Featured
-
-
Article
| Open Access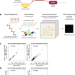 Mutational scanning pinpoints distinct binding sites of key ATGL regulators in lipolysis
Mutational scanning pinpoints distinct binding sites of key ATGL regulators in lipolysis
ATGL is a key enzyme in intracellular lipolysis. Here, the authors use deep mutational scanning to define the determinants of protein interaction between ATGL and its regulatory partners, gaining insights into lipolysis mechanisms in cells.
- Johanna M. Kohlmayr
- , Gernot F. Grabner
- & Ulrich Stelzl
-
Article
| Open Access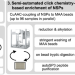 An integrated workflow for quantitative analysis of the newly synthesized proteome
An integrated workflow for quantitative analysis of the newly synthesized proteome
Analysis of newly synthesized proteins upon perturbation can provide detailed insights into immediate proteome remodeling, which drives cellular responses. Here, the authors report an optimized semi-automated workflow for the quantitative analysis of the newly synthesized proteome.
- Toman Borteçen
- , Torsten Müller
- & Jeroen Krijgsveld
-
Article
| Open Access Speos: an ensemble graph representation learning framework to predict core gene candidates for complex diseases
Speos: an ensemble graph representation learning framework to predict core gene candidates for complex diseases
Understanding phenotype-genotype relationships is a grand challenge of current biological research. Here, the authors use graph representation learning to identify human genes which display key characteristics of core genes for five complex diseases.
- Florin Ratajczak
- , Mitchell Joblin
- & Matthias Heinig
-
Article
| Open Access A genome-scale metabolic model of parasitic whipworm
A genome-scale metabolic model of parasitic whipworm
In this work, Bay et al describe the construction of the first genome-scale metabolic model for the parasitic whipworm, Trichuris muris and use it to identify novel metabolic pathways and predict critical enzymes and essential metabolites for worm survival.
- Ömer F. Bay
- , Kelly S. Hayes
- & Ian S. Roberts
-
Article
| Open Access Paired yeast one-hybrid assays to detect DNA-binding cooperativity and antagonism across transcription factors
Paired yeast one-hybrid assays to detect DNA-binding cooperativity and antagonism across transcription factors
Combinations of transcription factors (TFs) bind DNA to fine-tune gene expression. Here, the authors map cooperative and antagonistic DNA binding across hundreds of TF-pairs. TF-TF relationships vary depending on DNA targets and TF isoforms.
- Anna Berenson
- , Ryan Lane
- & Juan I. Fuxman Bass
-
Article
| Open Access Identification of gene function based on models capturing natural variability of Arabidopsis thaliana lipid metabolism
Identification of gene function based on models capturing natural variability of Arabidopsis thaliana lipid metabolism
The use of automated tools to reconstruct lipid metabolic pathways is not warranted in plants. Here, the authors construct Plant Lipid Module for Arabidopsis rosette using constraint-based modeling, demonstrate its integration in other plant metabolic models, and use it to dissect the genetic architecture of lipid metabolism.
- Sandra Correa Córdoba
- , Hao Tong
- & Zoran Nikoloski
-
Article
| Open Access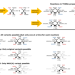 Network-wide thermodynamic constraints shape NAD(P)H cofactor specificity of biochemical reactions
Network-wide thermodynamic constraints shape NAD(P)H cofactor specificity of biochemical reactions
NADH and NADPH are redox cofactors coexisting in all living cells. Here, the authors present a computational study suggesting that evolved NAD(P)H reaction specificities in E. coli are largely shaped by metabolic network structure enabling maximal thermodynamic driving forces close to the theoretical optimum.
- Pavlos Stephanos Bekiaris
- & Steffen Klamt
-
Article
| Open Access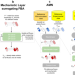 A neural-mechanistic hybrid approach improving the predictive power of genome-scale metabolic models
A neural-mechanistic hybrid approach improving the predictive power of genome-scale metabolic models
Mechanistic models estimate the phenotype of microorganisms in different environments but may have limited predictive capabilities. Here, authors develop trainable hybrid models with improved predictability using mechanistic insights and smaller training sets than conventional machine learning techniques.
- Léon Faure
- , Bastien Mollet
- & Jean-Loup Faulon
-
Article
| Open Access A general model-based causal inference method overcomes the curse of synchrony and indirect effect
A general model-based causal inference method overcomes the curse of synchrony and indirect effect
Traditional causal inference methods struggle to distinguish direct causation from synchrony and indirect effects. Here, authors present GOBI that overcomes this by testing a general model’s ability to reproduce data, providing accurate and broadly applicable causality inference for complex systems.
- Se Ho Park
- , Seokmin Ha
- & Jae Kyoung Kim
-
Article
| Open Access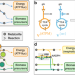 Functional decomposition of metabolism allows a system-level quantification of fluxes and protein allocation towards specific metabolic functions
Functional decomposition of metabolism allows a system-level quantification of fluxes and protein allocation towards specific metabolic functions
Quantifying the contribution of individual molecular components to complex cellular processes is a grand challenge in systems biology. Here, the authors present a general theoretical framework (Functional Decomposition of Metabolism, FDM) to quantify the contribution of every metabolic reaction to metabolic functions, e.g. the synthesis of biomass building blocks.
- Matteo Mori
- , Chuankai Cheng
- & Terence Hwa
-
Article
| Open Access Empowering drug off-target discovery with metabolic and structural analysis
Empowering drug off-target discovery with metabolic and structural analysis
The authors present a workflow integrating metabolic perturbations with protein structural analysis to identify drug off-targets, demonstrating how combining machine learning methods with mechanistic analyses can benefit off-target identification.
- Sourav Chowdhury
- , Daniel C. Zielinski
- & Eugene I. Shakhnovich
-
Article
| Open Access Universal structures for adaptation in biochemical reaction networks
Universal structures for adaptation in biochemical reaction networks
At the molecular level, the evolution of life is driven by the generation and diversification of adaptation mechanisms. Here Araujo and Liotta identify definitive and universal structural requirements for adaptation via intermolecular interactions.
- Robyn P. Araujo
- & Lance A. Liotta
-
Article
| Open Access Nucleocytoplasmic transport of active HER2 causes fractional escape from the DCIS-like state
Nucleocytoplasmic transport of active HER2 causes fractional escape from the DCIS-like state
HER2 receptor aberrations are more common in breast DCIS premalignancy than in breast cancer. Here the authors identify a feedback circuit involving HER2 nucleocytoplasmic transport that may explain why some DCIS lesions progress and others do not.
- Lixin Wang
- , B. Bishal Paudel
- & Kevin A. Janes
-
Article
| Open Access Tryptase β regulation of joint lubrication and inflammation via proteoglycan-4 in osteoarthritis
Tryptase β regulation of joint lubrication and inflammation via proteoglycan-4 in osteoarthritis
Altered expression and function of the extracellular matrix protein PRG4 have been associated with osteoarthritis. Here, the authors show that mast cell tryptase β cleaves PRG4, resulting in a reduction of lubrication and activation of inflammation in this context.
- Nabangshu Das
- , Luiz G. N. de Almeida
- & Antoine Dufour
-
Article
| Open Access A cybergenetic framework for engineering intein-mediated integral feedback control systems
A cybergenetic framework for engineering intein-mediated integral feedback control systems
Homeostasis and robust perfect adaptation are remarkable features of living cells. Here, to synthetically achieve this, the authors present a theoretical and experimental framework using inteins to implement compact biomolecular integral feedback controllers.
- Stanislav Anastassov
- , Maurice Filo
- & Mustafa Khammash
-
Article
| Open Access Size limits the sensitivity of kinetic schemes
Size limits the sensitivity of kinetic schemes
Living things rely on extremely sensitive molecular circuits. Here, authors uncover a universal structural limit on kinetic scheme sensitivity, with implications for gene regulation & the functions of condensates.
- Jeremy A. Owen
- & Jordan M. Horowitz
-
Article
| Open Access Genetically personalised organ-specific metabolic models in health and disease
Genetically personalised organ-specific metabolic models in health and disease
Here, the authors present a method to build genetically personalised metabolic models across tissues to estimate individualised reaction fluxes. A fluxome-wide association study in UK Biobank identifies fluxes associated with metabolites and coronary artery disease.
- Carles Foguet
- , Yu Xu
- & Michael Inouye
-
Article
| Open Access Quantitative fragmentomics allow affinity mapping of interactomes
Quantitative fragmentomics allow affinity mapping of interactomes
Protein networks have been widely explored but most binding affinities remain unknown, limiting the quantitative interpretation of interactomes. Here the authors measure affinities of 65,000 interactions involving human PDZ domains and target sequence motifs relevant for viral infection and cancer.
- Gergo Gogl
- , Boglarka Zambo
- & Gilles Travé
-
Article
| Open Access Scalable multiplex co-fractionation/mass spectrometry platform for accelerated protein interactome discovery
Scalable multiplex co-fractionation/mass spectrometry platform for accelerated protein interactome discovery
Co-fractionation/mass spectrometry (CF/MS) allows mapping protein interactomes but efficiency and quantitative accuracy are limited. Here, the authors develop a reproducible multiplexed CF/MS method and apply it to characterize interactome rewiring in breast cancer cells.
- Pierre C. Havugimana
- , Raghuveera Kumar Goel
- & Andrew Emili
-
Article
| Open Access Endosomal LC3C-pathway selectively targets plasma membrane cargo for autophagic degradation
Endosomal LC3C-pathway selectively targets plasma membrane cargo for autophagic degradation
Autophagy can selectively target cargo for degradation. Here the authors map the proximal interactome of ATG8-paralogs LC3B and LC3C uncovering an LC3C-Endocytic-Associated-Pathway that selectively recruits internalized plasma membrane cargo, Met and transferrin receptors, to nascent autophagosomes.
- Paula P. Coelho
- , Geoffrey G. Hesketh
- & Morag Park
-
Article
| Open Access Reconstruction of a catalogue of genome-scale metabolic models with enzymatic constraints using GECKO 2.0
Reconstruction of a catalogue of genome-scale metabolic models with enzymatic constraints using GECKO 2.0
Genome-scale metabolic models have been widely used for quantitative exploration of the relation between genotype and phenotype. Here the authors present GECKO 2, an automated framework for continuous and version controlled update of enzyme-constrained models of metabolism, producing an interesting catalogue of high-quality models for diverse yeasts, bacteria and human metabolism, aiming to facilitate their use in basic science, metabolic engineering and synthetic biology purposes.
- Iván Domenzain
- , Benjamín Sánchez
- & Jens Nielsen
-
Article
| Open Access A scalable, open-source implementation of a large-scale mechanistic model for single cell proliferation and death signaling
A scalable, open-source implementation of a large-scale mechanistic model for single cell proliferation and death signaling
Mechanistic models of how single cells respond to different perturbations can help integrate disparate big data sets or predict response to varied drug combinations. Here the authors develop a scalable, open-source pipeline for constructing and simulating large-scale, single-cell mechanistic models, an important building block for clinically-predictive mechanistic models and interpretable big data integration.
- Cemal Erdem
- , Arnab Mutsuddy
- & Marc R. Birtwistle
-
Article
| Open Access Global stable-isotope tracing metabolomics reveals system-wide metabolic alternations in aging Drosophila
Global stable-isotope tracing metabolomics reveals system-wide metabolic alternations in aging Drosophila
Stable-isotope tracing allows quantifying metabolic activity by measuring isotopically labeled metabolites, but its metabolome coverage has been limited. Here, the authors develop a global isotope tracing approach with metabolome-wide coverage and use it to characterize metabolic activities in aging Drosophila.
- Ruohong Wang
- , Yandong Yin
- & Zheng-Jiang Zhu
-
Article
| Open Access Artificial neural networks enable genome-scale simulations of intracellular signaling
Artificial neural networks enable genome-scale simulations of intracellular signaling
Many diseases are caused by disruptions to the network of biochemical reactions that allow cells to respond to external signals. Here Nilsson et al develop a method to simulate cellular signaling using artificial neural networks to predict cellular responses and activities of signaling molecules.
- Avlant Nilsson
- , Joshua M. Peters
- & Douglas A. Lauffenburger
-
Article
| Open Access In vitro reconstitution of Escherichia coli divisome activation
In vitro reconstitution of Escherichia coli divisome activation
In E. coli, FtsA and FtsZ control the place and time of cell division. Here, the authors use in vitro experiments to show how FtsA can follow FtsZ treadmilling and that downstream proteins form dynamic copolymers with FtsA to initiate division.
- Philipp Radler
- , Natalia Baranova
- & Martin Loose
-
Article
| Open Access Elucidating Human Milk Oligosaccharide biosynthetic genes through network-based multi-omics integration
Elucidating Human Milk Oligosaccharide biosynthetic genes through network-based multi-omics integration
Human milk oligosaccharides are fundamental to infant health. Here the authors deploy a multi-omics systems biology approach to elucidate their biosynthetic network, including the associated enzymes and likely structures of ambiguous oligosaccharides.
- Benjamin P. Kellman
- , Anne Richelle
- & Nathan E. Lewis
-
Article
| Open Access Expanding biochemical knowledge and illuminating metabolic dark matter with ATLASx
Expanding biochemical knowledge and illuminating metabolic dark matter with ATLASx
“Mapping the dark matter of metabolism remains an open challenge that can be addressed globally and systematically by existing computational solutions. Here the authors present ATLASx, a repository of known and predicted enzymatic reaction, connecting millions of compounds to help synthetic biologists and metabolic engineers to design and explore metabolic pathways.”
- Homa MohammadiPeyhani
- , Jasmin Hafner
- & Vassily Hatzimanikatis
-
Article
| Open Access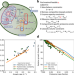 Whole-cell modeling in yeast predicts compartment-specific proteome constraints that drive metabolic strategies
Whole-cell modeling in yeast predicts compartment-specific proteome constraints that drive metabolic strategies
Metabolically active organelles compete for cytosolic space and resources during metabolism rewiring. Here, the authors develop a computational model of yeast metabolism and resource allocation to predict condition- and compartment-specific proteome constraints that govern metabolic strategies.
- Ibrahim E. Elsemman
- , Angelica Rodriguez Prado
- & Bas Teusink
-
Article
| Open Access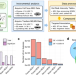 High-coverage metabolomics uncovers microbiota-driven biochemical landscape of interorgan transport and gut-brain communication in mice
High-coverage metabolomics uncovers microbiota-driven biochemical landscape of interorgan transport and gut-brain communication in mice
The gut microbiota harbours neuroactive potential with links to neurological disorders. Here, the authors apply global metabolomics with an integrated annotation strategy to comparatively profile fecal, blood serum and cerebral cortical brain tissues of eight-week-old germ-free mice vs. age-matched specific-pathogen-free mice, providing a snapshot of the metabolome status linked to the gut-brain axis.
- Yunjia Lai
- , Chih-Wei Liu
- & Kun Lu
-
Article
| Open Access Spatial localisation meets biomolecular networks
Spatial localisation meets biomolecular networks
Complex biomolecular networks are fundamental to the functioning of living systems, both at the cellular level and beyond. In this paper, the authors develop a systems framework to elucidate the interplay of networks and the spatial localisation of network components.
- Govind Menon
- & J. Krishnan
-
Article
| Open Access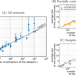 The basis of easy controllability in Boolean networks
The basis of easy controllability in Boolean networks
Boolean networks allow a simplified representation of interactions. Here, the authors systematically analyze regulation in dozens of biological Boolean networks, finding mathematical regularities that suggest biological systems could be controlled through a relatively small number of components.
- Enrico Borriello
- & Bryan C. Daniels
-
Article
| Open Access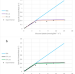 A genome-scale metabolic model of Saccharomyces cerevisiae that integrates expression constraints and reaction thermodynamics
A genome-scale metabolic model of Saccharomyces cerevisiae that integrates expression constraints and reaction thermodynamics
Formulating metabolic networks mathematically can help researchers study metabolic diseases and optimize the production of industrially important molecules. Here, the authors propose a framework that allows to model eukaryotic metabolism considering gene expression and thermodynamic constraints.
- Omid Oftadeh
- , Pierre Salvy
- & Vassily Hatzimanikatis
-
Article
| Open Access The quantitative metabolome is shaped by abiotic constraints
The quantitative metabolome is shaped by abiotic constraints
Evolution selects for the fittest but must operate within the realm of the physically possible. Here, the authors present a theoretical framework that allows them to explore how ten abiotic constraints can shape the operation, regulation, and adaptation of metabolism in E. coli.
- Amir Akbari
- , James T. Yurkovich
- & Bernhard O. Palsson
-
Article
| Open Access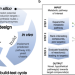 A computational workflow for the expansion of heterologous biosynthetic pathways to natural product derivatives
A computational workflow for the expansion of heterologous biosynthetic pathways to natural product derivatives
The top down cheminformatics method is usually used for the reconstitution of heterologous pathway to produce plant natural products. Here, the authors report a bottom up computational workflow for the identification of potential products and the enzymes required to make them in a noscapine pathway in yeast.
- Jasmin Hafner
- , James Payne
- & Christina Smolke
-
Article
| Open Access Protein context shapes the specificity of SH3 domain-mediated interactions in vivo
Protein context shapes the specificity of SH3 domain-mediated interactions in vivo
The SRC Homology 3 (SH3) domains mediate protein–protein interactions (PPIs). Here, the authors assess the SH3-mediated PPIs in yeast, and show that the identity of the protein itself and the position of the SH3 both affect the interaction specificity and thus the PPI-dependent cellular functions.
- Ugo Dionne
- , Émilie Bourgault
- & Christian R. Landry
-
Article
| Open Access A logical network-based drug-screening platform for Alzheimer’s disease representing pathological features of human brain organoids
A logical network-based drug-screening platform for Alzheimer’s disease representing pathological features of human brain organoids
Developing effective drugs for Alzheimer’s disease (AD), the most common cause of dementia, has been difficult because of complicated pathogenesis. Here, the authors report an efficient network-based drug-screening platform developed by integrating mathematical modeling and the pathological features of human cerebral organoids.
- Jong-Chan Park
- , So-Yeong Jang
- & Inhee Mook-Jung
-
Perspective
| Open Access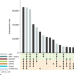 Towards a unified open access dataset of molecular interactions
Towards a unified open access dataset of molecular interactions
The IMEx consortium provides one of the largest resources of curated, experimentally verified molecular interaction data. Here, the authors review how IMEx evolved into a fundamental resource for life scientists and describe how IMEx data can support biomedical research.
- Pablo Porras
- , Elisabet Barrera
- & Sandra Orchard
-
Article
| Open Access The protein translation machinery is expressed for maximal efficiency in Escherichia coli
The protein translation machinery is expressed for maximal efficiency in Escherichia coli
The protein translation machinery is the most expensive cellular subsystem in fast growing bacteria. Providing a detailed mechanistic model for this complex system, the authors show that the translation machinery components are expressed such that their combined cost to the cell is minimal.
- Xiao-Pan Hu
- , Hugo Dourado
- & Martin J. Lercher
-
Article
| Open Access Multiplexed Optical Sensors in Arrayed Islands of Cells for multimodal recordings of cellular physiology
Multiplexed Optical Sensors in Arrayed Islands of Cells for multimodal recordings of cellular physiology
Existing fluorescent protein-based sensor measurements are limited to 4 or fewer simultaneously recorded modalities due to spectral overlap. Here the authors introduce Multiplexed Optical Sensors in Arrayed Islands of Cells (MOSAIC), which enables parallel recording of tens of physiological parameters using dense arrays of cell islands, each expressing a different fluorescent sensor.
- Christopher A. Werley
- , Stefano Boccardo
- & Adam E. Cohen
-
Article
| Open Access Sequential modification of bacterial chemoreceptors is key for achieving both accurate adaptation and high gain
Sequential modification of bacterial chemoreceptors is key for achieving both accurate adaptation and high gain
Bacterial chemoreceptors have multiple methylation sites, but whether the order of methylation matters is unclear. Here, the authors show that sequentially ordered methylation is critical for perfect adaptation and for attenuating the trade-off between accurate adaptation and high response gain.
- Bernardo A. Mello
- , Anderson B. Beserra
- & Yuhai Tu
-
Article
| Open Access A biochemically-interpretable machine learning classifier for microbial GWAS
A biochemically-interpretable machine learning classifier for microbial GWAS
Current machine learning classifiers have been applied to whole-genome sequencing data to identify determinants of antimicrobial resistance, but they lack interpretability. Here the authors present a metabolic machine learning classifier that uses flux balance analysis to estimate the biochemical effects of alleles.
- Erol S. Kavvas
- , Laurence Yang
- & Bernhard O. Palsson
-
Article
| Open Access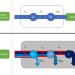 An analytical theory of balanced cellular growth
An analytical theory of balanced cellular growth
Genome-scale models of microbial metabolism largely ignore reaction kinetics. Here, the authors develop a general mathematical framework for modeling cellular growth with explicit non-linear reaction kinetics and use it to glean insights into the principles of cellular resource allocation and growth.
- Hugo Dourado
- & Martin J. Lercher
-
Article
| Open Access Discovering the genes mediating the interactions between chronic respiratory diseases in the human interactome
Discovering the genes mediating the interactions between chronic respiratory diseases in the human interactome
Complex diseases often share genetic determinants and symptoms, but the mechanistic basis of disease interactions remains elusive. Here, the authors propose a network topological measure to identify proteins linking complex diseases in the interactome, and identify mediators between COPD and asthma.
- Enrico Maiorino
- , Seung Han Baek
- & Amitabh Sharma
-
Article
| Open Access Genome-scale reconstructions of the mammalian secretory pathway predict metabolic costs and limitations of protein secretion
Genome-scale reconstructions of the mammalian secretory pathway predict metabolic costs and limitations of protein secretion
The secretory pathway is used in the production of most biopharmaceuticals, but the associated biosynthetic costs are little understood. Here, the authors integrate the core secretory pathway into genome-scale metabolic models of human, mouse, and CHO cells, enabling in silico analysis.
- Jahir M. Gutierrez
- , Amir Feizi
- & Nathan E. Lewis
-
Article
| Open Access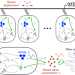 Emergence of collective oscillations in adaptive cells
Emergence of collective oscillations in adaptive cells
There are many examples of cell populations exhibiting density-dependent collective oscillatory behaviour. Here, the authors show that sustained collective oscillations emerge when cells anticipate variation in signal and attempt to amplify it, a property that can be linked to adaptation.
- Shou-Wen Wang
- & Lei-Han Tang
-
Article
| Open Access Microbial carbon use efficiency predicted from genome-scale metabolic models
Microbial carbon use efficiency predicted from genome-scale metabolic models
Microbial respiration releases carbon from the soil. Here, the authors estimate bacterial carbon use efficiency in soils for over 200 species using constraint-based modeling, incorporate the values into an ecosystem model, and find that shifts in community composition may impact carbon storage.
- Mustafa Saifuddin
- , Jennifer M. Bhatnagar
- & Adrien C. Finzi
-
Article
| Open Access A consensus S. cerevisiae metabolic model Yeast8 and its ecosystem for comprehensively probing cellular metabolism
A consensus S. cerevisiae metabolic model Yeast8 and its ecosystem for comprehensively probing cellular metabolism
Genome-scale metabolic models provide a platform to study metabolism through simulations and analysis of omics data. Here the authors introduce Yeast8 with its model ecosystem, a comprehensive computational resource for simulating the metabolism of Saccharomyces cerevisiae.
- Hongzhong Lu
- , Feiran Li
- & Jens Nielsen
-
Article
| Open Access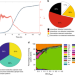 Regulatory mechanisms underlying coordination of amino acid and glucose catabolism in Escherichia coli
Regulatory mechanisms underlying coordination of amino acid and glucose catabolism in Escherichia coli
Bacteria must adapt their metabolism in the face of dynamically changing nutrient availability. Here, using their constraint-based modeling approach the authors analyze E. coli exometabolome data during growth in complex medium, revealing temporal coordination of glucose and amino acid catabolism.
- Mattia Zampieri
- , Manuel Hörl
- & Uwe Sauer
-
Article
| Open Access Estimating dispensable content in the human interactome
Estimating dispensable content in the human interactome
The fraction of protein-protein interactions (PPIs) that can be disrupted without fitness effect is unknown. Here, the authors model how disease-causing mutations and common mutations carried by healthy people perturb the interactome, and estimate that <20% of human PPIs are completely dispensable.
- Mohamed Ghadie
- & Yu Xia

 Network-based elucidation of colon cancer drug resistance mechanisms by phosphoproteomic time-series analysis
Network-based elucidation of colon cancer drug resistance mechanisms by phosphoproteomic time-series analysis