Featured
-
-
Article
| Open Access Proteogenomics of clear cell renal cell carcinoma response to tyrosine kinase inhibitor
Proteogenomics of clear cell renal cell carcinoma response to tyrosine kinase inhibitor
Many clear cell renal cell carcinoma (ccRCC) patients do not respond or develop resistance to tyrosine kinase inhibitors, such as Sunitinib. Here, the authors perform a proteogenomics analysis of Chinese ccRCC patients treated with Sunitinib and develop a multi-omics classifier to distinguish responders from non-responders.
- Hailiang Zhang
- , Lin Bai
- & Chen Ding
-
Article
| Open Access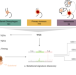 Mutational signature dynamics shaping the evolution of oesophageal adenocarcinoma
Mutational signature dynamics shaping the evolution of oesophageal adenocarcinoma
It is critical to understand what drives the progression of oesophageal adenocarcinoma (OAC) from a pre-cancerous state. Here, the authors use whole-genome sequencing to characterise the mutational processes and drivers of OAC progression from Barrett’s Oesophagus, as well as their prognostic associations.
- Sujath Abbas
- , Oriol Pich
- & Maria Secrier
-
Article
| Open Access The proteomic landscape of soft tissue sarcomas
The proteomic landscape of soft tissue sarcomas
Characterising the molecular profile of soft tissue sarcomas (STS) remains critical. Here, the authors analyse samples from 321 STS patients across 11 histological subtypes using proteomics and identify prognostic signatures that can be applied to multiple subtypes.
- Jessica Burns
- , Christopher P. Wilding
- & Paul H. Huang
-
Article
| Open Access The transcriptional landscape and diagnostic potential of long non-coding RNAs in esophageal squamous cell carcinoma
The transcriptional landscape and diagnostic potential of long non-coding RNAs in esophageal squamous cell carcinoma
Esophageal squamous cell carcinoma is difficult to detect at early stages, and late detection is often linked to poor prognosis. Here, the authors develop a lncRNA signature predictive of ESCC and validate across multiple external cohorts.
- Meng Zhou
- , Siqi Bao
- & Zhihua Liu
-
Article
| Open Access Systems-level analyses of protein-protein interaction network dysfunctions via epichaperomics identify cancer-specific mechanisms of stress adaptation
Systems-level analyses of protein-protein interaction network dysfunctions via epichaperomics identify cancer-specific mechanisms of stress adaptation
Epichaperomics allow the study of protein-protein interactions and their alterations, but probes have been limited to capturing HSP90 epichaperomes. Here, the authors introduce and validate a toolset of HSP70 epichaperome ligands, and use them in epichaperomics to identify a mechanism with which cancer cells can enhance the fitness of mitotic protein networks.
- Anna Rodina
- , Chao Xu
- & Gabriela Chiosis
-
Article
| Open Access Oncogenic structural aberration landscape in gastric cancer genomes
Oncogenic structural aberration landscape in gastric cancer genomes
Gastric cancers (GC) are driven by genomic alterations, but the underlying molecular mechanisms remain unclear. Here, the authors analyse the structural rearrangement landscape of 170 GCs using whole-genome sequencing, identify recurrent structural variant hotspots and find oncogene amplicons driven by extrachromosomal DNA.
- Mihoko Saito-Adachi
- , Natsuko Hama
- & Tatsuhiro Shibata
-
Article
| Open Access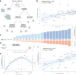 Cancer genomes tolerate deleterious coding mutations through somatic copy number amplifications of wild-type regions
Cancer genomes tolerate deleterious coding mutations through somatic copy number amplifications of wild-type regions
Most of the mutations accumulated in cancer cells are deleterious, and it is unclear how such alterations are tolerated. Here, the authors propose that copy number amplifications could increase the tolerance to deleterious mutations, and analyse the features that could determine the underlying selection process.
- Fabio Alfieri
- , Giulio Caravagna
- & Martin H. Schaefer
-
Article
| Open Access Tumour mutations in long noncoding RNAs enhance cell fitness
Tumour mutations in long noncoding RNAs enhance cell fitness
The role of mutations within long noncoding RNAs (lncRNAs) exons on tumour cell fitness remains to be explored. Here, the authors investigate the landscape of driver lncRNAs in primary and metastatic samples and validate the functional significance of single nucleotide variants in the NEAT1 oncogene in vitro and in vivo.
- Roberta Esposito
- , Andrés Lanzós
- & Rory Johnson
-
Article
| Open Access Multi-omic features of oesophageal adenocarcinoma in patients treated with preoperative neoadjuvant therapy
Multi-omic features of oesophageal adenocarcinoma in patients treated with preoperative neoadjuvant therapy
It remains critical to understand the genomic events in response to treatment of oesophageal adenocarcinoma (OAC). Here, the authors perform a multi-omics analysis of OAC patients from the DOCTOR phase II clinical trial, finding genomic features and immune clusters associated with survival.
- Marjan M. Naeini
- , Felicity Newell
- & Nicola Waddell
-
Article
| Open Access Single-cell transcriptomics reveals immune suppression and cell states predictive of patient outcomes in rhabdomyosarcoma
Single-cell transcriptomics reveals immune suppression and cell states predictive of patient outcomes in rhabdomyosarcoma
The cellular differentiation states of paediatric rhabdomyosarcoma (RMS) remain to be explored. Here, single-cell RNA sequencing analysis of RMS tumours reveals an immunosuppressive microenvironment and distinct transcriptional programs predictive of patient outcomes.
- Jeff DeMartino
- , Michael T. Meister
- & Jarno Drost
-
Article
| Open Access macroH2A2 antagonizes epigenetic programs of stemness in glioblastoma
macroH2A2 antagonizes epigenetic programs of stemness in glioblastoma
Self-renewing cells play an important role in initiation, progression, and therapy resistance in glioblastoma. Here, the authors identify histone variant macroH2A2 as a regulator of chromatin organisation resulting in the suppression of transcriptional programs of self-renewal in glioblastoma.
- Ana Nikolic
- , Francesca Maule
- & Marco Gallo
-
Article
| Open Access Detecting recurrent passenger mutations in melanoma by targeted UV damage sequencing
Detecting recurrent passenger mutations in melanoma by targeted UV damage sequencing
Genome sequencing has identified many recurrent mutations in melanoma. Here, we use targeted UV damage sequencing to show that many of these mutations are associated with UV damage hotspots that are linked to DNA binding by ETS transcription factors.
- Kathiresan Selvam
- , Smitha Sivapragasam
- & John J. Wyrick
-
Article
| Open Access Multiplatform molecular profiling uncovers two subgroups of malignant peripheral nerve sheath tumors with distinct therapeutic vulnerabilities
Multiplatform molecular profiling uncovers two subgroups of malignant peripheral nerve sheath tumors with distinct therapeutic vulnerabilities
Malignant peripheral nerve sheath tumours are an aggressive form of sarcoma, with limited treatment options. Here, the authors utilise DNA methylation and transcriptomic data to identify two subtypes of tumours with potential therapeutic vulnerabilities.
- Suganth Suppiah
- , Sheila Mansouri
- & Gelareh Zadeh
-
Article
| Open Access Integrated genomic analysis reveals aberrations in WNT signaling in germ cell tumors of childhood and adolescence
Integrated genomic analysis reveals aberrations in WNT signaling in germ cell tumors of childhood and adolescence
Genomic landscape studies of malignant germ cell tumors (GCTs) that occur in children, adolescents and young adults are limited. Here the authors perform multi-omics profiling of different types of GCTs across the age spectrum from 0–24 years and show that WNT signalling pathway is activated in GCTs and is associated with poor clinical outcomes.
- Lin Xu
- , Joshua L. Pierce
- & James F. Amatruda
-
Article
| Open Access Circulating tumor DNA reveals mechanisms of lorlatinib resistance in patients with relapsed/refractory ALK-driven neuroblastoma
Circulating tumor DNA reveals mechanisms of lorlatinib resistance in patients with relapsed/refractory ALK-driven neuroblastoma
Inhibition of ALK is initially effective in patients with ALK-driven lung cancer but resistance often arises. Here, the authors use circulating tumour DNA, collected as part of a phase I trial investigating lorlatinib (ALK inhibitor) in pediatric patients with ALK-driven neuroblastoma, to detect early resistance mechanisms.
- Esther R. Berko
- , Gabriela M. Witek
- & Yaël P. Mossé
-
Article
| Open Access Reversible transitions between noradrenergic and mesenchymal tumor identities define cell plasticity in neuroblastoma
Reversible transitions between noradrenergic and mesenchymal tumor identities define cell plasticity in neuroblastoma
Noradrenergic and mesenchymal cell states have been proposed in neuroblastoma, but their contributions to the tumour are not clearly understood. Here, the authors used in vitro and in vivo models, as well as single-cell RNA-seq, to characterise noradrenergic and mesenchymal cells and their phenotypic plasticity in neuroblastoma.
- Cécile Thirant
- , Agathe Peltier
- & Isabelle Janoueix-Lerosey
-
Article
| Open Access Re-convolving the compositional landscape of primary and recurrent glioblastoma reveals prognostic and targetable tissue states
Re-convolving the compositional landscape of primary and recurrent glioblastoma reveals prognostic and targetable tissue states
Glioblastoma (GBM) cells can infiltrate into the tumour microenvironment (TME) and contribute to recurrence. Here, the authors analyse primary and recurrent GBMs and their TME using single-nucleus and spatial transcriptomics, revealing tissue states defined by the combinations of neoplastic and non-neoplastic cells, which could be therapeutic targets.
- Osama Al-Dalahmah
- , Michael G. Argenziano
- & Peter Canoll
-
Article
| Open Access Genomic mutation landscape of skin cancers from DNA repair-deficient xeroderma pigmentosum patients
Genomic mutation landscape of skin cancers from DNA repair-deficient xeroderma pigmentosum patients
Xeroderma pigmentosum (XP) is a rare genetic disorder that is associated with a higher risk of skin cancer. Here, the authors analyse the genomes of skin cancers from patients across five different XP groups, revealing genetic and molecular factors related to the mutational profile and UV-related mutagenesis in XP.
- Andrey A. Yurchenko
- , Fatemeh Rajabi
- & Sergey I. Nikolaev
-
Article
| Open Access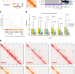 3D genome mapping identifies subgroup-specific chromosome conformations and tumor-dependency genes in ependymoma
3D genome mapping identifies subgroup-specific chromosome conformations and tumor-dependency genes in ependymoma
Ependymoma is a tumor of the brain or spinal cord with the two most common and aggressive types mainly occurring in children. Here the authors employ 3D genomics and epigenomics to reveal targets for aggressive ependymoma tumors in children.
- Konstantin Okonechnikov
- , Aylin Camgöz
- & Lukas Chavez
-
Article
| Open Access Bladder cancer organoids as a functional system to model different disease stages and therapy response
Bladder cancer organoids as a functional system to model different disease stages and therapy response
Bladder cancer heterogeneity can limit treatment efficacy in individual patients. Here, the authors use patient derived organoids to develop a drug screening pipeline and identify markers of treatment response.
- Martina Minoli
- , Thomas Cantore
- & Marianna Kruithof-de Julio
-
Article
| Open Access Integrative modeling of tumor genomes and epigenomes for enhanced cancer diagnosis by cell-free DNA
Integrative modeling of tumor genomes and epigenomes for enhanced cancer diagnosis by cell-free DNA
Despite advances in ctDNA cancer detection, early detection remains difficult. Here, the authors utilise whole genome sequencing of 2,125 patient samples to create a model for early cancer and tissue of origin detection.
- Mingyun Bae
- , Gyuhee Kim
- & Jung Kyoon Choi
-
Article
| Open Access Predicting response to enzalutamide and abiraterone in metastatic prostate cancer using whole-omics machine learning
Predicting response to enzalutamide and abiraterone in metastatic prostate cancer using whole-omics machine learning
Prostate cancer is known to have a variable response to androgen receptor signalling inhibitors. Here, the authors use machine learning to predict response to therapy from genomic, transcriptomic and clinical data.
- Anouk C. de Jong
- , Alexandra Danyi
- & Martijn P. Lolkema
-
Article
| Open Access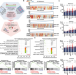 Targeting C/EBPα overcomes primary resistance and improves the efficacy of FLT3 inhibitors in acute myeloid leukaemia
Targeting C/EBPα overcomes primary resistance and improves the efficacy of FLT3 inhibitors in acute myeloid leukaemia
Resistance of FLT3-ITD acute myeloid leukaemia (AML) patients to FLT3 inhibitors (FLT3i) remains an urgent clinical challenge. Here, the authors identify C/EBPα activation as a mechanism of FLT3i resistance and therapeutically target C/EBPα activation in combination with FLT3i in preclinical models FLT3-ITD AML.
- Hanlin Wang
- , Guanghao Luo
- & Jia Li
-
Article
| Open Access Etiology of oncogenic fusions in 5,190 childhood cancers and its clinical and therapeutic implication
Etiology of oncogenic fusions in 5,190 childhood cancers and its clinical and therapeutic implication
Oncogenic gene fusions are frequent in childhood cancers but remain poorly understood and untargeted. Here, the authors identify 272 oncogenic fusions in transcriptomics data from 5190 childhood cancer patients, revealing their possible etiologies, their links with tumor progression and evolution, and their potential as therapeutic targets.
- Yanling Liu
- , Jonathon Klein
- & Xiaotu Ma
-
Article
| Open Access Comprehensive proteogenomic characterization of early duodenal cancer reveals the carcinogenesis tracks of different subtypes
Comprehensive proteogenomic characterization of early duodenal cancer reveals the carcinogenesis tracks of different subtypes
Duodenal cancer (DC) has complex subtypes and undergoes complicated morphological changes throughout progression, so understanding the molecular basis is crucial. Here, the authors perform a proteogenomics analysis of 156 DCs, revealing molecular subtypes as well as the roles of smoking, AARS1 and PARP1.
- Lingling Li
- , Dongxian Jiang
- & Chen Ding
-
Article
| Open Access Epigenetic and transcriptomic characterization reveals progression markers and essential pathways in clear cell renal cell carcinoma
Epigenetic and transcriptomic characterization reveals progression markers and essential pathways in clear cell renal cell carcinoma
Tumour heterogeneity in clear cell renal cell carcinoma (ccRCC) remains to be investigated. Here, the integration of spatial omics, transcriptional and chromatin accessibility profiling at the single-nucleus level and bulk proteogenomics data reveal markers and pathways important for ccRCC.
- Yige Wu
- , Nadezhda V. Terekhanova
- & Feng Chen
-
Article
| Open Access Integrative proteogenomic characterization of early esophageal cancer
Integrative proteogenomic characterization of early esophageal cancer
The progression of oesophageal squamous cell carcinoma (ESCC) from early to advanced stages requires comprehensive molecular characterisation. Here, the authors perform a proteogenomics analysis of ESCC patient samples across nine histopathological stages and three phases, identifying key alterations and paths for progression.
- Lingling Li
- , Dongxian Jiang
- & Chen Ding
-
Article
| Open Access Clonal origin and development of high hyperdiploidy in childhood acute lymphoblastic leukaemia
Clonal origin and development of high hyperdiploidy in childhood acute lymphoblastic leukaemia
High hyperdiploid acute lymphoblastic leukaemia (HeH ALL) is driven by nonrandom chromosomal gains, which have been suggested to arise early - even before birth. Here, the authors use single-cell whole genome sequencing and in silico modelling to show that HeH ALL aneuploidies could originate early and follow punctuated evolution.
- Eleanor L. Woodward
- , Minjun Yang
- & Kajsa Paulsson
-
Article
| Open Access Genomic characterization of DICER1-associated neoplasms uncovers molecular classes
Genomic characterization of DICER1-associated neoplasms uncovers molecular classes
DICER1 syndrome is associated with a predisposition to multiple tumor types. Here, the authors identify and characterize 3 molecular subgroups of mesenchymal tumors with DICER1 mutations.
- Felix K. F. Kommoss
- , Anne-Sophie Chong
- & William D. Foulkes
-
Article
| Open Access Pan-cancer classification of single cells in the tumour microenvironment
Pan-cancer classification of single cells in the tumour microenvironment
The accuracy and granularity of classifying cell types in the tumour microenvironment (TME) from single-cell RNA-seq data is impacted by heterogeneity among cancer cells and similarities among functionally related immune cells. Here, the authors develop scATOMIC, a tumour and TME cell type classifier based on a hierarchical approach that can be applied to pan-cancer datasets.
- Ido Nofech-Mozes
- , David Soave
- & Sagi Abelson
-
Article
| Open Access Integrated analysis of genomic and transcriptomic data for the discovery of splice-associated variants in cancer
Integrated analysis of genomic and transcriptomic data for the discovery of splice-associated variants in cancer
Analysing the regulatory consequences of mutations and splice variants at large scale in cancer requires efficient computational tools. Here, the authors develop RegTools, a software package that can identify splice-associated variants from large-scale genomics and transcriptomics data with efficiency and flexibility.
- Kelsy C. Cotto
- , Yang-Yang Feng
- & Malachi Griffith
-
Article
| Open Access Integrated transcriptome study of the tumor microenvironment for treatment response prediction in male predominant hypopharyngeal carcinoma
Integrated transcriptome study of the tumor microenvironment for treatment response prediction in male predominant hypopharyngeal carcinoma
Many patients with hypopharyngeal carcinoma (HPC) do not respond to first-line combination therapy. Here, the authors analyse the tumour and the tumour microenvironment of HPC patients treated with combination therapy using single-cell RNA-seq, and train a classifier to distinguish responders based on cell type composition.
- Yang Zhang
- , Gan Liu
- & Zhigang Huang
-
Article
| Open Access Towards routine chromosome-scale haplotype-resolved reconstruction in cancer genomics
Towards routine chromosome-scale haplotype-resolved reconstruction in cancer genomics
The precise inference of structural variants (SVs) requires suitable sequencing technologies and computational tools. Here, in order to analyse SVs with haplotype resolution, the author applies high-resolution long-read sequencing and long-range Hi-C to a melanoma cell line and develops an efficient graph-based computational framework, pstools.
- Shilpa Garg
-
Article
| Open Access Genomic and immune landscape Of metastatic pheochromocytoma and paraganglioma
Genomic and immune landscape Of metastatic pheochromocytoma and paraganglioma
The molecular mechanisms underlying metastasis in pheochromocytoma/paraganglioma (mPPGL) remain to be explored. Here, the authors perform genomic and immunogenomic profiling of mPPGL tumors and suggest potential biomarkers for risk of metastasis and immunotherapy response.
- Bruna Calsina
- , Elena Piñeiro-Yáñez
- & Mercedes Robledo
-
Article
| Open Access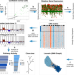 A variational algorithm to detect the clonal copy number substructure of tumors from scRNA-seq data
A variational algorithm to detect the clonal copy number substructure of tumors from scRNA-seq data
The inference of clonal architectures in cancer using single-cell RNA-seq data remains challenging. Here, the authors develop SCEVAN, a variational algorithm for copy number-based clonal structure inference in single-cell RNA-seq data that can characterise evolution and heterogeneity in the tumour and the microenvironment.
- Antonio De Falco
- , Francesca Caruso
- & Michele Ceccarelli
-
Article
| Open Access Single-cell transcriptome profiling of the stepwise progression of head and neck cancer
Single-cell transcriptome profiling of the stepwise progression of head and neck cancer
Head and neck squamous cell carcinomas (HNSCCs) undergo a stepwise progression from normal tissues. In order to better understand the molecular mechanisms behind such progression, here the authors profile HNSCC tumors at different stages using single-cell RNA-seq, and observe the role of interactions with the tumor microenvironment.
- Ji-Hye Choi
- , Bok-Soon Lee
- & Chul-Ho Kim
-
Article
| Open Access Spatial transcriptomics reveals niche-specific enrichment and vulnerabilities of radial glial stem-like cells in malignant gliomas
Spatial transcriptomics reveals niche-specific enrichment and vulnerabilities of radial glial stem-like cells in malignant gliomas
The spatial organisation of diffuse midline glioma-H3K27M mutant (DMG) and glioblastoma (GBM) remains to be investigated. Here, the authors integrate short-read and long-read spatial profiling of DMG and GBM to identify regulatory programs and cellular ecosystems in distinct glioma niches.
- Yanming Ren
- , Zongyao Huang
- & Yuan Wang
-
Article
| Open Access Reconstructing clonal tree for phylo-phenotypic characterization of cancer using single-cell transcriptomics
Reconstructing clonal tree for phylo-phenotypic characterization of cancer using single-cell transcriptomics
The functional changes of individual clones in single cell RNA sequencing (scRNA-seq) data remain elusive. Here, the authors develop PhylEx that integrates bulk genomics data with co-occurrences of mutations revealed by scRNA-seq data and apply it to high-grade serous ovarian cancer cell line and breast cancer datasets.
- Seong-Hwan Jun
- , Hosein Toosi
- & Jens Lagergren
-
Article
| Open Access Single-cell biological network inference using a heterogeneous graph transformer
Single-cell biological network inference using a heterogeneous graph transformer
Single-cell multi-omics and deep learning could lead to the inference of biological networks across specific cell types. Here, the authors develop DeepMAPS, a deep learning, graph-based approach for cell-type specific network inference from single-cell multi-omics data that is tested on healthy and tumour tissue datasets.
- Anjun Ma
- , Xiaoying Wang
- & Qin Ma
-
Article
| Open Access Reconstruction of the tumor spatial microenvironment along the malignant-boundary-nonmalignant axis
Reconstruction of the tumor spatial microenvironment along the malignant-boundary-nonmalignant axis
Delineating the cellular composition of tumour boundaries in spatial transcriptomics (ST) data is challenging. Here, the authors develop Cottrazm to integrate ST with histological imaging and single-cell data, identify the malignant and non-malignant tissue boundaries, deconvolute cell-type composition, and reconstruct cell type-specific gene expression profiles.
- Zhenzhen Xun
- , Xinyu Ding
- & Youqiong Ye
-
Comment
| Open AccessLarge-scale genomic analyses reveal alterations and mechanisms underlying clonal evolution and immune evasion in esophageal cancer
Esophageal cancers feature distinct manifestations between and within patients which complicate precision diagnosis, prognosis, and patient care. New genomic and epigenomic research uncovers novel mechanisms underlying both inter- and intra-tumoral heterogeneity in esophageal cancer, with significant biological and translational implications.
- De-Chen Lin
-
Article
| Open Access Tracking the evolution of esophageal squamous cell carcinoma under dynamic immune selection by multi-omics sequencing
Tracking the evolution of esophageal squamous cell carcinoma under dynamic immune selection by multi-omics sequencing
It is essential to understand heterogeneity and evolution at different omics levels in oesophageal squamous cell carcinoma (ESCC). Here, the authors use multi-omics to analyse heterogeneity and evolution in ESCC patient samples, and characterise the levels of immune infiltration as well as selective pressure from the tumour microenvironment.
- Sijia Cui
- , Nicholas McGranahan
- & Shixiu Wu
-
Article
| Open Access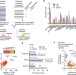 Widespread perturbation of ETS factor binding sites in cancer
Widespread perturbation of ETS factor binding sites in cancer
Few cancer drivers in non-coding regions have been identified so far. Here, the authors develop a transcription factor-aware burden test to predict non-coding variants and analyze the impact on transcription factor binding - especially ETS factors - as well as their impact on transcriptional activity.
- Sebastian Carrasco Pro
- , Heather Hook
- & Juan Ignacio Fuxman Bass
-
Article
| Open Access Genomic and microenvironmental heterogeneity shaping epithelial-to-mesenchymal trajectories in cancer
Genomic and microenvironmental heterogeneity shaping epithelial-to-mesenchymal trajectories in cancer
The intermediate states of the epithelial-to-mesenchymal transition (EMT) in cancer require further molecular characterisation. Here, the authors develop a method to evaluate EMT transformation and trajectories in cancer transcriptomics data, characterising EMT macro-states, including a hybrid state, and EMT hallmarks.
- Guidantonio Malagoli Tagliazucchi
- , Anna J. Wiecek
- & Maria Secrier
-
Article
| Open Access Loss of phosphatase CTDNEP1 potentiates aggressive medulloblastoma by triggering MYC amplification and genomic instability
Loss of phosphatase CTDNEP1 potentiates aggressive medulloblastoma by triggering MYC amplification and genomic instability
Group 3 medulloblastomas (MBs) have the worst prognosis amongst the subtypes of MBs and are associated with MYC amplifications. Here the authors identify that mutations in CTDNEP1 cause MYC activation, amplification, and genomic instability in this subtype of MBs.
- Zaili Luo
- , Dazhuan Xin
- & Q. Richard Lu
-
Article
| Open Access Application of high-throughput single-nucleus DNA sequencing in pancreatic cancer
Application of high-throughput single-nucleus DNA sequencing in pancreatic cancer
Implementing high-throughput single-cell DNA sequencing for the study of solid tumours has been challenging. Here, the authors present an optimised approach for snap-frozen tissue single nuclei extraction and DNA sequencing, which can be applied to study pancreatic ductal adenocarcinoma evolution and heterogeneity.
- Haochen Zhang
- , Elias-Ramzey Karnoub
- & Christine A. Iacobuzio-Donahue
-
Article
| Open Access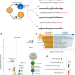 Multi-omics and machine learning reveal context-specific gene regulatory activities of PML::RARA in acute promyelocytic leukemia
Multi-omics and machine learning reveal context-specific gene regulatory activities of PML::RARA in acute promyelocytic leukemia
The PML-RARA gene fusion is the characteristic driver of Acute Promyelocytic Leukaemia (APL) and is known to bind to the genome. Here, the authors characterise the impact of PML-RARA on gene regulation in APL cell lines and patient samples using transcriptomics, epigenomics, and machine learning.
- William Villiers
- , Audrey Kelly
- & Cameron S. Osborne
-
Article
| Open Access Parallelized multidimensional analytic framework applied to mammary epithelial cells uncovers regulatory principles in EMT
Parallelized multidimensional analytic framework applied to mammary epithelial cells uncovers regulatory principles in EMT
Epithelial-to-mesenchymal transition (EMT) is a complex process regulated at multiple molecular levels. Here, the authors implement an analytic framework - PAMAF - to integrate data from twelve distinct omics modalities, which they use to understand the molecular changes and regulation during EMT in vitro.
- Indranil Paul
- , Dante Bolzan
- & Andrew Emili
-
Article
| Open Access Antitumour activity of neratinib in patients with HER2-mutant advanced biliary tract cancers
Antitumour activity of neratinib in patients with HER2-mutant advanced biliary tract cancers
In biliary tract cancer HER2 alterations correlate with poor prognosis. Here, the authors present the results of a phase II clinical trial reporting the efficacy and safety of the tyrosine kinase inhibitor neratinib in patients with HER2-mutation positive advanced biliary tract cancers.
- James J. Harding
- , Sarina A. Piha-Paul
- & Ghassan K. Abou-Alfa

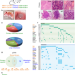 Distinct shared and compartment-enriched oncogenic networks drive primary versus metastatic breast cancer
Distinct shared and compartment-enriched oncogenic networks drive primary versus metastatic breast cancer