Article
|
Open Access
Featured
-
-
Article
| Open Access Network enhancement as a general method to denoise weighted biological networks
Network enhancement as a general method to denoise weighted biological networks
Technical noise in experiments is unavoidable, but it introduces inaccuracies into the biological networks we infer from the data. Here, the authors introduce a diffusion-based method for denoising undirected, weighted networks, and show that it improves the performances of downstream analyses.
- Bo Wang
- , Armin Pourshafeie
- & Jure Leskovec
-
Article
| Open Access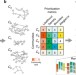 Prioritizing network communities
Prioritizing network communities
Community detection allows one to decompose a network into its building blocks. While communities can be identified with a variety of methods, their relative importance can’t be easily derived. Here the authors introduce an algorithm to identify modules which are most promising for further analysis.
- Marinka Zitnik
- , Rok Sosič
- & Jure Leskovec
-
Article
| Open Access Analysis of sensitive information leakage in functional genomics signal profiles through genomic deletions
Analysis of sensitive information leakage in functional genomics signal profiles through genomic deletions
Functional genomics data from many studies are widely shared publicly for their value in biomedical and disease research. Here, the authors show sensitive information leakage is possible by analyzing functional genomics signal profiles, and develop an anonymization procedure for privacy protection.
- Arif Harmanci
- & Mark Gerstein
-
Article
| Open Access Multi-omics profiling of younger Asian breast cancers reveals distinctive molecular signatures
Multi-omics profiling of younger Asian breast cancers reveals distinctive molecular signatures
While breast cancer incidence in the Asia Pacific region is rising, the molecular basis remains poorly characterized. Here the authors perform genomic screening of 187 Korean breast cancer patients and find differences in molecular subtype distribution, mutation pattern and prevalence, and gene expression signature when compared to TCGA.
- Zhengyan Kan
- , Ying Ding
- & Yeon Hee Park
-
Article
| Open Access A comprehensive evaluation of module detection methods for gene expression data
A comprehensive evaluation of module detection methods for gene expression data
Modules composed of groups of genes with similar expression profiles tend to be functionally related and co-regulated. Here, Saelens et al evaluate the performance of 42 computational methods and provide practical guidelines for module detection in gene expression data.
- Wouter Saelens
- , Robrecht Cannoodt
- & Yvan Saeys
-
Article
| Open Access Exploiting genetic variation to uncover rules of transcription factor binding and chromatin accessibility
Exploiting genetic variation to uncover rules of transcription factor binding and chromatin accessibility
Single nucleotide variants (SNVs) can affect chromatin occupancy by transcription factors (TF). Here the authors mine naturally occurring SNVs to probe cis elements and define contextual sequences that govern in vivo transcription factor chromatin occupancy and chromatin accessibility.
- Vivek Behera
- , Perry Evans
- & Gerd A. Blobel
-
Article
| Open Access The effects of death and post-mortem cold ischemia on human tissue transcriptomes
The effects of death and post-mortem cold ischemia on human tissue transcriptomes
RNA levels in post-mortem tissue can differ greatly from those before death. Studying the effect of post-mortem interval on the transcriptome in 36 human tissues, Ferreira et al. find that the response to death is largely tissue-specific and develop a model to predict time since death based on RNA data.
- Pedro G. Ferreira
- , Manuel Muñoz-Aguirre
- & Roderic Guigó
-
Article
| Open Access Subcortical evidence for a contribution of arousal to fMRI studies of brain activity
Subcortical evidence for a contribution of arousal to fMRI studies of brain activity
Resting cortical activity fluctuates, but it is unclear what underlies these variations in activity. Here, the authors show that large-scale fluctuations in fMRI cortical activity are associated with momentary decreases in cortical arousal and opposite activity changes in the basal forebrain and thalamus.
- Xiao Liu
- , Jacco A. de Zwart
- & Jeff H. Duyn
-
Article
| Open Access PAX7 target genes are globally repressed in facioscapulohumeral muscular dystrophy skeletal muscle
PAX7 target genes are globally repressed in facioscapulohumeral muscular dystrophy skeletal muscle
Facioscapulohumeral muscular dystrophy is a myopathy linked to ectopic expression of the DUX4 transcription factor. The authors show that the suppression of targets genes of the myogenesis regulator PAX7 is a signature of FSHD, and might explain oxidative stress sensitivity and epigenetic changes.
- Christopher R. S. Banerji
- , Maryna Panamarova
- & Peter S. Zammit
-
Article
| Open Access NSD1- and NSD2-damaging mutations define a subset of laryngeal tumors with favorable prognosis
NSD1- and NSD2-damaging mutations define a subset of laryngeal tumors with favorable prognosis
The authors use an integrative clustering approach to identify two laryngeal cancer clusters with distinct prognosis and show that mutations damaging the NSD1 and NSD2 methyltransferases segregate to the cluster with favorable prognosis, and independently predict longer survival in patients with laryngeal, but not other head and neck cancers.
- Suraj Peri
- , Evgeny Izumchenko
- & Erica A. Golemis
-
Article
| Open Access Mutational signatures reveal the dynamic interplay of risk factors and cellular processes during liver tumorigenesis
Mutational signatures reveal the dynamic interplay of risk factors and cellular processes during liver tumorigenesis
Tumorigenesis is a complex process driven by numerous risk factors. Here, genomic analysis of liver cancer reveals the evolution of mutational signatures during tumor development, highlighting mutational and structural signatures linked to environmental exposures and endogenous cellular processes.
- Eric Letouzé
- , Jayendra Shinde
- & Jessica Zucman-Rossi
-
Article
| Open Access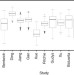 On the performance of pre-microRNA detection algorithms
On the performance of pre-microRNA detection algorithms
As the experimental discovery of microRNAs (miRNAs) is cumbersome, computational tools have been developed for the prediction of pre-miRNAs. Here the authors develop a framework to assess the performance of existing and novel pre-miRNA prediction tools and provide guidelines for selecting an appropriate approach for a given data set.
- Müşerref Duygu Saçar Demirci
- , Jan Baumbach
- & Jens Allmer
-
Article
| Open Access A pan-cancer genome-wide analysis reveals tumour dependencies by induction of nonsense-mediated decay
A pan-cancer genome-wide analysis reveals tumour dependencies by induction of nonsense-mediated decay
Nonsense-mediated decay (NMD) eliminates transcripts with premature stop codons and has been linked to cancer genesis. Here, the authors develop an algorithm to predict NMD and perform a pan-cancer analysis that finds that some hypermutated cancers are dependent on mutations that elicit NMD.
- Zhiyuan Hu
- , Christopher Yau
- & Ahmed Ashour Ahmed
-
Article
| Open Access Reconstructing cell cycle pseudo time-series via single-cell transcriptome data
Reconstructing cell cycle pseudo time-series via single-cell transcriptome data
In single-cell RNA sequencing data of heterogeneous cell populations, cell cycle stage of individual cells would often be informative. Here, the authors introduce a computational model to reconstruct a pseudo-time series from single cell transcriptome data, identify the cell cycle stages, identify candidate cell cycle-regulated genes and recover the methylome changes during the cell cycle.
- Zehua Liu
- , Huazhe Lou
- & Ting Chen
-
Article
| Open Access Systematic discovery of mutation-specific synthetic lethals by mining pan-cancer human primary tumor data
Systematic discovery of mutation-specific synthetic lethals by mining pan-cancer human primary tumor data
There are no robust methods for systematically identifying mutation-specific synthetic lethal (SL) partners in cancer. Here, the authors develop a computational algorithm that uses pan-cancer data to detect mutation-andcancer-specific SL partners and they validate a novel SL interaction between mutant IDH and loss of ACACA in leukaemia.
- Subarna Sinha
- , Daniel Thomas
- & David L. Dill
-
Article
| Open Access Analysis of renal cancer cell lines from two major resources enables genomics-guided cell line selection
Analysis of renal cancer cell lines from two major resources enables genomics-guided cell line selection
Cell lines are central to cancer research, but knowing which cell lines are the best representative of actual tumours is a major challenge. Here the authors provide a resource assessment of 65 renal cell lines to assist researchers in selecting suitable lines for studying specific renal carcinoma subtypes.
- Rileen Sinha
- , Andrew G. Winer
- & A. Ari Hakimi

 Predicting the evolution of Escherichia coli by a data-driven approach
Predicting the evolution of Escherichia coli by a data-driven approach