Abstract
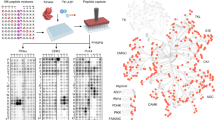
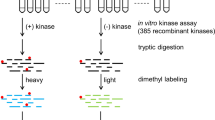
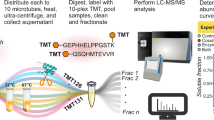
Elucidating kinase-substrate relationships is critical for understanding how phosphorylation affects signal transduction and regulatory cascades. Using the α catalytic subunit of protein kinase CK2 (CK2α) as a paradigm, we developed an in-gel method to facilitate identification of physiologic kinase substrates. In this approach, the roles of kinase and substrate in a classic in-gel kinase assay are reversed. In the reverse in-gel kinase assay (RIKA), a kinase is copolymerized in a denaturing polyacrylamide gel used to resolve a tissue or cell protein extract. Restoration of kinase activity and substrate structure followed by an in situ kinase reaction and mass spectrometric analyses results in identification of potential kinase substrates. We demonstrate that this method can be used to profile both known and novel human and mouse substrates of CK2α and cAMP-dependent protein kinase (PKA). Using widely available straightforward technology, the RIKA has the potential to facilitate discovery of physiologic kinase substrates in any biological system.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Johnson, S.A. & Hunter, T. Kinomics: methods for deciphering the kinome. Nat. Methods 2, 17–25 (2005).
Manning, G., Whyte, D.B., Martinez, R., Hunter, T. & Sudarsanam, S. The protein kinase complement of the human genome. Science 298, 1912–1934 (2002).
Cohen, P. The role of protein phosphorylation in human health and disease. The Sir Hans Krebs Medal Lecture. Eur. J. Biochem. 268, 5001–5010 (2001).
Cohen, P. Protein kinases–the major drug targets of the twenty-first century? Nat. Rev. Drug Discov. 1, 309–315 (2002).
Berwick, D.C. & Tavare, J.M. Identifying protein kinase substrates: hunting for the organ-grinder's monkeys. Trends Biochem. Sci. 29, 227–232 (2004).
Knebel, A., Morrice, N. & Cohen, P. A novel method to identify protein kinase substrates: eEF2 kinase is phosphorylated and inhibited by SAPK4/p38delta. EMBO J. 20, 4360–4369 (2001).
Cohen, P. & Knebel, A. KESTREL: a powerful method for identifying the physiological substrates of protein kinases. Biochem. J. 393, 1–6 (2006).
Shah, K. & Shokat, K.M. A chemical genetic screen for direct v-Src substrates reveals ordered assembly of a retrograde signaling pathway. Chem. Biol. 9, 35–47 (2002).
Geahlen, R.L., Anostario, M. Jr., Low, P.S. & Harrison, M.L. Detection of protein kinase activity in sodium dodecyl sulfate-polyacrylamide gels. Anal. Biochem. 153, 151–158 (1986).
Wooten, M.W. In-gel kinase assay as a method to identify kinase substrates. Sci. STKE 2002, PL15 (2002).
Burnett, G. & Kennedy, E.P. The enzymatic phosphorylation of proteins. J. Biol. Chem. 211, 969–980 (1954).
Horoszewicz, J.S. et al. The LNCaP cell line–a new model for studies on human prostatic carcinoma. Prog. Clin. Biol. Res. 37, 115–132 (1980).
Shabb, J.B. Physiological substrates of cAMP-dependent protein kinase. Chem. Rev. 101, 2381–2411 (2001).
Litchfield, D.W. Protein kinase CK2: structure, regulation and role in cellular decisions of life and death. Biochem. J. 369, 1–15 (2003).
Cohen, P.T. & Cohen, P. Discovery of a protein phosphatase activity encoded in the genome of bacteriophage lambda. Probable identity with open reading frame 221. Biochem. J. 260, 931–934 (1989).
Sarno, S. et al. Selectivity of 4,5,6,7-tetrabromobenzotriazole, an ATP site-directed inhibitor of protein kinase CK2 ('casein kinase-2'). FEBS Lett. 496, 44–48 (2001).
Wilson, M.J. et al. Protein kinase CK2 activities in human prostatic and seminal-vesicle secretions and seminal plasma. J. Androl. 19, 754–760 (1998).
Meggio, F. & Pinna, L.A. One-thousand-and-one substrates of protein kinase CK2? FASEB J. 17, 349–368 (2003).
Kobayashi, T. et al. Regulation of cytosolic prostaglandin E synthase by phosphorylation. Biochem. J. 381, 59–69 (2004).
Okabe, M., Enomoto, M., Maeda, H., Kuroki, K. & Ohtsuki, K. Biochemical characterization of suramin as a selective inhibitor for the PKA-mediated phosphorylation of HBV core protein in vitro. Biol. Pharm. Bull. 29, 1810–1814 (2006).
Hovland, R., Doskeland, A.P., Eikhom, T.S., Robaye, B. & Doskeland, S.O. cAMP induces co-translational modification of proteins in IPC-81 cells. Biochem. J. 342, 369–377 (1999).
Zhu, H. et al. Analysis of yeast protein kinases using protein chips. Nat. Genet. 26, 283–289 (2000).
Zhu, H. & Snyder, M. Protein chip technology. Curr. Opin. Chem. Biol. 7, 55–63 (2003).
Blencowe, B.J. Alternative splicing: new insights from global analyses. Cell 126, 37–47 (2006).
Guerrera, I.C. & Kleiner, O. Application of mass spectrometry in proteomics. Biosci. Rep. 25, 71–93 (2005).
Troiani, S. et al. Searching for biomarkers of Aurora-A kinase activity: identification of in vitro substrates through a modified KESTREL approach. J. Proteome Res. 4, 1296–1303 (2005).
Brownell, J.E. & Allis, C.D. An activity gel assay detects a single, catalytically active histone acetyltransferase subunit in Tetrahymena macronuclei. Proc. Natl. Acad. Sci. USA 92, 6364–6368 (1995).
Acknowledgements
We thank D. Litchfield, (University of Western Ontario) for providing the CK2α clones, P. Jayachandran for cloning the PGES gene, and L. Mutton for technical assistance. This work was supported by grant DAMD17-03-1-0091 from the Congressionally Directed Medical Research Program, and by grant R21-CA122884 from the US National Institutes of Health, National Cancer Institute Innovative Technologies for the Molecular Analysis of Cancer program.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
C.J.B. and X.L. are named as inventors on US and PCT patent applications entitled “Devices and Methods for Profiling Enzyme Substrates” which were filed on June 6, 2006 (PCT publication number WO 2007 046882).
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5, Supplementary Tables 1–2, Supplementary Methods (PDF 2740 kb)
Rights and permissions
About this article
Cite this article
Li, X., Guan, B., Srivastava, M. et al. The reverse in-gel kinase assay to profile physiological kinase substrates. Nat Methods 4, 957–962 (2007). https://doi.org/10.1038/nmeth1106
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth1106
This article is cited by
-
Mapping phospho-catalytic dependencies of therapy-resistant tumours reveals actionable vulnerabilities
Nature Cell Biology (2019)
-
Global Screening of CK2 Kinase Substrates by an Integrated Phosphoproteomics Workflow
Scientific Reports (2013)