Article
|
Open Access
Featured
-
-
Article
| Open Access Genomic insights unveil the plasmid transfer mechanism and epidemiology of hypervirulent Klebsiella pneumoniae in Vietnam
Genomic insights unveil the plasmid transfer mechanism and epidemiology of hypervirulent Klebsiella pneumoniae in Vietnam
Hypervirulent Klebsiella pneumoniae (hvKp) is a significant cause of severe community-acquired infection, primarily in Asia. Here, the authors characterise the genetic profile, phylogenetic structure, and plasmid features of hvKp in Vietnam.
- Quynh Nguyen
- , Nguyen Yen Thi Phuong
- & Duy Thanh Pham
-
Article
| Open Access Dynamic microfluidic single-cell screening identifies pheno-tuning compounds to potentiate tuberculosis therapy
Dynamic microfluidic single-cell screening identifies pheno-tuning compounds to potentiate tuberculosis therapy
Tuberculosis is a major global health threat. Here, the authors develop a single-cell drug discovery approach and identify a compound that tunes bacterial phenotypic variation. This enhances the activity of anti-tubercular drugs against the pathogen.
- Maxime Mistretta
- , Mena Cimino
- & Giulia Manina
-
Article
| Open Access Evolution of triclosan resistance modulates bacterial permissiveness to multidrug resistance plasmids and phages
Evolution of triclosan resistance modulates bacterial permissiveness to multidrug resistance plasmids and phages
In this work, Yang et al. provide evidence of triclosan exposure resulting in increased evolvability of K. pneumoniae in experimental evolution studies. They utilize sequencing and transcriptomics to explore the chromosomally and horizontally acquired antimicrobial resistance mechanisms.
- Qiu E. Yang
- , Xiaodan Ma
- & Timothy R. Walsh
-
Article
| Open Access A secondary mechanism of action for triazole antifungals in Aspergillus fumigatus mediated by hmg1
A secondary mechanism of action for triazole antifungals in Aspergillus fumigatus mediated by hmg1
Triazole antifungals are widely used and exert their action by inhibiting ergosterol biosynthesis. Here, Rybak et al show that these drugs both inhibit ergosterol biosynthesis and induce accumulation of pathway intermediates that directly induce inhibition of sterol synthesis.
- Jeffrey M. Rybak
- , Jinhong Xie
- & Jarrod R. Fortwendel
-
Article
| Open Access Colonisation of hospital surfaces from low- and middle-income countries by extended spectrum β-lactamase- and carbapenemase-producing bacteria
Colonisation of hospital surfaces from low- and middle-income countries by extended spectrum β-lactamase- and carbapenemase-producing bacteria
In hospitals, surfaces present as a reservoir for bacteria pathogens, potentially leading to nosocomial infections. In this work, authors aim to profile extended-spectrum β lactamase- and carbapenemase-carrying bacterial species colonising neonatal hospital wards and causing neonatal sepsis.
- Maria Nieto-Rosado
- , Kirsty Sands
- & Timothy R. Walsh
-
Article
| Open Access Mutations in the efflux pump regulator MexZ shift tissue colonization by Pseudomonas aeruginosa to a state of antibiotic tolerance
Mutations in the efflux pump regulator MexZ shift tissue colonization by Pseudomonas aeruginosa to a state of antibiotic tolerance
Mutations in mexZ, encoding a negative regulator of efflux pump genes, are frequently acquired by Pseudomonas aeruginosa during early lung infection, but do not confer high antibiotic resistance as measured in lab tests. Here, Laborda et al. show that mexZ mutations affect quorum sensing pathways, thus promoting tissue invasiveness and protecting bacteria from the action of antibiotics within tissues.
- Pablo Laborda
- , Signe Lolle
- & Helle Krogh Johansen
-
Article
| Open Access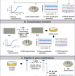 Bacteria can compensate the fitness costs of amplified resistance genes via a bypass mechanism
Bacteria can compensate the fitness costs of amplified resistance genes via a bypass mechanism
Antibiotic heteroresistance, in which a susceptible bacterial population includes a small resistant subpopulation, can arise by tandem amplification of resistance genes, which often carry fitness costs. Here, Pal and Andersson show that these fitness costs can be ameliorated by the acquisition of compensatory mutations and a reduction in copy number of the resistance genes.
- Ankita Pal
- & Dan I. Andersson
-
Article
| Open Access Characterisation of colistin resistance in Gram-negative microbiota of pregnant women and neonates in Nigeria
Characterisation of colistin resistance in Gram-negative microbiota of pregnant women and neonates in Nigeria
Here, the authors report the results of a BARNARDS sub-study identifying a 1% mobile colistin resistance gene (mcr) carriage rate in around 5000 rectal swabs from mothers and neonates across Nigeria, of which 90% were mcr-10 (mostly Enterobacter spp.) and 10% were mcr-1 and mcr9.
- E. A. R. Portal
- , K. Sands
- & O. B. Spiller
-
Article
| Open Access Deep learning model for personalized prediction of positive MRSA culture using time-series electronic health records
Deep learning model for personalized prediction of positive MRSA culture using time-series electronic health records
Identification of patients at high risk of methicillin-resistant Staphylococcus aureus (MRSA) infection could improve treatment outcomes by optimising antimicrobial therapy. Here the authors develop a deep learning model that uses electronic health record data from the United States to predict MRSA culture positivity.
- Masayuki Nigo
- , Laila Rasmy
- & Degui Zhi
-
Article
| Open Access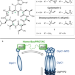 Homo-BacPROTAC-induced degradation of ClpC1 as a strategy against drug-resistant mycobacteria
Homo-BacPROTAC-induced degradation of ClpC1 as a strategy against drug-resistant mycobacteria
Antimicrobial resistance is a global health threat and the development of alternative strategies to overcome it is of high interest. Here, the authors report proteolysis targeting chimeras active in bacteria (BacPROTACs) that bind to ClpC1, a component of the mycobacterial protein degradation machinery, and apply them for targeting a range of mycobacterial strains, including antibiotic-resistant ones.
- Lukas Junk
- , Volker M. Schmiedel
- & Guido Boehmelt
-
Article
| Open Access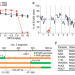 Deaggregation of mutant Plasmodium yoelii de-ubiquitinase UBP1 alters MDR1 localization to confer multidrug resistance
Deaggregation of mutant Plasmodium yoelii de-ubiquitinase UBP1 alters MDR1 localization to confer multidrug resistance
Here, the authors show that two mutations in the Plasmodium de-ubiquitinase UBP1 alter the ubiquitination level, membrane localization, and ligand transport direction of multidrug resistance transporter 1 (MDR1), leading to multiple drug resistances.
- Ruixue Xu
- , Lirong Lin
- & Jian Li
-
Article
| Open Access The plasmidome associated with Gram-negative bloodstream infections: A large-scale observational study using complete plasmid assemblies
The plasmidome associated with Gram-negative bloodstream infections: A large-scale observational study using complete plasmid assemblies
Plasmids carry antimicrobial resistance genes and contribute to the rapid dissemination of resistance. Here, the authors sequence 1,880 complete plasmids from 738 isolates from bloodstream infections, shedding light on the links between plasmid types, bacterial hosts and antimicrobial resistance.
- Samuel Lipworth
- , William Matlock
- & Nicole Stoesser
-
Article
| Open Access Exploiting lung adaptation and phage steering to clear pan-resistant Pseudomonas aeruginosa infections in vivo
Exploiting lung adaptation and phage steering to clear pan-resistant Pseudomonas aeruginosa infections in vivo
In this work, authors utilise a pan-resistant Pseudomonas aeruginosa in vivo infection model to demonstrate antibiotic re-sensitisation with bacteriophage therapy.
- Eleri A. Ashworth
- , Rosanna C. T. Wright
- & Joanne L. Fothergill
-
Article
| Open Access Duplicated antibiotic resistance genes reveal ongoing selection and horizontal gene transfer in bacteria
Duplicated antibiotic resistance genes reveal ongoing selection and horizontal gene transfer in bacteria
Mobile genetic elements can promote the duplication of antibiotic resistance genes which may in turn accelerate the evolution of resistance to new drugs. Here, the authors show that duplicated antibiotic resistance genes are enriched in bacterial isolates from environments associated with rampant antibiotic use.
- Rohan Maddamsetti
- , Yi Yao
- & Lingchong You
-
Article
| Open Access Hormonal steroids induce multidrug resistance and stress response genes in Neisseria gonorrhoeae by binding to MtrR
Hormonal steroids induce multidrug resistance and stress response genes in Neisseria gonorrhoeae by binding to MtrR
Transcriptional regulator MtrR inhibits the expression of the multidrug efflux pump operon mtrCDE in Neisseria gonorrhoeae. Here, Hooks et al. show that hormonal steroids bind to MtrR and decrease its affinity for cognate promoters, thus leading to increased mtrCDE expression and enhanced antimicrobial resistance.
- Grace M. Hooks
- , Julio C. Ayala
- & Richard G. Brennan
-
Article
| Open Access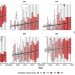 Global surveillance of antimicrobial resistance in food animals using priority drugs maps
Global surveillance of antimicrobial resistance in food animals using priority drugs maps
Monitoring antimicrobial resistance in food animals is challenging due to limited surveillance systems. Here, the authors combine data from point prevalence surveys in lower- and middle-income settings to map resistance to seven antimicrobials and predict which are likely to exceed key resistance thresholds.
- Cheng Zhao
- , Yu Wang
- & Thomas P. Van Boeckel
-
Article
| Open Access Rapid and visual identification of β-lactamase subtypes for precision antibiotic therapy
Rapid and visual identification of β-lactamase subtypes for precision antibiotic therapy
The rapid identification of drug-resistant bacteria is vital for effective treatment and to avoid antibiotic misuse. Here authors report a paper-based sensor which utilises chromogenic carbapenem and cephalosporin substrates for the identification and discrimination of β-lactamase subtypes.
- Wenshuai Li
- , Jingqi Li
- & Dingbin Liu
-
Article
| Open Access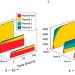 Host- plasmid network structure in wastewater is linked to antimicrobial resistance genes
Host- plasmid network structure in wastewater is linked to antimicrobial resistance genes
Authors apply theory and microbial ecology modelling to a wastewater sample, and show that antimicrobial resistance carrying plasmids interact with a higher number and more diverse range of bacteria than plasmids that do not carry resistance genes.
- Alice Risely
- , Arthur Newbury
- & Dirk Sanders
-
Article
| Open Access Quantitative measurement of antibiotic resistance in Mycobacterium tuberculosis reveals genetic determinants of resistance and susceptibility in a target gene approach
Quantitative measurement of antibiotic resistance in Mycobacterium tuberculosis reveals genetic determinants of resistance and susceptibility in a target gene approach
Molecular diagnostics for tuberculosis have focused on predicting drug susceptibilities in a binary manner (i.e., strains are either susceptible or resistant). Here, CRyPTIC Consortium researchers use whole genome sequencing and a quantitative assay to identify associations between genomic mutations and minimum inhibitory concentrations in over 15,000 Mycobacterium tuberculosis clinical isolates.
- Ivan Barilar
- , Simone Battaglia
- & Baoli Zhu
-
Article
| Open Access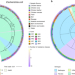 Convergence of resistance and evolutionary responses in Escherichia coli and Salmonella enterica co-inhabiting chicken farms in China
Convergence of resistance and evolutionary responses in Escherichia coli and Salmonella enterica co-inhabiting chicken farms in China
Bacteria in the same environment can share genetic material but the extent to which this influences development of antimicrobial resistance is unclear. Here, the authors investigate the evidence for co-evolution of antimicrobial resistance in bacteria found coexisting in animals and the environment in chicken farms and slaughterhouses in China.
- Michelle Baker
- , Xibin Zhang
- & Tania Dottorini
-
Article
| Open Access The antibiotic resistance reservoir of the lung microbiome expands with age in a population of critically ill patients
The antibiotic resistance reservoir of the lung microbiome expands with age in a population of critically ill patients
Here, by performing tracheal aspirate RNA sequencing of critically ill patients, the authors find that older age associates with a greater number of detectably expressed antimicrobial resistance genes in the lower respiratory tract microbiome.
- Victoria T. Chu
- , Alexandra Tsitsiklis
- & Charles R. Langelier
-
Article
| Open Access Within-host genetic diversity of extended-spectrum beta-lactamase-producing Enterobacterales in long-term colonized patients
Within-host genetic diversity of extended-spectrum beta-lactamase-producing Enterobacterales in long-term colonized patients
The diversity of ESBL-producing Klebsiella pneumoniae species complex and ESBL-Escherichia coli within patients is low and colonization with the same strain may persist for long periods. Authors utilise clinical and microbiological data from electronic health records to investigate genetic diversity of colonizing and infecting strains.
- Lisandra Aguilar-Bultet
- , Ana B. García-Martín
- & Sarah Tschudin-Sutter
-
Article
| Open Access ANCA: artificial nucleic acid circuit with argonaute protein for one-step isothermal detection of antibiotic-resistant bacteria
ANCA: artificial nucleic acid circuit with argonaute protein for one-step isothermal detection of antibiotic-resistant bacteria
Antibiotic-resistant bacteria pose a growing threat to global health. Here, the authors present an artificial nucleic acid circuit with argonaute protein (ANCA) for one-step, amplification-free, and isothermal detection of carbapenemase-producing Klebsiella pneumoniae.
- Hyowon Jang
- , Jayeon Song
- & Taejoon Kang
-
Article
| Open Access National genomic surveillance integrating standardized quantitative susceptibility testing clarifies antimicrobial resistance in Enterobacterales
National genomic surveillance integrating standardized quantitative susceptibility testing clarifies antimicrobial resistance in Enterobacterales
Kayama et al. present a blueprint for a national genomic surveillance study that conducts genome sequencing of thousands of strains, integrates standardized quantitative antimicrobial susceptibility testing, and characterizes antimicrobial resistance determinants.
- Shizuo Kayama
- , Koji Yahara
- & Motoyuki Sugai
-
Article
| Open Access Klebsiella pneumoniae clinical isolates with features of both multidrug-resistance and hypervirulence have unexpectedly low virulence
Klebsiella pneumoniae clinical isolates with features of both multidrug-resistance and hypervirulence have unexpectedly low virulence
Convergent strains, those containing characteristics of both multidrug-resistant & hypervirulent Klebsiella pneumoniae, are a global threat to public health. In this work, authors analyse convergent isolates from the United States and reveal unexpectantly low virulence.
- Travis J. Kochan
- , Sophia H. Nozick
- & Alan R. Hauser
-
Article
| Open Access Global pathogenomic analysis identifies known and candidate genetic antimicrobial resistance determinants in twelve species
Global pathogenomic analysis identifies known and candidate genetic antimicrobial resistance determinants in twelve species
A global analysis of antimicrobial resistance (AMR) across 27,155 genomes and 69 drugs reveals patterns in AMR gene transfer between species and identifies 142 AMR gene candidates, two of which were tested and confirmed as contributing to AMR.
- Jason C. Hyun
- , Jonathan M. Monk
- & Bernhard O. Palsson
-
Article
| Open Access Clinically relevant antibiotic resistance genes are linked to a limited set of taxa within gut microbiome worldwide
Clinically relevant antibiotic resistance genes are linked to a limited set of taxa within gut microbiome worldwide
Antimicrobial resistance genes (ARGs) in commensal gut bacteria may act as a reservoir for acquisition by pathogens. Here, the authors assess the distribution and transfer potential of ARGs in gut microbiomes and find that clinically important ARGs are taxonomically restricted despite being associated with mobile plasmids
- Peter J. Diebold
- , Matthew W. Rhee
- & Ilana L. Brito
-
Article
| Open Access Staphylococcus aureus sacculus mediates activities of M23 hydrolases
Staphylococcus aureus sacculus mediates activities of M23 hydrolases
In this work, the authors provide structural insights into the interaction of two evolutionarily related peptidoglycan hydrolases, lysostaphin and LytM with S. aureus sacculus, and propose a model in which PG crosslinking affects their activity differently.
- Alicja Razew
- , Cedric Laguri
- & Jean-Pierre Simorre
-
Article
| Open Access Assessing the global risk of typhoid outbreaks caused by extensively drug resistant Salmonella Typhi
Assessing the global risk of typhoid outbreaks caused by extensively drug resistant Salmonella Typhi
Extensively drug resistant (XDR) typhoid fever is an emerging global health threat. This study compares data on air travel patterns and typhoid incidence to identify countries at high risk for XDR typhoid outbreaks.
- Joseph Walker
- , Chrispin Chaguza
- & Virginia E. Pitzer
-
Article
| Open Access Deep mutational scanning reveals the molecular determinants of RNA polymerase-mediated adaptation and tradeoffs
Deep mutational scanning reveals the molecular determinants of RNA polymerase-mediated adaptation and tradeoffs
Mutations in an RNA polymerase fragment, frequently found in lab adaptation, cluster in two modules favoring growth or maintenance via loss of interactions. Combining mutations in both modules enhances both traits, promoting compensatory evolution.
- Alaksh Choudhury
- , Benoit Gachet
- & Olivier Tenaillon
-
Perspective
| Open Access The uncertain role of substandard and falsified medicines in the emergence and spread of antimicrobial resistance
The uncertain role of substandard and falsified medicines in the emergence and spread of antimicrobial resistance
Substandard and falsified medicines are a problem, particularly in low- and middle-income countries, and effects on antimicrobial resistance development aren’t well understood. Here, the authors discuss mechanisms by which they can increase or decrease levels of resistance and the need for improved data collection and analytical approaches.
- Sean Cavany
- , Stella Nanyonga
- & Ben S. Cooper
-
Article
| Open Access Bacterial motility can govern the dynamics of antibiotic resistance evolution
Bacterial motility can govern the dynamics of antibiotic resistance evolution
In nature, bacteria experience gradients of antibiotics, but we know little about how such heterogeneity affects bacterial adaptation. Piskovsky and Oliveira develop quantitative models of bacterial adaptation in antibiotic landscapes and find that bacterial motility can govern the spatiotemporal dynamics of antibiotic resistance evolution.
- Vit Piskovsky
- & Nuno M. Oliveira
-
Article
| Open Access Metallo-sideromycin as a dual functional complex for combating antimicrobial resistance
Metallo-sideromycin as a dual functional complex for combating antimicrobial resistance
Here, the authors utilise cefiderocol, a sideromycin, in complex with colloidal bismuth citrate, to demonstrate antimicrobial efficacy against Pseudomonas aeruginosa in vivo.
- Chenyuan Wang
- , Yushan Xia
- & Hongzhe Sun
-
Article
| Open Access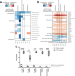 Antibiotics promote intestinal growth of carbapenem-resistant Enterobacteriaceae by enriching nutrients and depleting microbial metabolites
Antibiotics promote intestinal growth of carbapenem-resistant Enterobacteriaceae by enriching nutrients and depleting microbial metabolites
Broad-spectrum antibiotics can kill harmless bacteria in our intestine, thus facilitating invasion by antibiotic-resistant bacteria such as carbapenem-resistant Enterobacteriaceae (CRE). Here, Yip et al. show that killing gut bacteria with antibiotics leads to enrichment of nutrients and depletion of inhibitory microbial metabolites, which overall potentiates CRE growth.
- Alexander Y. G. Yip
- , Olivia G. King
- & Julie A. K. McDonald
-
Article
| Open Access Profiling cell envelope-antibiotic interactions reveals vulnerabilities to β-lactams in a multidrug-resistant bacterium
Profiling cell envelope-antibiotic interactions reveals vulnerabilities to β-lactams in a multidrug-resistant bacterium
The bacterial pathogen Burkholderia cenocepacia and related species are often multidrug resistant because their cell envelope restricts antibiotic penetration. Here, Hogan et al systematically identify genes associated with resistance and susceptibility to cell envelope-targeting antibiotics, providing insights into underlying mechanisms and suggesting avenues for development of improved antibacterial therapies.
- Andrew M. Hogan
- , A. S. M. Zisanur Rahman
- & Silvia T. Cardona
-
Article
| Open Access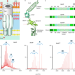 Conformational restriction shapes the inhibition of a multidrug efflux adaptor protein
Conformational restriction shapes the inhibition of a multidrug efflux adaptor protein
Multidrug efflux protein pumps are key players in bacterial antimicrobial resistance. Here, the authors show how dynamics of a periplasmic pump component can be targeted for efflux inhibition.
- Benjamin Russell Lewis
- , Muhammad R. Uddin
- & Eamonn Reading
-
Article
| Open Access Structural insights into the mechanism of overcoming Erm-mediated resistance by macrolides acting together with hygromycin-A
Structural insights into the mechanism of overcoming Erm-mediated resistance by macrolides acting together with hygromycin-A
The authors discovered that hygromycin A not only enhances the cell-killing properties of macrolides but also renders them active against resistant bacteria. The provided structures of antibiotic pairs in complex with WT and macrolide-resistant ribosomes rationalize binding cooperativity of these drugs.
- Chih-Wei Chen
- , Nadja Leimer
- & Maxim S. Svetlov
-
Article
| Open Access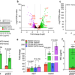 Decoding a cryptic mechanism of metronidazole resistance among globally disseminated fluoroquinolone-resistant Clostridioides difficile
Decoding a cryptic mechanism of metronidazole resistance among globally disseminated fluoroquinolone-resistant Clostridioides difficile
Detection of resistance to the antibiotic metronidazole in C. difficile often requires the presence of heme in the media, for unclear reasons. Here, the authors show that most metronidazole-resistant strains carry a mutation that promotes expression of a heme-dependent enzyme that degrades nitroimidazoles, and the mutation often co-occurs with an amino-acid substitution in DNA gyrase that confers resistance to another class of antibiotics, fluoroquinolones.
- Abiola O. Olaitan
- , Chetna Dureja
- & Julian G. Hurdle
-
Article
| Open Access Mixed strain pathogen populations accelerate the evolution of antibiotic resistance in patients
Mixed strain pathogen populations accelerate the evolution of antibiotic resistance in patients
Here, Caballero et al. provide an in depth characterisation of patients colonized with single or mixed strains of Pseudomonas aeruginosa to demonstrate the impact of within-host diversity on the development of antibiotic resistance.
- Julio Diaz Caballero
- , Rachel M. Wheatley
- & R. Craig MacLean
-
Article
| Open Access Molecular mechanism of plasmid-borne resistance to sulfonamide antibiotics
Molecular mechanism of plasmid-borne resistance to sulfonamide antibiotics
Bacterial resistance to sulfonamide antibiotics (sulfas) is mediated by acquisition of sul genes, which encode sulfa-insensitive versions of the target enzyme, dihydropteroate synthase. Here, Venkatesan et al. study Sul enzymes using biochemical, structural, mutational and functional analyses, revealing the molecular basis for Sul-mediated drug resistance.
- Meenakshi Venkatesan
- , Michael Fruci
- & Alexei Savchenko
-
Article
| Open Access Architecture of a complete Bce-type antimicrobial peptide resistance module
Architecture of a complete Bce-type antimicrobial peptide resistance module
Here the authors present cryo-EM structures of the Bacillus subtilis bacitracin resistance module BceABS, providing insight into the assembly and conformational dynamics within the complex, and coregulation between individual protein components.
- Natasha L. George
- & Benjamin J. Orlando
-
Article
| Open Access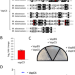 Mycobacterium abscessus VapC5 toxin potentiates evasion of antibiotic killing by ribosome overproduction and activation of multiple resistance pathways
Mycobacterium abscessus VapC5 toxin potentiates evasion of antibiotic killing by ribosome overproduction and activation of multiple resistance pathways
Mycobacterium abscessus (Mab) infections are difficult to clear with antibiotics. Here the authors show that clinical Mab strains can acquire a toxin-antitoxin system that enhances survival upon treatment with current first-line antibiotics through depletion of tRNASerCGA and subsequent ribosome overproduction.
- Eduardo A. Troian
- , Heather M. Maldonado
- & Nancy A. Woychik
-
Article
| Open Access Timing of antibiotic administration determines the spread of plasmid-encoded antibiotic resistance during microbial range expansion
Timing of antibiotic administration determines the spread of plasmid-encoded antibiotic resistance during microbial range expansion
Plasmids are the main vector by which antibiotic resistance is transferred between bacterial cells within surface-associated communities. Here, Ma et al. show that plasmid spread peaks at intermediate antibiotic administration times, when the intermixing of plasmid donors and potential recipients is maximal.
- Yinyin Ma
- , Josep Ramoneda
- & David R. Johnson
-
Article
| Open Access Efflux pump gene amplifications bypass necessity of multiple target mutations for resistance against dual-targeting antibiotic
Efflux pump gene amplifications bypass necessity of multiple target mutations for resistance against dual-targeting antibiotic
Antibiotics that attack multiple targets in bacteria are thought to reduce the frequency of resistance. The authors show that genomic amplifications of poorly characterized efflux pumps can instead lead to high-frequency antibiotic cross-resistance.
- Kalinga Pavan T. Silva
- , Ganesh Sundar
- & Anupama Khare
-
Article
| Open Access A point mutation in recC associated with subclonal replacement of carbapenem-resistant Klebsiella pneumoniae ST11 in China
A point mutation in recC associated with subclonal replacement of carbapenem-resistant Klebsiella pneumoniae ST11 in China
Authors carry out a genomic analysis of carbapenem-resistant Klebsiella pneumoniae bloodstream isolates, noting subclonal expansion, associated with the emergence of a hypervirulent subpopulation.
- Kai Zhou
- , Chun-Xu Xue
- & Yonghong Xiao
-
Article
| Open Access Transcription tuned by S-nitrosylation underlies a mechanism for Staphylococcus aureus to circumvent vancomycin killing
Transcription tuned by S-nitrosylation underlies a mechanism for Staphylococcus aureus to circumvent vancomycin killing
Antibiotic resistance in Staphylococcus aureus is increasingly emerging. Here, Shu et al demonstrate that transcriptional regulation by S-nitrosylation underlies vancomycin resistance.
- Xueqin Shu
- , Yingying Shi
- & Baolin Sun
-
Article
| Open Access Associate toxin-antitoxin with CRISPR-Cas to kill multidrug-resistant pathogens
Associate toxin-antitoxin with CRISPR-Cas to kill multidrug-resistant pathogens
CRISPR-regulated toxin-antitoxin (CreTA), safeguards CRISPR-Cas immune systems. Here the authors characterize a bacterial CreTA and use this to generate a proof-of-concept antimicrobial strategy, ATTACK, which associates TA and CRISPR-Cas to kill multidrug resistant pathogens.
- Rui Wang
- , Xian Shu
- & Ming Li
-
Article
| Open Access Collateral sensitivity profiling in drug-resistant Escherichia coli identifies natural products suppressing cephalosporin resistance
Collateral sensitivity profiling in drug-resistant Escherichia coli identifies natural products suppressing cephalosporin resistance
Collateral sensitivity (CS), whereby resistance to one drug is accompanied by increased sensitivity to another, provides new opportunities for antimicrobial drug discovery. Here, Liu et al. screen large chemical libraries across 29 drug-resistant E. coli strains to identify multiple CS interactions, including natural products with potent CS activities against cephalosporin-resistant strains.
- Dennis Y. Liu
- , Laura Phillips
- & Roger G. Linington
-
Article
| Open Access The relative transmission fitness of multidrug-resistant Mycobacterium tuberculosis in a drug resistance hotspot
The relative transmission fitness of multidrug-resistant Mycobacterium tuberculosis in a drug resistance hotspot
Geographical hotspots with high frequency of multi-drug resistant tuberculosis (MDR-TB) have been observed in several locations, such as the country of Georgia. Here, the authors analyse genomic sequences from tuberculosis bacteria isolated from Georgia to show that the transmission fitness of MDR-TB strains is heterogeneous, and highly drug-resistant and transmissible isolates contribute to the emergence and maintenance of MDR-TB hotspots.
- Chloé Loiseau
- , Etthel M. Windels
- & Sebastien Gagneux

 β-lactamase expression induces collateral sensitivity in Escherichia coli
β-lactamase expression induces collateral sensitivity in Escherichia coli