in response to DNA damage.
- Qian Chen
- , Muzaffer Ahmad Kassab
- & Xiaochun Yu
-
Article
| Open Access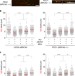 MRE11 and EXO1 nucleases degrade reversed forks and elicit MUS81-dependent fork rescue in BRCA2-deficient cells
MRE11 and EXO1 nucleases degrade reversed forks and elicit MUS81-dependent fork rescue in BRCA2-deficient cells
BRCA proteins have emerged as key stabilizing factors for the maintenance of replication forks following replication stress. Here the authors describe how reversed replication forks are degraded in the absence of BRCA2, and a MUS81 and POLD3-dependent mechanism of rescue following the withdrawal of genotoxic agent.
- Delphine Lemaçon
- , Jessica Jackson
- & Alessandro Vindigni
-
Article
| Open Access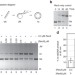 The cohesin-like RecN protein stimulates RecA-mediated recombinational repair of DNA double-strand breaks
The cohesin-like RecN protein stimulates RecA-mediated recombinational repair of DNA double-strand breaks
RecN is a cohesin-like molecule involved in the bacterial response to DNA double-strand breaks. Here the authors provide evidence that RecN stimulates the DNA strand invasion step of RecA-mediated recombination.
- Lee A. Uranga
- , Emigdio D. Reyes
- & Shelley L. Lusetti
-
Article
| Open Access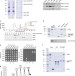 Identification and characterization of a heterotrimeric archaeal DNA polymerase holoenzyme
Identification and characterization of a heterotrimeric archaeal DNA polymerase holoenzyme
The current model for B-family DNA polymerases in archaea is one of single-subunit enzymes in contrast to the multi-subunit complexes in eukaryotes. Here the authors show that PolB1 fromSulfolobus solfataricusexists as a heterotrimeric complex in cell extracts.
- Jiangyu Yan
- , Thomas R. Beattie
- & Stephen D. Bell
-
Article
| Open Access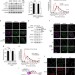 Coordinated nuclease activities counteract Ku at single-ended DNA double-strand breaks
Coordinated nuclease activities counteract Ku at single-ended DNA double-strand breaks
Homologous recombination requires end resection of the DNA at the site of the break, however the Ku dimer can sequester single-ended double-strand breaks. Here the authors show that ATM-dependent phosphorylation of CtIP, along with the actions of Mre11, impair the stable loading of Ku onto DNA.
- Pauline Chanut
- , Sébastien Britton
- & Patrick Calsou
-
Article
| Open Access RNF168 and USP10 regulate topoisomerase IIα function via opposing effects on its ubiquitylation
RNF168 and USP10 regulate topoisomerase IIα function via opposing effects on its ubiquitylation
The E3 ligase RNF168 is essential for the signalling of DNA double strand break and its mutations are associated with the RIDDLE syndrome. Here the authors identify TOP2a as substrate for RNF168 and USP10; providing a link between the RNF168/USP10 axis, TOP2a and the response to anti-cancer drugs that target TOP2.
- Kiran Kumar Naidu Guturi
- , Miyuki Bohgaki
- & Razqallah Hakem
-
Article
| Open Access PARP3 is a sensor of nicked nucleosomes and monoribosylates histone H2BGlu2
PARP3 is a sensor of nicked nucleosomes and monoribosylates histone H2BGlu2
Chromosomal single-strand DNA breaks occur frequently and require repair to avoid disease outcomes. Here, the authors show that in bird cells, PARP3 accelerates this repair, and use structural biology and cell biology techniques to reveal details of the mechanism of action.
- Gabrielle J. Grundy
- , Luis M. Polo
- & Keith W. Caldecott
-
Article
| Open Access Impact of ribonucleotide incorporation by DNA polymerases β and λ on oxidative base excision repair
Impact of ribonucleotide incorporation by DNA polymerases β and λ on oxidative base excision repair
Oxidative stress is a common source of DNA damage and is repaired by the base excision repair machinery, including polymerase beta. Here the authors find that polymerase beta, and to a lesser extent lambda, can mistakenly incorporate ribonucleotides during synthesis.
- Emmanuele Crespan
- , Antonia Furrer
- & Giovanni Maga
-
Article
| Open Access Wnt/β-catenin pathway regulates MGMT gene expression in cancer and inhibition of Wnt signalling prevents chemoresistance
Wnt/β-catenin pathway regulates MGMT gene expression in cancer and inhibition of Wnt signalling prevents chemoresistance
The high expression of the DNA repair enzyme O6-methylguanine DNA methyltransferase (MGMT) often confers resistance to chemotherapy in several cancers. In this study, the authors propose the inhibition of the Wnt signalling pathway as an alternative strategy to modulate MGMT expression and sensitize tumours to chemotherapy.
- Malin Wickström
- , Cecilia Dyberg
- & John Inge Johnsen
-
Article
| Open Access Crystal structure, biochemical and cellular activities demonstrate separate functions of MTH1 and MTH2
Crystal structure, biochemical and cellular activities demonstrate separate functions of MTH1 and MTH2
Dysfunctional redox regulation in cancer can damage dNTPs so inhibiting dNTP pool sanitizing enzymes, such as MTH1, is a potential cancer treatment. Here, Carter et al.characterize MTH2 (NUDT15) and show that it is not a dNTP sanitizer, and so is unlikely to influence the efficacy of MTH1 inhibitors.
- Megan Carter
- , Ann-Sofie Jemth
- & Pål Stenmark
-
Article |
 The molecular origin of high DNA-repair efficiency by photolyase
The molecular origin of high DNA-repair efficiency by photolyase
Photolyase is an enzyme responsible for repairing DNA which is damaged after exposure to UV light. Here, the authors use site directed mutagenesis and femtosecond spectroscopy to study how photolyase achieves its maximal repair efficiency.
- Chuang Tan
- , Zheyun Liu
- & Dongping Zhong
-
Article |
 The DNA repair endonuclease Mus81 facilitates fast DNA replication in the absence of exogenous damage
The DNA repair endonuclease Mus81 facilitates fast DNA replication in the absence of exogenous damage
Several mechanisms are in place to ensure the accurate and timely replication of the genome as cells progress through S-phase. Here, the authors show that Mus81, an endonuclease involved in the response to DNA damage during replicative stress, also regulates the rate of DNA replication during normal growth.
- Haiqing Fu
- , Melvenia M. Martin
- & Mirit I. Aladjem
-
Article |
 DNA repair choice defines a common pathway for recruitment of chromatin regulators
DNA repair choice defines a common pathway for recruitment of chromatin regulators
Chromatin regulators facilitate repair of DNA double-strand breaks in chromosomal DNA. The authors show that the recruitment of such chromatin regulators to DNA lesions is controlled by the choice of DNA repair pathway.
- Gwendolyn Bennett
- , Manolis Papamichos-Chronakis
- & Craig L. Peterson
-
Article |
 ALKBH8-mediated formation of a novel diastereomeric pair of wobble nucleosides in mammalian tRNA
ALKBH8-mediated formation of a novel diastereomeric pair of wobble nucleosides in mammalian tRNA
Uridines at the wobble position of transfer RNA anticodons are usually modified to allow efficient decoding of messenger RNA codons. In this study, ALKBH8 is shown to be a bifunctional transfer RNA modification enzyme required for the formation of a novel diastereomeric pair of modified wobble uridines.
- Erwin van den Born
- , Cathrine B. Vågbø
- & Pål Ø. Falnes

 PARP2 promotes Break Induced Replication-mediated telomere fragility in response to replication stress
PARP2 promotes Break Induced Replication-mediated telomere fragility in response to replication stress