Article
|
Open Access
Featured
-
-
Article
| Open Access A large-scale binding and functional map of human RNA-binding proteins
A large-scale binding and functional map of human RNA-binding proteins
A combination of five assays is used to produce a catalogue of RNA elements to which RNA-binding proteins bind in human cells.
- Eric L. Van Nostrand
- , Peter Freese
- & Gene W. Yeo
-
Letter |
 Spliceosomal disruption of the non-canonical BAF complex in cancer
Spliceosomal disruption of the non-canonical BAF complex in cancer
A range of SF3B1 mutations promote tumorigenesis through the repression of BRD9, a core component of the non-canonical BAF complex, and correcting BRD9 mis-splicing in these SF3B1-mutant cells suppresses tumour growth.
- Daichi Inoue
- , Guo-Liang Chew
- & Robert K. Bradley
-
Letter |
 Prespliceosome structure provides insights into spliceosome assembly and regulation
Prespliceosome structure provides insights into spliceosome assembly and regulation
The cryo-electron microscopy structure of the Saccharomyces cerevisiae prespliceosome provides insights into splice-site selection and early spliceosome assembly events.
- Clemens Plaschka
- , Pei-Chun Lin
- & Kiyoshi Nagai
-
Letter |
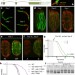 Splicing factor 1 modulates dietary restriction and TORC1 pathway longevity in C. elegans
Splicing factor 1 modulates dietary restriction and TORC1 pathway longevity in C. elegans
Precursor mRNA splicing homeostasis is a biomarker and predictor of life expectancy in Caenorhabditis elegans and defects in global pre-mRNA splicing associated with age are reduced by dietary restriction via splicing factor 1.
- Caroline Heintz
- , Thomas K. Doktor
- & William B. Mair
-
Letter |
 m6A potentiates Sxl alternative pre-mRNA splicing for robust Drosophila sex determination
m6A potentiates Sxl alternative pre-mRNA splicing for robust Drosophila sex determination
Two complementary studies describe how the pervasive N6-methyladenosine modification in mRNA can affect Drosophila sex determination, neuronal function and behaviour.
- Irmgard U. Haussmann
- , Zsuzsanna Bodi
- & Matthias Soller
-
Letter |
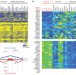 MBNL proteins repress ES-cell-specific alternative splicing and reprogramming
MBNL proteins repress ES-cell-specific alternative splicing and reprogramming
This study identifies MBNL proteins as negative regulators of alternative splicing events that are differentially regulated between ES cells and other cell types; several lines of evidence show that these proteins repress an ES cell alternative splicing program and the reprogramming of somatic cells to induced pluripotent stem cells.
- Hong Han
- , Manuel Irimia
- & Benjamin J. Blencowe
-
Letter |
 Cryptic peroxisomal targeting via alternative splicing and stop codon read-through in fungi
Cryptic peroxisomal targeting via alternative splicing and stop codon read-through in fungi
Translocation of glycolytic enzymes to peroxisomes in fungi suggests broader metabolic role for this organelle.
- Johannes Freitag
- , Julia Ast
- & Michael Bölker
-
Article |
 CTCF-promoted RNA polymerase II pausing links DNA methylation to splicing
CTCF-promoted RNA polymerase II pausing links DNA methylation to splicing
- Sanjeev Shukla
- , Ersen Kavak
- & Shalini Oberdoerffer
-
Letter |
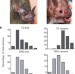 Ganglion-specific splicing of TRPV1 underlies infrared sensation in vampire bats
Ganglion-specific splicing of TRPV1 underlies infrared sensation in vampire bats
- Elena O. Gracheva
- , Julio F. Cordero-Morales
- & David Julius
-
Article |
 Role of the ubiquitin-like protein Hub1 in splice-site usage and alternative splicing
Role of the ubiquitin-like protein Hub1 in splice-site usage and alternative splicing
- Shravan Kumar Mishra
- , Tim Ammon
- & Stefan Jentsch
-
Letter |
 A methyl transferase links the circadian clock to the regulation of alternative splicing
A methyl transferase links the circadian clock to the regulation of alternative splicing
Various biological processes are entrained by the day–night cycle to occur at a specific time of day. One way the circadian system exerts these effects is through post-transcriptional regulation. These authors show that a protein that transfers methyl groups onto several spliceosome subunits, PRMT5, is regulated by the light–dark cycle. Methylation of these subunits affects alternative splicing of some genes, thus making them subject to circadian control.
- Sabrina E. Sanchez
- , Ezequiel Petrillo
- & Marcelo J. Yanovsky
-
Article |
 Deciphering the splicing code
Deciphering the splicing code
The coding capacity of the genome is greatly expanded by the process of alternative splicing, which enables a single gene to produce more than one distinct protein. Can the expression of these different proteins be predicted from sequence data? Here, modelling based on information theory has been used to develop a 'splicing code', which can predict, with good accuracy, tissue-dependent changes in alternative splicing.
- Yoseph Barash
- , John A. Calarco
- & Brendan J. Frey
-
Review Article |
 Expansion of the eukaryotic proteome by alternative splicing
Expansion of the eukaryotic proteome by alternative splicing
- Timothy W. Nilsen
- & Brenton R. Graveley

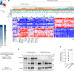 RBFOX2 modulates a metastatic signature of alternative splicing in pancreatic cancer
RBFOX2 modulates a metastatic signature of alternative splicing in pancreatic cancer