Featured
-
-
Article
| Open Access Multi-omic and functional analysis for classification and treatment of sarcomas with FUS-TFCP2 or EWSR1-TFCP2 fusions
Multi-omic and functional analysis for classification and treatment of sarcomas with FUS-TFCP2 or EWSR1-TFCP2 fusions
The molecular characteristics and therapeutic vulnerabilities of TFCP2-rearranged rhabdomyosarcomas (RMS) require further exploration. Here, the authors use multi-omics analyses and functional and mechanistic investigations to characterize TFCP2-rearranged RMS – including cases with FUS/EWSR1-TFCP2 fusions – across two precision oncology programs.
- Julia Schöpf
- , Sebastian Uhrig
- & Claudia Scholl
-
Article
| Open Access Single-cell multi-omics analysis of human testicular germ cell tumor reveals its molecular features and microenvironment
Single-cell multi-omics analysis of human testicular germ cell tumor reveals its molecular features and microenvironment
The molecular features and tumour microenvironment (TME) in seminoma remain to be characterised. Here, the authors perform single cell multi-omics and spatial transcriptomics and identify key gene expression programmes, immune cell subtypes and potential therapeutic targets.
- Xiaojian Lu
- , Yanwei Luo
- & Jingtao Guo
-
Article
| Open Access Genomic and epigenomic integrative subtypes of renal cell carcinoma in a Japanese cohort
Genomic and epigenomic integrative subtypes of renal cell carcinoma in a Japanese cohort
Renal cell carcinoma (RCC) subtypes are associated with different molecular alterations and clinical outcomes, but they need to be characterised in diverse cohorts. Here, the authors perform genomic, transcriptomic, and epigenomic profiling in a large cohort of Japanese RCC cases, and identify epi-subtypes associated with a particular immune environment.
- Akihiko Fukagawa
- , Natsuko Hama
- & Tatsuhiro Shibata
-
Article
| Open Access Whole-genome sequencing reveals the molecular implications of the stepwise progression of lung adenocarcinoma
Whole-genome sequencing reveals the molecular implications of the stepwise progression of lung adenocarcinoma
Current sequencing technologies can shed light on the stepwise progression of lung adenocarcinoma. Here, the authors characterize tumor progression in lung adenocarcinomas from an early stage using short and long read whole-genome sequencing, bulk and spatial transcriptomics, and epigenomics.
- Yasuhiko Haga
- , Yoshitaka Sakamoto
- & Ayako Suzuki
-
Article
| Open Access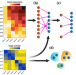 Accurate integration of single-cell DNA and RNA for analyzing intratumor heterogeneity using MaCroDNA
Accurate integration of single-cell DNA and RNA for analyzing intratumor heterogeneity using MaCroDNA
Here, the authors develop MaCroDNA, an algorithm to integrate single-cell DNA and RNA sequencing data from the same tissue. They use MaCroDNA to show—in agreement with previous studies—that copy number changes can predict progression from Barrett’s esophagus to esophageal adenocarcinoma.
- Mohammadamin Edrisi
- , Xiru Huang
- & Luay Nakhleh
-
Article
| Open Access Single cell multi-omics reveal intra-cell-line heterogeneity across human cancer cell lines
Single cell multi-omics reveal intra-cell-line heterogeneity across human cancer cell lines
Intra-cell line heterogeneity remains to be characterized. Here, the use of single multi-omics on a large panel of human cell lines identifies copy number variation, epigenetic variation and extrachromosomal DNA distribution as the main contributors to intra-cell line heterogeneity.
- Qionghua Zhu
- , Xin Zhao
- & Liang Wu
-
Article
| Open Access Delineating the early dissemination mechanisms of acral melanoma by integrating single-cell and spatial transcriptomic analyses
Delineating the early dissemination mechanisms of acral melanoma by integrating single-cell and spatial transcriptomic analyses
Acral melanoma (AM) is a rare melanoma subtype with unique features, where lymph node metastasis is closely associated with clinical outcomes. Here, the authors use single-cell and spatial transcriptomics to analyse early dissemination, tumour microenvironment, and heterogeneity in AM, and infer metabolic shifts with therapeutic implications.
- Chuanyuan Wei
- , Wei Sun
- & Jianying Gu
-
Article
| Open Access Mosaic chromosomal alterations in peripheral blood leukocytes of children in sub-Saharan Africa
Mosaic chromosomal alterations in peripheral blood leukocytes of children in sub-Saharan Africa
Mosaic chromosomal alterations (mCAs) in peripheral blood leukocytes are associated with an increased risk of malignancy. Here, the authors use genome-wide genotyping array data to investigate the prevalence of mCAs in sub-Saharan African children with versus those without Burkitt lymphoma.
- Weiyin Zhou
- , Anja Fischer
- & Sam M. Mbulaiteye
-
Article
| Open Access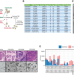 Mapping and modeling human colorectal carcinoma interactions with the tumor microenvironment
Mapping and modeling human colorectal carcinoma interactions with the tumor microenvironment
Tumour-microenvironment interactions, pivotal in cancer progression, are challenging to replicate in vitro. Here, the authors use single-cell RNA-seq to analyse these interactions in colorectal cancer within organoid models, and aim to emulate and understand these crucial interactions by introducing specific microenvironmental components.
- Ning Li
- , Qin Zhu
- & Christopher J. Lengner
-
Article
| Open Access Delineating the interplay between oncogenic pathways and immunity in anaplastic Wilms tumors
Delineating the interplay between oncogenic pathways and immunity in anaplastic Wilms tumors
Treatment of diffuse anaplastic Wilms tumours (DAWT) remains a challenge. Here, the authors perform multi-omic analysis and identify a desert-like DAWT subtype accounting for one third of DAWT cases and suggest treating them with HDAC and/or WEE1 inhibitors.
- Xiaoping Su
- , Xiaofan Lu
- & Gabriel G. Malouf
-
Article
| Open Access Contribution of pks+ E. coli mutations to colorectal carcinogenesis
Contribution of pks+ E. coli mutations to colorectal carcinogenesis
Common driver mutations in colorectal cancer (CRC) are not always consistent with frequent mutational signatures. Here, the authors analyse spatially annotated colon crypts in CRC patients and find mutational signatures of pks+ E. coli that are consistent with driver mutations, suggesting a potential role of pks+ E. coli in carcinogenesis.
- Bingjie Chen
- , Daniele Ramazzotti
- & Andrea Sottoriva
-
Article
| Open Access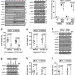 HDAC8-mediated inhibition of EP300 drives a transcriptional state that increases melanoma brain metastasis
HDAC8-mediated inhibition of EP300 drives a transcriptional state that increases melanoma brain metastasis
The drivers of melanoma brain metastases (MBM) remain poorly understood. Here, the authors identify stress-induced HDAC8 activity as the driver of a neural crest-stem cell like transcriptional state that leads to MBM, and explore the molecular mechanism that drives this transition.
- Michael F. Emmons
- , Richard L. Bennett
- & Keiran S. M. Smalley
-
Article
| Open Access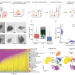 Fragment-sequencing unveils local tissue microenvironments at single-cell resolution
Fragment-sequencing unveils local tissue microenvironments at single-cell resolution
Experimentally preserving tissue microenvironments remains challenging for single-cell sequencing methods. Here, the authors introduce fragment-sequencing, a method that can preserve three-dimensional microenvironments in single-cell RNA-seq, thus allowing the reconstruction of spatial tissue niches.
- Kristina Handler
- , Karsten Bach
- & Andreas E. Moor
-
Article
| Open Access Genomic profiling of subcutaneous patient-derived xenografts reveals immune constraints on tumor evolution in childhood solid cancer
Genomic profiling of subcutaneous patient-derived xenografts reveals immune constraints on tumor evolution in childhood solid cancer
Subcutaneous patient-derived xenografts are a common tool in cancer research. Here, the authors compare 65 paired early passage xenografts to their original paediatric tumour and show clonal evolution determines seeding of the xenograft.
- Funan He
- , Abhik M. Bandyopadhyay
- & Siyuan Zheng
-
Article
| Open Access Targetable lesions and proteomes predict therapy sensitivity through disease evolution in pediatric acute lymphoblastic leukemia
Targetable lesions and proteomes predict therapy sensitivity through disease evolution in pediatric acute lymphoblastic leukemia
The role of clonal evolution on the actionable proteome and response to therapy in childhood acute lymphoblastic leukemia (ALL) remains unknown. Here, targeted sequencing and proteomic analysis of paired ALL diagnosis and relapsed samples revealed PARP1 as a potential therapeutic target.
- Amanda C. Lorentzian
- , Jenna Rever
- & Philipp F. Lange
-
Article
| Open Access Genomic profiling and pre-clinical modelling of breast cancer leptomeningeal metastasis reveals acquisition of a lobular-like phenotype
Genomic profiling and pre-clinical modelling of breast cancer leptomeningeal metastasis reveals acquisition of a lobular-like phenotype
Breast cancer leptomeningeal metastasis is difficult to assess due to the site. Here, the authors assess cerebrospinal fluid cell-free DNA to show specific mutations present in leptomeningeal metastasis, and develop organoids of the disease.
- Amanda Fitzpatrick
- , Marjan Iravani
- & Clare M. Isacke
-
Article
| Open Access Tracing cancer evolution and heterogeneity using Hi-C
Tracing cancer evolution and heterogeneity using Hi-C
It is challenging to analyse chromosomal rearrangements in heterogeneous solid cancers. Here the authors present HiDENSEC, a method to jointly infer absolute copy number, ploidy, tumor purity and large-scale rearrangements from Hi-C data. The increased statistical power afforded by joint inference enables novel insights into cancer genome evolution.
- Dan Daniel Erdmann-Pham
- , Sanjit Singh Batra
- & Dirk Hockemeyer
-
Comment
| Open AccessMoving closer towards a comprehensive view of tumor biology and microarchitecture using spatial transcriptomics
Spatial transcriptomic profiling of cancer has enabled spatial delineation of malignant transcriptional heterogeneity, intercellular communication, and organization of microanatomical structures within the tumor microenvironment. This technical breakthrough paves the way for the development of precision diagnostic methods and targeted therapies.
- Young Min Park
- & De-Chen Lin
-
Article
| Open Access The chromatin network helps prevent cancer-associated mutagenesis at transcription-replication conflicts
The chromatin network helps prevent cancer-associated mutagenesis at transcription-replication conflicts
Epigenetic alterations are frequent in human malignancies and have been shown to threaten genome integrity. Here the authors show that a chromatin network prevents R-loops and transcription-replication conflicts from genomic instability and mutagenic signatures frequently associated with cancer.
- Aleix Bayona-Feliu
- , Emilia Herrera-Moyano
- & Andrés Aguilera
-
Article
| Open Access Chromatin accessibility landscape of relapsed pediatric B-lineage acute lymphoblastic leukemia
Chromatin accessibility landscape of relapsed pediatric B-lineage acute lymphoblastic leukemia
The molecular mechanisms underlying relapse in pediatric B-lineage acute lymphoblastic leukemia (B-ALL) patients remain to be explored. Here, the authors characterise the chromatin accessibility landscape of B-ALL and identify subtype and drug response specific patterns.
- Han Wang
- , Huiying Sun
- & Yu Liu
-
Article
| Open Access Expanding PROTACtable genome universe of E3 ligases
Expanding PROTACtable genome universe of E3 ligases
Proteolysis-targeting chimeras (PROTACs) offer new avenues for drug development. Here the authors investigate E3 ligases—key to PROTAC function—and identify candidate targets for cancer drivers such as KRAS and EGFR.
- Yuan Liu
- , Jingwen Yang
- & Leng Han
-
Article
| Open Access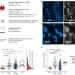 Molecular landscape and functional characterization of centrosome amplification in ovarian cancer
Molecular landscape and functional characterization of centrosome amplification in ovarian cancer
The prevalence of centrosome amplification (CA) and the genomic landscape of chromosomal instability in high-grade serous ovarian carcinoma (HGSOC) remain to be explored. Here the authors suggest CA as a potential driver of tumour evolution and a biomarker for treatment response in HGSOC.
- Carolin M. Sauer
- , James A. Hall
- & James D. Brenton
-
Article
| Open Access Rewiring of the promoter-enhancer interactome and regulatory landscape in glioblastoma orchestrates gene expression underlying neurogliomal synaptic communication
Rewiring of the promoter-enhancer interactome and regulatory landscape in glioblastoma orchestrates gene expression underlying neurogliomal synaptic communication
The integration of transcriptomics and epigenomics data helps to better understand the regulatory and topological changes in glioblastoma subtypes. Here, the authors map the promoter-enhancer interactome and regulatory landscape and show changes in promoter-enhancer interactions, chromatin accessibility, and redistribution of histone marks across four glioblastoma subtypes.
- Chaitali Chakraborty
- , Itzel Nissen
- & Silvia Remeseiro
-
Article
| Open Access Super-enhancer hijacking drives ectopic expression of hedgehog pathway ligands in meningiomas
Super-enhancer hijacking drives ectopic expression of hedgehog pathway ligands in meningiomas
Hedgehog signalling is known to be linked to oncogenic proliferation. Here, the authors identify structural events as a mechanism of Hedgehog activation in over one-third of driver unknown meningiomas.
- Mark W. Youngblood
- , Zeynep Erson-Omay
- & Murat Günel
-
Article
| Open Access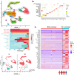 Single-cell analysis reveals altered tumor microenvironments of relapse- and remission-associated pediatric acute myeloid leukemia
Single-cell analysis reveals altered tumor microenvironments of relapse- and remission-associated pediatric acute myeloid leukemia
Single-cell RNA-seq could help identify acute myeloid leukaemia (AML) patients at high risk of relapse after therapy. Here, the authors use single-cell RNA-seq from paediatric AML samples to construct a 7-gene signature that can identify malignant cells at diagnosis, which are distinctly associated with relapse or complete remission.
- Hope Mumme
- , Beena E. Thomas
- & Manoj Bhasin
-
Article
| Open Access Genomic signatures of past and present chromosomal instability in Barrett’s esophagus and early esophageal adenocarcinoma
Genomic signatures of past and present chromosomal instability in Barrett’s esophagus and early esophageal adenocarcinoma
Genome complexity is a distinguishing feature of advanced cancers in contrast to precancerous conditions. Here, by analysing chromosomal copy-number evolution in early cancers and precancerous lesions of the oesophagus, the authors reveal signatures of ongoing chromosomal instability and its role in promoting tumour progression.
- Chunyang Bao
- , Richard W. Tourdot
- & Cheng-Zhong Zhang
-
Article
| Open Access Integrated molecular and multiparametric MRI mapping of high-grade glioma identifies regional biologic signatures
Integrated molecular and multiparametric MRI mapping of high-grade glioma identifies regional biologic signatures
Glioma tumours are known to be heterogenous in mutation and gene expression patterns, but sampling limitations can lead to inaccurate detection of evolutionary events. Here, the authors carry out multi-omics analysis of multi-regional biopsies from 68 patients and show differential mutations in non-enhancing regions.
- Leland S. Hu
- , Fulvio D’Angelo
- & Nhan L. Tran
-
Article
| Open Access Evolutionary signatures of human cancers revealed via genomic analysis of over 35,000 patients
Evolutionary signatures of human cancers revealed via genomic analysis of over 35,000 patients
The identification of cancer type-specific evolutionary signatures could be used for patient stratification. Here, the authors develop a method, ASCETIC, that utilises bulk and single-cell sequencing data and identifies evolutionary signatures associated with different prognostic outcomes across different cancer types.
- Diletta Fontana
- , Ilaria Crespiatico
- & Daniele Ramazzotti
-
Article
| Open Access Genome-wide enhancer-gene regulatory maps link causal variants to target genes underlying human cancer risk
Genome-wide enhancer-gene regulatory maps link causal variants to target genes underlying human cancer risk
Here, the authors apply the Activity-by-Contact (ABC) model to infer enhancer-gene regulation and the effect of associated variants across multiple cancer types, integrating genetic and multi-omics data. Then, they explore the mechanisms associated with ABC regulatory variants in colorectal cancer.
- Pingting Ying
- , Can Chen
- & Xiaoping Miao
-
Article
| Open Access Targetable NOTCH1 rearrangements in reninoma
Targetable NOTCH1 rearrangements in reninoma
Reninomas are very rare kidney tumours of juxtaglomerular cells. Here, the authors analyse reninomas using whole-genome and transcriptome sequencing, and reveal the presence and functional effects of NOTCH1 rearrangements.
- Taryn D. Treger
- , John E. G. Lawrence
- & Tanzina Chowdhury
-
Article
| Open Access Epigenomic analysis of formalin-fixed paraffin-embedded samples by CUT&Tag
Epigenomic analysis of formalin-fixed paraffin-embedded samples by CUT&Tag
Conducting epigenomic studies on FFPE samples is traditionally challenging due to chromatin damage caused due to exposure to formaldehyde. Here, the authors show that an optimisation of their previous CUTAC method allows the production of high-resolution maps of regulatory elements from FFPE samples.
- Steven Henikoff
- , Jorja G. Henikoff
- & Eric C. Holland
-
Article
| Open Access Global impact of somatic structural variation on the cancer proteome
Global impact of somatic structural variation on the cancer proteome
The relevance of non-coding somatic mutations in cancer remains elusive. Here, the combination of mass spectrometry-based proteomics and whole genome sequencing data across multiple cancer types helps to assess the effects of somatic structural variant breakpoint patterns on protein expression of nearby genes.
- Fengju Chen
- , Yiqun Zhang
- & Chad J. Creighton
-
Article
| Open Access Transposable elements as tissue-specific enhancers in cancers of endodermal lineage
Transposable elements as tissue-specific enhancers in cancers of endodermal lineage
A comprehensive characterisation of transposable elements (TEs) in cancer is lacking. Here, the authors identify distinct TEs with different genomic features as tissue-specific enhancers in colorectal and liver cancer.
- Konsta Karttunen
- , Divyesh Patel
- & Biswajyoti Sahu
-
Article
| Open Access Barcoded multiple displacement amplification for high coverage sequencing in spatial genomics
Barcoded multiple displacement amplification for high coverage sequencing in spatial genomics
Spatial genomics offers insights into cellular interactions within tissues. Here, the authors develop barcoded multiple displacement amplification, achieving high-coverage sequencing to map complex genomic variations within cellular landscapes.
- Jinhyun Kim
- , Sungsik Kim
- & Sunghoon Kwon
-
Article
| Open Access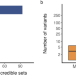 Androgen receptor binding sites enabling genetic prediction of mortality due to prostate cancer in cancer-free subjects
Androgen receptor binding sites enabling genetic prediction of mortality due to prostate cancer in cancer-free subjects
The prediction of mortality due to prostate cancer remains challenging. Here, the authors perform trans-ancestry metaanalysis with a focus on binding sites of the androgen receptor and develop a polygenic risk score.
- Shuji Ito
- , Xiaoxi Liu
- & Chikashi Terao
-
Article
| Open Access A biallelic multiple nucleotide length polymorphism explains functional causality at 5p15.33 prostate cancer risk locus
A biallelic multiple nucleotide length polymorphism explains functional causality at 5p15.33 prostate cancer risk locus
Here, the authors functionally characterize a complex genetic variant relevant in prostate cancer that regulates IRX4 expression through epigenetic activation. This work highlights the significance of non-single nucleotide polymorphism causal variants in explaining disease risk.
- Sandor Spisak
- , Viktoria Tisza
- & Matthew L. Freedman
-
Comment
| Open Access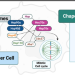 Epichaperomics reveals dysfunctional chaperone protein networks
Epichaperomics reveals dysfunctional chaperone protein networks
Molecular chaperones establish essential protein-protein interaction networks. Modified versions of these assemblies are generally enriched in certain maladies. A study published in Nature Communications used epichaperomics to identify unique changes occurring in chaperone-formed protein networks during mitosis in cancer cells.
- Mark R. Woodford
- , Dimitra Bourboulia
- & Mehdi Mollapour
-
Article
| Open Access Molecular features and clinical implications of the heterogeneity in Chinese patients with HER2-low breast cancer
Molecular features and clinical implications of the heterogeneity in Chinese patients with HER2-low breast cancer
HER2-low breast cancer has recently been defined as a potential subtype for sensitivity to novel antibody-drug conjugates. Here, the authors analyse a multiomics cohort of 434 HER2-low patients and find an altered molecular status compared to other subtypes and the interpatient heterogeneity within this subtype.
- Lei-Jie Dai
- , Ding Ma
- & Zhi-Ming Shao
-
Article
| Open Access NFIB facilitates replication licensing by acting as a genome organizer
NFIB facilitates replication licensing by acting as a genome organizer
The precise rule of replication origin selection and activation in metazoans remains unclear. Here, the authors identify NFIB as a genome organizer and replication pioneer by facilitating nucleosome remodeling and chromatin assembly of the pre-RC.
- Wenting Zhang
- , Yue Wang
- & Yongfeng Shang
-
Article
| Open Access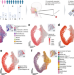 Spatial transcriptomics reveals distinct and conserved tumor core and edge architectures that predict survival and targeted therapy response
Spatial transcriptomics reveals distinct and conserved tumor core and edge architectures that predict survival and targeted therapy response
Oral squamous cell carcinoma is known to contain altered tumour cells within both the tumour core and leading edge. Here, the authors utilise spatial transcriptomics to characterise differences in gene expression and ligand-receptor architecture between areas of the tumour.
- Rohit Arora
- , Christian Cao
- & Pinaki Bose
-
Article
| Open Access Resolving the spatial architecture of myeloma and its microenvironment at the single-cell level
Resolving the spatial architecture of myeloma and its microenvironment at the single-cell level
The spatial architecture of multiple myeloma remains to be explored. Here, the authors perform bulk and single cell sequencing for samples from newly diagnosed patients and reveal gene signatures associated with focal lesions and spatial heterogeneity in the tumour microenvironment.
- Lukas John
- , Alexandra M. Poos
- & Niels Weinhold
-
Article
| Open Access Genomic analysis and clinical correlations of non-small cell lung cancer brain metastasis
Genomic analysis and clinical correlations of non-small cell lung cancer brain metastasis
The genomic landscape of brain metastasis (BM) in patients with non-small cell lung cancer (NSCLC) remains to be explored. Here, the authors analyse a cohort of 233 patients with BM including 47 primary tumour, 42 extracranial metastatic matched samples and reveal distinct mutational patterns.
- Anna Skakodub
- , Henry Walch
- & Luke R. G. Pike
-
Article
| Open Access Copy number architectures define treatment-mediated selection of lethal prostate cancer clones
Copy number architectures define treatment-mediated selection of lethal prostate cancer clones
The heterogeneity of androgen receptor (AR) gene alterations across metastases in prostate cancer remains unresolved. Here, the authors characterise AR genomic complexity across spatially separated lethal metastases from 10 prostate cancer patients and investigate how AR alterations evolve.
- A. M. Mahedi Hasan
- , Paolo Cremaschi
- & Gerhardt Attard
-
Article
| Open Access MAPK inhibitor sensitivity scores predict sensitivity driven by the immune infiltration in pediatric low-grade gliomas
MAPK inhibitor sensitivity scores predict sensitivity driven by the immune infiltration in pediatric low-grade gliomas
The MAPK pathway is a key driver of pediatric low-grade gliomas (pLGG); however, response to MAPK inhibitors (MAPKi) in pLGG patients is not consistent. Here, the authors develop MAPKi sensitivity scores (MSS) to predict response to MAPKi and apply them to bulk and single-cell sequencing datasets from pLGG patients and preclinical models.
- Romain Sigaud
- , Thomas K. Albert
- & Till Milde
-
Article
| Open Access Obesity-associated changes in molecular biology of primary breast cancer
Obesity-associated changes in molecular biology of primary breast cancer
The association between obesity and breast cancer biology remains understudied in humans. Here, using a large retrospective data collection, the authors identify obesity associated changes in the genomic, transcriptomic profile, and the tumor microenvironment of primary untreated breast tumors.
- Ha-Linh Nguyen
- , Tatjana Geukens
- & Christine Desmedt
-
Article
| Open Access The copy number and mutational landscape of recurrent ovarian high-grade serous carcinoma
The copy number and mutational landscape of recurrent ovarian high-grade serous carcinoma
‘Treatment resistance is common in ovarian high grade serous carcinoma, often leading to relapse. Here, the authors leverage shallow whole genome and panel sequencing of 276 patients with available diagnostic and relapse samples and show high concordance of copy number and mutation status.
- Philip Smith
- , Thomas Bradley
- & Iain A. McNeish
-
Article
| Open Access Identification of transcriptional programs using dense vector representations defined by mutual information with GeneVector
Identification of transcriptional programs using dense vector representations defined by mutual information with GeneVector
In single-cell RNA-seq analyses, it would be critical to measure the relationships between genes. Here, the authors develop a framework for single-cell dimensionality reduction that incorporates gene-specific relationships - GeneVector -, and use it for tasks such as annotating cell types and analysing pathway variation after treatment.
- Nicholas Ceglia
- , Zachary Sethna
- & Andrew McPherson
-
Article
| Open Access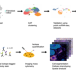 Cancer-associated fibroblast classification in single-cell and spatial proteomics data
Cancer-associated fibroblast classification in single-cell and spatial proteomics data
Cancer-associated fibroblasts (CAFs) have different subtypes and play diverse roles in the tumour microenvironment. Here, the authors use single-cell RNA-seq and multiplex imaging mass cytometry data to propose a CAF classification scheme of nine subtypes across different cancer types.
- Lena Cords
- , Sandra Tietscher
- & Bernd Bodenmiller
-
Article
| Open Access Next generation pan-cancer blood proteome profiling using proximity extension assay
Next generation pan-cancer blood proteome profiling using proximity extension assay
Comprehensive and scalable proteomic profiling of plasma samples can improve the screening and diagnosis of cancer patients. Here, the authors use the Olink Proximity Extension Assay technology to characterise the plasma proteomes of 1477 patients across twelve cancer types, and use machine learning to obtain a protein panel for cancer classification.
- María Bueno Álvez
- , Fredrik Edfors
- & Mathias Uhlén

 The ALT pathway generates telomere fusions that can be detected in the blood of cancer patients
The ALT pathway generates telomere fusions that can be detected in the blood of cancer patients